HIER Buffer Protocols: Optimizing Sodium Citrate, Tris, and EDTA Antigen Retrieval for IHC/ISH
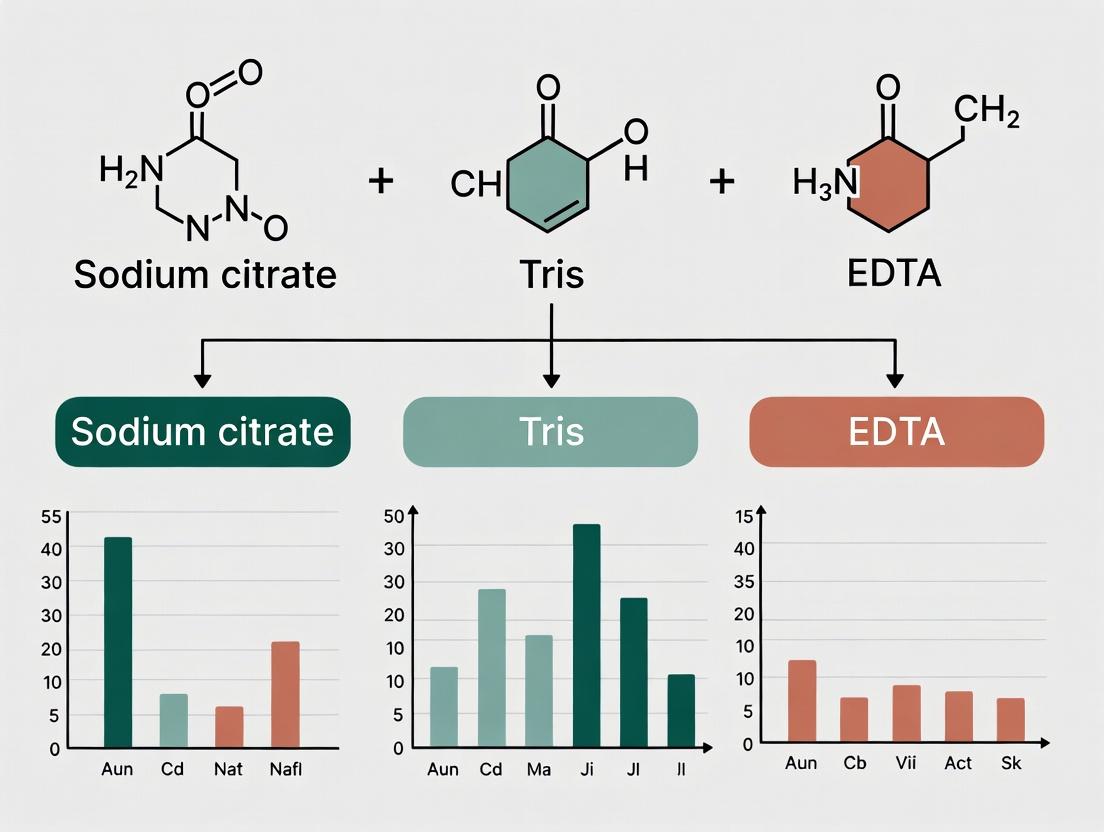
This comprehensive guide details the preparation, application, and optimization of HIER buffers, focusing on the critical roles of sodium citrate, Tris, and EDTA.
HIER Buffer Protocols: Optimizing Sodium Citrate, Tris, and EDTA Antigen Retrieval for IHC/ISH
Abstract
This comprehensive guide details the preparation, application, and optimization of HIER buffers, focusing on the critical roles of sodium citrate, Tris, and EDTA. Targeted at researchers and drug development professionals, it covers foundational chemistry, step-by-step methodological protocols, troubleshooting for common artifacts, and comparative validation against commercial kits and alternative retrieval methods. The article synthesizes current best practices to ensure reproducible, high-quality results in immunohistochemistry (IHC) and in situ hybridization (ISH) for biomedical research and clinical diagnostics.
Understanding HIER Buffers: The Science Behind Sodium Citrate, Tris, and EDTA in Antigen Unmasking
Heat-Induced Epitope Retrieval (HIER) is a foundational technique in immunohistochemistry (IHC) and immunofluorescence (IF). Formalin fixation, while preserving tissue architecture, creates methylene bridges that cross-link proteins, masking antigenic epitopes and impairing antibody binding. HIER reverses these cross-links by applying heat in a specific chemical buffer, restoring antigen immunoreactivity. This article details the application and protocols of HIER within a broader thesis investigating the efficacy of sodium citrate, Tris, and EDTA-based retrieval buffers.
Core Principles & Quantitative Buffer Comparison
The efficacy of HIER depends on the combined effect of heat energy and the buffer's chemical action. Different buffers operate via distinct mechanisms: chelation of calcium ions (EDTA) or hydrolysis of cross-links (citrate, Tris). The optimal buffer varies by antigen.
Table 1: Comparative Analysis of Common HIER Buffers
| Buffer (pH) | Primary Mechanism | Common Antigen Targets | Typical Heating Time (at 95-100°C) | Key Advantages | Considerations |
|---|---|---|---|---|---|
| Sodium Citrate (6.0) | Hydrolysis of cross-links via pH and heat | Nuclear antigens (ER, PR, p53), Cytoplasmic | 20-40 minutes | Gentle, robust for many nuclear antigens | May be insufficient for tightly cross-linked epitopes |
| Tris-EDTA (9.0) | Chelation of calcium ions & hydrolysis | Membrane proteins, Cytokeratins, CD markers | 20-40 minutes | Potent, effective for challenging epitopes | Can damage tissue morphology if overused |
| Tris-HCl (8.0-10.0) | Hydrolysis at alkaline pH | Phospho-epitopes, Viral antigens | 15-30 minutes | Good for phospho-specific antibodies | pH must be carefully optimized |
Detailed HIER Protocols
The following protocols assume the use of formalin-fixed, paraffin-embedded (FFPE) tissue sections mounted on slides.
General Deparaffinization and Rehydration (Pre-HIER)
- Materials: Xylene or xylene substitute, 100% ethanol, 95% ethanol, 70% ethanol, deionized water (dH2O).
- Protocol:
- Bake slides at 60°C for 20-60 minutes (optional but recommended).
- Deparaffinize in 3 changes of xylene, 5 minutes each.
- Rehydrate in series: 100% ethanol (2x, 3 min each) → 95% ethanol (2 min) → 70% ethanol (2 min) → dH2O (2 min).
Sodium Citrate Buffer HIER Protocol (pH 6.0)
- Buffer Preparation (10mM Sodium Citrate, 0.05% Tween 20, pH 6.0):
- Dissolve 2.94g of tri-sodium citrate dihydrate in 1L of dH2O.
- Adjust pH to 6.0 with 1N HCl.
- Add 0.5mL of Tween 20 and mix.
- HIER Procedure:
- Place slides in a coplin jar or heat-resistant slide holder filled with pre-heated sodium citrate buffer.
- Perform retrieval in a decloaking chamber/pressure cooker (125°C, 3-5 min) or a water bath (95-100°C, 20-40 min).
- After heating, cool slides in buffer at room temperature for 20-30 minutes.
- Rinse slides in dH2O, then transfer to PBS or TBS for subsequent staining.
Tris-EDTA Buffer HIER Protocol (pH 9.0)
- Buffer Preparation (10mM Tris Base, 1mM EDTA, 0.05% Tween 20, pH 9.0):
- Dissolve 1.21g Tris base and 0.37g EDTA disodium salt in 1L of dH2O.
- Adjust pH to 9.0 with 1N NaOH.
- Add 0.5mL of Tween 20 and mix.
- HIER Procedure:
- Follow steps 1-4 from the Sodium Citrate Protocol (3.2), using the Tris-EDTA buffer.
Visualization of HIER Principles & Workflow
Title: HIER Experimental Workflow in IHC
Title: Decision Tree for HIER Buffer Selection
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for HIER Buffer Preparation & Validation
| Reagent/Material | Function/Description | Key Consideration |
|---|---|---|
| Tris Base (C4H11NO3) | Alkaline buffer component. Hydrolyzes cross-links at high pH. | Use high-purity grade. pH is temperature-dependent. |
| Sodium Citrate Dihydrate (C6H5Na3O7·2H2O) | Acidic buffer component. Effective hydrolysis via heat and pH. | Common standard at 10mM concentration, pH 6.0. |
| EDTA Disodium Salt (C10H14N2Na2O8·2H2O) | Chelating agent. Binds calcium ions crucial for cross-link stability. | Use with alkaline buffers (pH 8-9) for full efficacy. |
| Tween 20 (Polysorbate 20) | Non-ionic surfactant. Reduces surface tension, improves buffer penetration. | Typically used at 0.05% v/v. Avoid excessive foaming. |
| pH Meter & Calibrated Electrodes | Critical for accurate buffer pH adjustment. | Calibrate daily with standard buffers (pH 4.0, 7.0, 10.0). |
| Decloaking Chamber/Pressure Cooker | Provides consistent, high-temperature (110-125°C) heating. | Reduces retrieval time, often improves consistency. |
| Water Bath or Steamer | Provides lower temperature (95-100°C) heating. | Gentle but requires longer incubation; monitor water level. |
| Positive Control Tissue Slides | Tissue known to express target antigen. Essential for protocol validation. | Should show strong specific staining when HIER is optimal. |
| Negative Control (No Primary Antibody) | Controls for non-specific binding of detection system. | Should show no signal when HIER is correctly performed. |
Within a broader thesis on Heat-Induced Epitope Retrieval (HIER) buffer optimization for immunohistochemistry, the precise chemical profiling of sodium citrate, Tris, and EDTA-based buffers is paramount. The efficacy of HIER in reversing formaldehyde cross-links is critically dependent on buffer pH, ionic strength, and chelation capacity. This article details the application notes and protocols for characterizing these key parameters, providing a foundation for reproducible and effective antigen retrieval in research and drug development.
Quantitative Profiles of Key HIER Buffers
The following table summarizes the critical chemical parameters for standard HIER buffer formulations at working concentrations (10-100 mM). Data is compiled from current manufacturer specifications and published studies.
Table 1: Chemical Profiles of Common HIER Buffers
| Buffer System | Typical pH Range (25°C) | pKa (25°C) | Ionic Strength (μ) @ 10 mM, pH 6.0 | Key Chelation Properties | Common HIER Concentration |
|---|---|---|---|---|---|
| Sodium Citrate | 3.0 - 6.2 | 3.13, 4.76, 6.40 | ~24 mM (as Na+) | Moderate chelator of Ca²⁺, Mg²⁺ via carboxylates. | 10 mM, pH 6.0 |
| Tris-HCl | 7.0 - 9.0 | 8.07 | ~10 mM (as Cl⁻) | Non-chelating. | Rarely used alone for HIER. |
| Tris-EDTA (TE) | 8.0 - 9.0 | Tris: 8.07; EDTA: 0, 1.5, 2.0, 2.69, 6.13, 10.37 | Variable, depends on counterions | Powerful chelation of divalent cations (Ca²⁺, Mg²⁺, Zn²⁺) via EDTA. | 10 mM Tris, 1 mM EDTA, pH 9.0 |
| Citrate-EDTA | 6.0 - 8.0 | Citrate: as above; EDTA: as above | Variable | Combined chelation from citrate and EDTA; synergistic for metal-dependent epitopes. | 10 mM Citrate, 1 mM EDTA, pH 6.0-8.0 |
Detailed Application Notes
pH & Buffer Capacity: Sodium citrate (pH 6.0) is the historical standard for HIER, effective for many epitopes. Tris-EDTA (pH 8.0-9.0) is essential for retrieving nuclear antigens and phospho-epitopes, as the higher pH and EDTA work synergistically to disrupt cross-links involving metal ions. The buffer capacity must be sufficient to maintain pH during the heating process.
Ionic Strength (I): Ionic strength impacts the electrostatic shielding between proteins and cross-links. Moderate ionic strength (e.g., from sodium citrate) can aid in breaking ionic interactions. Excessively high I may cause non-specific protein aggregation. It is calculated using I = 1/2Σcᵢzᵢ², where c is concentration and z is charge.
Chelation Properties: EDTA is a hexadentate chelator crucial for sequestering divalent cations (Zn²⁺, Ca²⁺, Mg²⁺) that stabilize protein structure and cross-links. Citrate acts as a tridentate chelator, less potent than EDTA but contributory. Chelation is critical for unmasking epitopes in metalloprotein-rich tissues.
Experimental Protocols
Protocol 1: Preparation and Standardization of 10 mM Sodium Citrate Buffer (pH 6.0)
Purpose: To prepare 1 L of a standardized, reproducible sodium citrate HIER buffer. Research Reagent Solutions & Materials:
- Tri-sodium citrate dihydrate (C₆H₅Na₃O₇·2H₂O): Primary buffering agent.
- Citric Acid (anhydrous): For pH adjustment.
- Deionized Water (18.2 MΩ·cm): Solvent to minimize interference.
- pH Meter (calibrated with pH 4.01, 7.00, 10.01 buffers): For accurate pH measurement.
- Magnetic Stirrer and Stir Bar: For mixing.
- Volumetric Flask (1 L): For precise final volume.
Methodology:
- Weigh 2.941 g of tri-sodium citrate dihydrate and transfer to a 1 L beaker.
- Add approximately 800 mL of deionized water and stir until fully dissolved.
- Using a calibrated pH meter, adjust the pH to 6.0 by dropwise addition of a 1 M citric acid solution (prepared by dissolving 19.21 g in 100 mL water).
- Transfer the solution quantitatively to a 1 L volumetric flask. Bring to the final volume with deionized water and mix thoroughly.
- Verify the final pH. Store at 4°C for up to 1 month.
Protocol 2: Determination of Effective Chelation Capacity via Metal-Sensitive Dye Assay
Purpose: To compare the relative chelation strength of citrate, EDTA, and Citrate-EDTA buffers. Research Reagent Solutions & Materials:
- Test Buffers: 10 mM Sodium Citrate (pH 6.0), 10 mM Tris / 1 mM EDTA (pH 9.0), 10 mM Citrate / 1 mM EDTA (pH 6.0).
- Magnesium Green dye (5 mM stock in DMSO): Fluorescent dye whose signal decreases upon Mg²⁺ chelation.
- MgCl₂ Solution (100 mM): Source of divalent cations.
- Microplate Reader (Fluorescence-capable): For high-throughput measurement.
- Black 96-well Plate: To minimize signal crosstalk.
Methodology:
- Prepare a 5 µM working solution of Magnesium Green in each test buffer.
- Pipette 90 µL of each dye-buffer solution into triplicate wells of a 96-well plate.
- Record initial fluorescence (λex ~506 nm, λem ~531 nm).
- Add 10 µL of 100 mM MgCl₂ solution to each well (final [Mg²⁺] = 10 mM).
- Incubate for 5 minutes at room temperature and record final fluorescence.
- Calculate: % Chelation = [1 - (Fsample / Fbuffer control)] * 100, where F_buffer control is fluorescence of dye in buffer without Mg²⁺. TE buffer will show the highest chelation capacity.
Protocol 3: HIER Protocol for FFPE Tissue Sections Using Optimized Buffers
Purpose: To perform antigen retrieval for immunohistochemistry using chemically profiled buffers. Research Reagent Solutions & Materials:
- Prepared HIER Buffer (e.g., Sodium Citrate pH 6.0 or Tris-EDTA pH 9.0): Retrieval medium.
- Deparaffinized & Rehydrated FFPE Tissue Sections on slides: Sample.
- Pressure Cooker or Decloaking Chamber: For standardized heating.
- Slide Rack and Coplin Jars: For holding slides.
- PBS (pH 7.4): For washing.
Methodology:
- Preheat the pressure cooker containing the chosen HIER buffer (enough to cover slides).
- Place slides in a slide rack and submerge in the preheated buffer.
- Process under pressure for a standardized time (e.g., 2.5 minutes at full pressure for citrate buffer).
- Carefully remove the container and allow it to cool at room temperature for 20-30 minutes.
- Transfer slides to Coplin jars and wash 3 x 5 minutes in PBS.
- Proceed immediately with standard immunohistochemistry staining protocols.
Visualizations
Diagram Title: HIER Buffer Selection & Mechanism Flowchart
Diagram Title: Chelation Capacity Assay Workflow
Heat-Induced Epitope Retrieval (HIER) is a cornerstone technique in immunohistochemistry (IHC) that reverses formaldehyde-induced cross-links, enabling antibody binding to masked epitopes. The selection of retrieval buffer directly impacts staining intensity, specificity, and background. Within the broader thesis on HIER buffer preparation (sodium citrate vs. Tris-EDTA), sodium citrate buffer (pH 6.0) emerges as the empirically validated gold standard for the majority of IHC applications. Its mild acidic nature is optimal for preserving tissue morphology while effectively retrieving a wide array of nuclear, cytoplasmic, and membranous antigens.
Comparative Performance Data of HIER Buffers
The following table summarizes key quantitative performance metrics from recent studies comparing sodium citrate (pH 6.0) with other common HIER buffers.
Table 1: Comparative Analysis of HIER Buffer Performance for General IHC
| Buffer Type & pH | Optimal Antigen Classes | Median Retrieval Efficacy Score (1-10) | Tissue Morphology Preservation (1-5) | Reported Use in Key Publications (%) |
|---|---|---|---|---|
| Sodium Citrate, pH 6.0 | Broad-spectrum (Nuclear: ER, p53; Cytoplasmic: CK; Membranous: HER2) | 9.2 | 4.5 | 68% |
| Tris-EDTA, pH 9.0 | Phospho-epitopes, Challenging nuclear antigens (e.g., FoxP3) | 8.7 | 4.0 | 25% |
| EDTA, pH 8.0 | Tightly cross-linked epitopes | 8.5 | 3.8 | 5% |
| Glycine-HCl, pH 3.5 | Select viral antigens | 6.0 | 4.2 | <2% |
Efficacy Score: Composite metric of staining intensity and specificity. Morphology: 5=Excellent, 1=Poor. Data compiled from published literature (2020-2024).
Detailed Protocol: Preparation and Use of Sodium Citrate Buffer (pH 6.0)
Reagent Solutions and Materials
Table 2: The Scientist's Toolkit for Sodium Citrate HIER
| Item | Function in Protocol | Specification/Note |
|---|---|---|
| Sodium Citrate, Dihydrate | Buffer component, provides chelating action and pH stability | ACS grade, 99.0% minimum purity |
| Citric Acid, Anhydrous | pH adjustment | Used for fine-tuning pH to 6.0 ± 0.1 |
| Deionized Water | Solvent | Nuclease-free, 18.2 MΩ·cm resistivity |
| pH Meter | Accurate pH measurement | Calibrated daily with pH 4.01, 7.00, 10.01 buffers |
| Microwave Oven or Pressure Cooker | Heat source for retrieval | Must provide consistent, controllable heating |
| Glass or Plastic Coplin Jars | Container for slides during retrieval | Chemically resistant, dedicated to IHC |
| Target Retrieval Solution (Commercial) | Optional comparison/backup | Pre-mixed, standardized formulation |
Buffer Preparation Protocol
Title: Preparation of 10x Sodium Citrate Stock Solution (100 mM, pH 6.0)
- Weigh 29.41 g of trisodium citrate dihydrate (C₆H₅Na₃O₇·2H₂O, MW 294.1).
- Add to a 1 L graduated beaker containing approximately 800 mL of deionized water.
- Stir on a magnetic stirrer until completely dissolved.
- Using a calibrated pH meter, adjust the pH to 6.0 by adding drops of 1M citric acid solution (21.0 g/L in water). Do not use HCl.
- Transfer the solution to a 1 L volumetric flask and bring to volume with deionized water.
- Filter the 10x stock through a 0.22 μm membrane. Store at 4°C for up to 6 months.
- For working solution (10 mM), dilute 100 mL of 10x stock with 900 mL deionized water. Check pH before use.
Standard HIER Protocol Using Sodium Citrate Buffer
Title: IHC Heat-Induced Epitope Retrieval with Sodium Citrate
- Dewax and Hydrate: Process formalin-fixed, paraffin-embedded (FFPE) tissue sections through xylene and graded ethanol series to water.
- Buffer Pre-heat: Preheat the sodium citrate working solution (10 mM, pH 6.0) in a microwave-safe container or pressure cooker. For microwave: Heat to 95-100°C. For pressure cooker: Bring to a boil.
- Retrieval:
- Microwave Method: Place slides in pre-heated buffer. Microwave at 95-100°C for 15-20 minutes, ensuring slides remain submerged. Replace evaporated water with pre-warmed deionized water.
- Pressure Cooker Method: Place slides in boiling buffer. Secure lid and bring to full pressure. Maintain at full pressure for 2-5 minutes (optimize per antibody).
- Water Bath/Steamer Method: Incubate at 95-100°C for 30-40 minutes.
- Cooling: After heating, remove the container from the heat source and allow slides to cool in the buffer for 20-30 minutes at room temperature.
- Rinse: Rinse slides thoroughly with deionized water, then proceed to PBS or TBS wash.
- Immunostaining: Continue with standard IHC protocol (peroxide block, protein block, primary/secondary antibody, detection, counterstain, mount).
Experimental Workflow and Buffer Selection Logic
Title: HIER Buffer Selection Workflow for IHC
Mechanism of Action and Comparison Diagram
Title: HIER Buffer Mechanisms: Citrate vs. Tris-EDTA
Application Notes
Heat-Induced Epitope Retrieval (HIER) is a critical step in immunohistochemistry (IHC) for unmasking antigens cross-linked by formalin fixation. While sodium citrate buffer (pH 6.0) is a standard retrieval solution, many challenging targets, particularly nuclear antigens, phosphorylated epitopes, and proteins in densely cross-linked tissues, require more robust retrieval conditions. Tris-EDTA and its high-pH variant (pH 9.0) provide an effective alternative by utilizing alkaline conditions and chelation to reverse formaldehyde adducts and extract calcium ions, respectively. This approach is essential within the broader research thesis on optimizing HIER buffer systems (sodium citrate vs. Tris-EDTA) for maximal signal-to-noise ratios across diverse biomarker panels.
Tris-EDTA buffers work through a dual mechanism:
- Alkaline Hydrolysis: The high pH (8.0-9.0) promotes hydrolysis of methylene bridges formed during fixation.
- Chelation: EDTA chelates divalent cations (e.g., Ca²⁺, Zn²⁺) that are involved in stabilizing protein structures and cross-links, further loosening the tissue matrix.
The selection between Tris-EDTA (pH 8.0) and Tris-EDTA (pH 9.0) is antigen-dependent. A pH of 9.0 is often necessary for the most recalcitrant nuclear targets, such as certain transcription factors, but may increase background for some cytoplasmic antigens.
Quantitative Comparison of Common HIER Buffers
The following table summarizes key performance data for standard retrieval buffers, synthesized from current literature and laboratory protocols.
Table 1: Comparative Analysis of Common HIER Buffers
| Buffer Formulation | Typical pH Range | Optimal Antigen Types | Typical Retrieval Time/Temp | Key Advantages | Potential Limitations |
|---|---|---|---|---|---|
| Sodium Citrate | 6.0 | Many cytoplasmic and membrane antigens (e.g., CD markers, cytokeratins) | 20-40 min at 95-100°C | Gentle, low background; widely established standard. | Ineffective for many nuclear and phosphorylated targets. |
| Tris-EDTA | 8.0 - 8.5 | A broad range, including some nuclear antigens (e.g., ER, PR) and select phospho-epitopes. | 20-40 min at 95-100°C | More effective than citrate for many targets; good balance of potency and tissue integrity. | May not be sufficient for highly cross-linked epitopes. |
| Tris-EDTA (pH 9.0) | 9.0 - 9.5 | Challenging nuclear targets (e.g., FoxP3, p53), many phosphorylated proteins (e.g., pSTATs), and antigens in over-fixed tissue. | 15-30 min at 95-100°C or 10-20 min at 110-121°C (pressure cooker). | High retrieval power for difficult epitopes; often the last resort for unsuccessful retrieval. | Can increase non-specific background; may damage tissue morphology if overheated. |
| EDTA-only | 8.0 - 9.0 | Very challenging nuclear antigens. | 20-40 min at 95-100°C | Powerful chelation can be effective where other buffers fail. | Can be harsh on tissue morphology; requires careful optimization. |
Detailed Protocols
Protocol 1: Preparation of Tris-EDTA Buffers (pH 8.0 and pH 9.0)
Research Reagent Solutions & Materials:
- Tris base (C₄H₁₁NO₃): Provides the alkaline buffer component.
- EDTA, Disodium Salt, Dihydrate (C₁₀H₁₄N₂Na₂O₈·2H₂O): Chelating agent to bind metal ions.
- HCl (1M or concentrated): For pH adjustment.
- Deionized or Distilled Water: Solvent.
- pH Meter: For accurate pH adjustment.
- Beaker and Stir Plate: For mixing.
- Volumetric Flask/Graduated Cylinder: For accurate volume measurement.
Procedure:
- For 1L of 10x Tris-EDTA Stock Solution:
- Weigh 60.55 g of Tris base and 37.2 g of EDTA disodium salt dihydrate.
- Add to a beaker containing approximately 800 mL of deionized water. Stir vigorously using a magnetic stirrer to dissolve.
- Adjust the pH to the desired target (8.0 or 9.0) using concentrated HCl. Note: Adding HCl is exothermic; allow the solution to cool before final pH measurement.
- Once the pH is stable at the target value, transfer the solution to a 1L volumetric flask and bring to the final volume with deionized water. Mix thoroughly.
- For working solution, dilute the 10x stock 1:10 with deionized water (e.g., 100 mL stock + 900 mL water). Confirm the pH of the 1x working solution.
- Store at 4°C for up to 6 months.
Protocol 2: HIER Using Tris-EDTA (pH 9.0) for Challenging Nuclear Antigens (e.g., FoxP3)
Research Reagent Solutions & Materials:
- Pre-cut formalin-fixed, paraffin-embedded (FFPE) tissue sections on charged slides.
- Tris-EDTA (pH 9.0) 1x Working Solution (from Protocol 1).
- Slide staining rack and glass or plastic Coplin jars, OR a dedicated decloaking chamber/pressure cooker.
- Microwave, steamer, or commercial antigen retrieval system.
- PBS (pH 7.4): For washing.
- ImmEdge Hydrophobic Barrier Pen: To create a liquid barrier around sections.
Workflow:
- Dewax and Hydrate: Deparaffinize slides in xylene (or substitute) and rehydrate through a graded ethanol series (100%, 95%, 70%) to distilled water.
- Antigen Retrieval:
- Fill a Coplin jar or appropriate container with ~200-250 mL of Tris-EDTA (pH 9.0) working solution. Place slides in a staining rack and submerge.
- Using a Microwave: Heat at full power (~900W) until the solution boils (approx. 3-5 min), then reduce power to 20-30% to maintain a gentle boil for 15 minutes. Ensure slides remain submerged; replenish with pre-warmed buffer if needed.
- Using a Pressure Cooker/Decloaker: Heat until the internal temperature reaches 110-121°C and maintain for 10 minutes. Allow the system to cool naturally under pressure for 20-30 minutes before opening.
- Cooling: Remove the jar from the heat source and allow it to cool at room temperature for 20-30 minutes. Do not cool rapidly on ice, as this may promote non-specific antibody binding.
- Wash: Carefully transfer the slides to a wash bath containing PBS (pH 7.4). Wash for 5 minutes.
- Proceed with Staining: Continue with the standard protocol for immunohistochemistry (blocking, primary antibody incubation, detection, counterstaining, dehydration, and mounting).
Visualizations
HIER with Tris-EDTA (pH 9.0) Mechanism
Tris-EDTA HIER Experimental Workflow
The Scientist's Toolkit: Key Reagents for Tris-EDTA HIER
| Item | Function in Protocol |
|---|---|
| Tris Base | The primary buffering agent that maintains the alkaline pH (8.0-9.0) critical for hydrolyzing formalin-induced cross-links. |
| EDTA (Disodium Salt) | A chelating agent that binds to divalent cations (Ca²⁺, Mg²⁺, Zn²⁺), destabilizing protein complexes and aiding in epitope unmasking. |
| Hydrochloric Acid (HCl) | Used to titrate and precisely adjust the pH of the Tris-EDTA buffer to the required target value (8.0 or 9.0). |
| pH Meter | Essential laboratory instrument for accurately measuring and adjusting the pH of retrieval buffers. Calibration is critical. |
| Pressure Cooker / Decloaking Chamber | Preferred heating device for high-temperature (110-121°C) retrieval, providing consistent and powerful unmasking for difficult targets. |
| Microwave or Steamer | Alternative heating devices for performing HIER at 95-100°C. Must be capable of maintaining a stable, gentle boil. |
| Hydrophobic Barrier Pen | Used to draw a barrier around tissue sections on slides, minimizing reagent volume requirements during subsequent IHC steps. |
| Charged Microscope Slides | Provide strong adhesion for FFPE tissue sections, preventing detachment during the high-temperature, high-pH retrieval process. |
Within the broader thesis on HIER buffer optimization, the choice between sodium citrate and Tris-EDTA based antigen retrieval (AR) buffers is a fundamental determinant of immunohistochemistry (IHC) success. This document provides application notes and protocols, derived from current research, to guide researchers in selecting the optimal buffer for specific antigen targets based on their chemical stabilization mechanism.
Core Buffer Properties & Selection Guidelines
Table 1: Comparative Properties of Citrate and Tris-EDTA HIER Buffers
| Property | Sodium Citrate Buffer (pH 6.0) | Tris-EDTA Buffer (pH 9.0) |
|---|---|---|
| Typical pH | 6.0 | 8.0 - 9.0 |
| Primary Chelator | Citrate ions (weaker) | EDTA (strong) |
| Key Mechanism | Breaks protein cross-links via moderate calcium chelation and heat. | Strong chelation of divalent cations (Ca²⁺, Mg²⁺, Zn²⁺); disrupts stronger cross-links. |
| Optimal For | Formalin-induced methylene bridges; many nuclear and cytoplasmic antigens (e.g., ER, PR, Ki-67). | Zinc-finger proteins, tightly cross-linked epitopes, many transmembrane proteins (e.g., p53, CD44, MUC1). |
| Epitope Recovery | Effective for ~70-80% of common antigens. | Critical for ~20-30% of antigens unreactive with citrate. |
| Tissue Morphology | Excellent preservation. | Good preservation; can be harsher on delicate tissues. |
Decision Workflow: Begin with citrate buffer (pH 6.0) for routine IHC. If staining is weak or negative, switch to Tris-EDTA (pH 9.0), especially for nuclear transcription factors, phosphorylated epitopes, or tightly fixed membrane proteins.
Detailed Experimental Protocols
Protocol 1: Preparation and Use of 10mM Sodium Citrate Buffer (pH 6.0)
- Purpose: Recovery of most formaldehyde-fixed, paraffin-embedded (FFPE) antigens.
- Reagents: Tri-sodium citrate dihydrate, distilled water, HCl/NaOH for pH adjustment.
- Procedure:
- Dissolve 2.94 g of tri-sodium citrate dihydrate in 1 L of distilled water to make a 10 mM solution.
- Adjust pH to 6.0 ± 0.1 using 1M HCl or 1M NaOH.
- For AR, fill a plastic Coplin jar with buffer, ensuring slides are fully immersed.
- Heat in a microwave or pressure cooker: Microwave: 20 min at full power, maintaining boiling; replenish evaporated water. Pressure Cooker: 3-5 min at full pressure (≈121°C).
- Cool slides in the buffer for 20-30 minutes at room temperature before proceeding to immunohistochemistry staining.
Protocol 2: Preparation and Use of 1mM EDTA Buffer (pH 8.0-9.0)
- Purpose: Retrieval of challenging antigens, especially those stabilized by zinc or strong cross-links.
- Reagents: EDTA disodium salt, Tris Base, distilled water.
- Procedure:
- To 900 mL distilled water, add 0.37 g EDTA disodium salt (1 mM final) and 0.61 g Tris Base (5 mM final).
- Stir to dissolve and adjust pH to 8.0 or 9.0 using NaOH, as required by the target antigen.
- Bring final volume to 1 L.
- Perform heat-induced retrieval as described in Protocol 1 (steps 3-5).
Protocol 3: Sequential Buffer Testing for Optimal Antigen Retrieval
- Purpose: Empirically determine the optimal AR buffer for a novel or recalcitrant antigen.
- Procedure:
- Cut consecutive sections from the same FFPE block onto charged slides.
- Deparaffinize and hydrate all slides simultaneously.
- Divide slides into three groups: Group A (Citrate, pH 6.0), Group B (Tris-EDTA, pH 9.0), Group C (Optional: Tris-EDTA, pH 8.0).
- Perform HIER on each group using its respective buffer in separate containers.
- Process all slides identically through the same IHC run (primary antibody, detection, chromogen).
- Compare staining intensity, specificity, and background across groups to identify the optimal buffer.
Visualizations
Title: HIER Buffer Selection Workflow for IHC
Title: Chemical Mechanisms of Antigen Retrieval Buffers
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for HIER Optimization Experiments
| Reagent/Material | Function in HIER Protocol |
|---|---|
| Tri-Sodium Citrate Dihydrate | Primary component of citrate buffer; weak chelator that reverses formaldehyde cross-links at low pH. |
| Tris Base (Tris(hydroxymethyl)aminomethane) | Provides buffering capacity for high-pH Tris-EDTA solutions, maintaining stable alkaline conditions. |
| EDTA (Ethylenediaminetetraacetic Acid) Disodium Salt | Strong chelator of divalent cations (Ca²⁺, Mg²⁺, Zn²⁺); critical for disrupting metal-ion stabilized protein complexes. |
| pH Meter & Standard Buffers | Essential for precise pH adjustment of retrieval buffers, a critical parameter for efficacy. |
| HIER-Compatible Slide Holder & Coplin Jars | Withstand high heat and pressure; ensure uniform buffer exposure and prevent slide damage. |
| Pressure Cooker or Commercial Decloaking Chamber | Provides consistent, high-temperature heating (≈121°C) for uniform and robust antigen retrieval. |
| Heat-Resistant Plastic Container | For microwave-based HIER; prevents buffer boil-over and allows for safe handling. |
| Target Retrieval Validation Antibody Set | Positive control antibodies for known citrate-sensitive (e.g., ER) and EDTA-requiring (e.g., p53) antigens. |
Step-by-Step Guide: Preparing and Applying HIER Buffers for Robust IHC/ISH Staining
Within the broader thesis on Heat-Induced Epitope Retrieval (HIER) buffer preparation, the standardization of stock solutions is paramount for reproducibility in immunohistochemistry (IHC) and molecular biology. 10x Sodium Citrate (pH 6.0) and 10x Tris-EDTA (TE, pH 9.0) are fundamental buffers used for antigen retrieval and nucleic acid handling, respectively. This protocol provides detailed, reliable methods for preparing these critical stock solutions, ensuring consistency for researchers, scientists, and drug development professionals.
The Scientist's Toolkit: Essential Research Reagent Solutions
| Item | Function in Protocol / Field |
|---|---|
| Tris(hydroxymethyl)aminomethane (Tris) | A primary amine buffer, maintains alkaline pH in TE buffer essential for DNA stability. |
| Ethylenediaminetetraacetic Acid (EDTA) | A chelating agent that binds divalent cations (Mg²⁺), inactivating nucleases to protect nucleic acids. |
| Sodium Citrate Dihydrate | The buffering component for the antigen retrieval solution; the citrate ions are responsible for chelation and pH stabilization. |
| Hydrochloric Acid (HCl) | Used for precise downward adjustment of pH for both buffer solutions. |
| Sodium Hydroxide (NaOH) | Used for precise upward adjustment of pH for both buffer solutions. |
| Nuclease-free Water | Water treated to remove nucleases, essential for preparing TE buffer to prevent nucleic acid degradation. |
| pH Meter | Critical instrument for accurate pH adjustment to the specified target values (pH 6.0 ± 0.1, pH 9.0 ± 0.1). |
| Autoclave | Used for sterilizing solutions (TE buffer) and ensuring they are nuclease-free for long-term storage. |
Detailed Experimental Protocols
Protocol 1: Preparation of 10x Sodium Citrate Buffer (pH 6.0)
Principle: This buffer provides a mildly acidic, chelating environment optimal for unlocking many protein epitopes masked by formalin fixation during HIER.
Materials:
- Sodium citrate tribasic dihydrate (C₆H₅Na₃O₇ · 2H₂O)
- Hydrochloric acid (HCl), concentrated and/or 1M solution
- Sodium hydroxide (NaOH), 1M solution (if needed for minor adjustment)
- Deionized or distilled water
- pH meter, calibrated
- Beaker (1 L), stir bar, graduated cylinder
- Volumetric flask (1 L) or measuring cylinder
- Sterile bottle for storage
Method:
- Weigh 29.41 g of sodium citrate tribasic dihydrate.
- Transfer to a 1 L beaker and add approximately 800 mL of deionized water. Stir on a magnetic stirrer until completely dissolved.
- Using a calibrated pH meter, adjust the pH of the solution to 6.0. This is typically done by carefully adding concentrated HCl dropwise while stirring. Caution: The initial solution is alkaline; adding acid will lower the pH. Monitor closely.
- Once pH 6.0 ± 0.1 is achieved, transfer the solution quantitatively to a 1 L graduated cylinder. Bring the final volume to 1000 mL with deionized water. Mix thoroughly.
- Filter the solution through a 0.22 μm or 0.45 μm membrane filter into a sterile bottle. Label with name, concentration (10x), pH 6.0, and date.
- Store at room temperature. For 1x working solution, dilute 1 part 10x stock with 9 parts deionized water.
Protocol 2: Preparation of 10x Tris-EDTA (TE) Buffer (pH 9.0)
Principle: TE buffer maintains DNA and RNA integrity by providing an alkaline pH (from Tris) and chelating nuclease cofactors (via EDTA). The 10x stock is concentrated for convenience and dilution flexibility.
Materials:
- Tris(hydroxymethyl)aminomethane (Tris base)
- Ethylenediaminetetraacetic acid disodium salt dihydrate (EDTA-Na₂·2H₂O)
- Hydrochloric acid (HCl), concentrated and/or 1M solution
- Nuclease-free water
- pH meter, calibrated
- Beaker (500 mL), stir bar
- Volumetric flask (500 mL)
- Autoclave and sterile bottles
Method:
- Weigh 60.55 g of Tris base and 18.61 g of EDTA disodium salt dihydrate.
- Transfer to a 500 mL beaker and add approximately 400 mL of nuclease-free water. Stir vigorously to dissolve. EDTA will require some time and may need slight pH adjustment (>8.0) to fully dissolve.
- Using a calibrated pH meter, adjust the pH of the solution to 9.0 by carefully adding concentrated HCl dropwise while stirring.
- Once pH 9.0 ± 0.1 is achieved, quantitatively transfer the solution to a 500 mL volumetric flask. Bring the final volume to 500 mL with nuclease-free water. Mix thoroughly.
- Dispense the solution into suitable bottles and autoclave at 121°C for 20 minutes for sterilization. Alternatively, filter sterilize through a 0.22 μm membrane.
- Label with name, concentration (10x TE), pH 9.0, and date. Store at room temperature. For 1x TE buffer, dilute 1:10 with nuclease-free water.
Data Presentation: Buffer Composition and Properties
Table 1: 10x Stock Buffer Compositions and Final 1x Concentrations
| Buffer | 10x Stock Formula (per Liter) | Final 1x Working Concentration | Primary Function |
|---|---|---|---|
| Sodium Citrate, pH 6.0 | 29.41 g Sodium Citrate · 2H₂O | 10 mM Sodium Citrate, pH 6.0 | Antigen retrieval for IHC (HIER). |
| Tris-EDTA (TE), pH 9.0 | 121.1 g Tris base, 37.22 g EDTA-Na₂·2H₂O | 10 mM Tris-HCl, 1 mM EDTA, pH 9.0 | Nucleic acid solubilization and storage; antigen retrieval for select epitopes. |
Table 2: Key Buffer Properties and Storage Guidelines
| Property | 10x Sodium Citrate (pH 6.0) | 10x Tris-EDTA (pH 9.0) |
|---|---|---|
| Target pH | 6.0 ± 0.1 | 9.0 ± 0.1 |
| Sterilization Method | Filtration (0.22/0.45 μm) | Autoclaving or filtration |
| Recommended Storage | Room temperature, sterile | Room temperature, sterile |
| Shelf Life | 6-12 months | >1 year |
| Critical QC Step | pH verification after dilution to 1x | Verification of nuclease-free status (gel electrophoresis). |
Experimental Workflow and Context
Working Solution Preparation, Storage, and Shelf-Life Considerations
This document provides application notes and protocols for the preparation, storage, and stability assessment of working solutions critical to immunohistochemical research, specifically within the context of a broader thesis investigating Heat-Induced Epitope Retrieval (HIER) buffers, focusing on sodium citrate, Tris, and EDTA formulations. The reliability of experimental outcomes in epitope retrieval studies is fundamentally dependent on the consistency and stability of these working solutions. These protocols are designed for researchers and drug development professionals to ensure reproducibility and data integrity in longitudinal studies.
Key Research Reagent Solutions
| Reagent Solution | Primary Function in HIER Research | Critical Storage Parameter |
|---|---|---|
| 10x Sodium Citrate Buffer (pH 6.0) | Standard HIER buffer for a wide range of antigens. Provides acidic pH and chelating activity. | Store at 15-25°C; stable for 12 months. Protect from CO₂ absorption. |
| 10x Tris-EDTA Buffer (pH 9.0) | High-pH HIER buffer for challenging nuclear and phospho-antigens. EDTA enhances chelation. | Store at 15-25°C; stable for 12 months. |
| 1x Working HIER Buffer | Diluted ready-to-use buffer for the retrieval process. Stability is concentration-dependent. | Store at 2-8°C; recommended shelf-life 1 month. |
| Protease Inhibitor Cocktail (100x) | Added to working buffers for labile epitopes to prevent protein degradation during retrieval. | Store at -20°C; stable for 24 months. After thawing, store at 2-8°C for 2 weeks. |
| Antibody Diluent, Buffered | Preserves antibody stability and prevents non-specific binding during post-retrieval incubations. | Store at 2-8°C; stable as per manufacturer (typically 6-12 months). |
Quantitative Stability Data for Common HIER Buffers
Table 1: Shelf-life of Concentrated (10x) Stock HIER Buffers at Room Temperature (15-25°C)
| Buffer Formulation | Initial pH | pH at 12 Months | Visible Precipitation | Recommended Max Shelf-Life |
|---|---|---|---|---|
| 10mM Sodium Citrate, 0.05% Tween 20, pH 6.0 | 6.0 ± 0.1 | 5.9 ± 0.2 | No | 12 months |
| 100mM Tris Base, 10mM EDTA, 0.05% Tween 20, pH 9.0 | 9.0 ± 0.1 | 8.7 ± 0.3 | No (if filtered) | 12 months |
| 1mM EDTA, 0.05% Tween 20, pH 8.0 | 8.0 ± 0.1 | 7.8 ± 0.2 | No | 12 months |
Table 2: Stability of 1x Working Solution Buffers at 2-8°C
| Working Solution | Initial Parameter | Parameter at 30 Days | Recommended Shelf-Life | Key Degradation Sign |
|---|---|---|---|---|
| 1x Sodium Citrate (pH 6.0) | pH 6.0, Clear | pH 6.1, Clear | 30 days | Microbial growth, pH drift >0.3 |
| 1x Tris-EDTA (pH 9.0) | pH 9.0, Clear | pH 8.6, Clear | 14 days | pH drift due to CO₂ absorption |
| 1x Buffer + Protease Inhibitors | Full Activity | ~80% Activity | 7 days | Protease inhibitor oxidation/deactivation |
Experimental Protocols
Protocol 3.1: Preparation of 10x Sodium Citrate Stock Buffer (1 L, pH 6.0)
Purpose: To prepare a stable, concentrated stock for HIER. Materials: Tri-sodium citrate dihydrate, Citric acid anhydrous, Tween 20, DI water, pH meter. Method:
- Dissolve 29.41 g of tri-sodium citrate dihydrate in 900 mL of deionized water with stirring.
- Adjust to pH 6.0 using a 1M solution of citric acid (approximately 4-5 mL required).
- Add 0.5 mL of Tween 20.
- Bring the final volume to 1 L with deionized water.
- Filter through a 0.22 µm PES membrane into a sterile, sealed container.
- Label with date, pH, and formulation. Store at room temperature.
Protocol 3.2: Accelerated Stability Testing for Working Solutions
Purpose: To empirically determine the shelf-life of a 1x working HIER buffer. Materials: Freshly prepared 1x working buffer, sealed containers, pH meter, spectrophotometer, microbial growth plates. Method:
- Aliquot Preparation: Aseptically prepare twelve 50 mL aliquots of the 1x buffer.
- Storage Conditions: Store nine aliquots at 2-8°C, and three aliquots at 25°C (accelerated condition).
- Time Points: Test aliquots at t=0, 7, 14, 30, 60, and 90 days (for 2-8°C) and at t=0, 3, and 7 days (for 25°C).
- Test Parameters: a. pH Measurement: Using a calibrated pH meter. b. Clarity/Turbidity: Visual inspection and absorbance at 600 nm (A600 < 0.1 acceptable). c. Microbial Testing: Plate 100 µL onto LB agar, incubate at 37°C for 48h. d. Functional Assay: Use the buffer in a standard HIER/IHC protocol with a control tissue and antibody. Score staining intensity and background.
- Analysis: Shelf-life is the longest time point before a critical parameter (pH drift >±0.5, microbial growth, significant loss of staining intensity) fails.
Protocol 3.3: HIER Procedure Using Prepared Working Buffer
Purpose: Standardized epitope retrieval for immunohistochemistry. Materials: Deparaffinized tissue sections, 1x working HIER buffer, heat source (water bath, steamer, or pressure cooker), staining racks/coplin jars. Method:
- Pre-heat the HIER buffer (typically 200-500 mL) in a dedicated plastic coplin jar or container in a 95-100°C water bath, steamer, or pressure cooker until the target temperature is reached.
- Place slide racks with deparaffinized and rehydrated tissue sections into the pre-heated buffer.
- Incubate for 20 minutes (water bath/steamer) or 5 minutes (pressure cooker at 15 psi). Maintain consistent temperature.
- After retrieval, remove the container from heat and cool at room temperature for 20-30 minutes.
- Rinse slides gently in deionized water. Proceed immediately with immunohistochemical staining.
Visualizations
Diagram Title: Workflow for HIER Buffer Stability Assessment in Thesis Research
Diagram Title: Buffer Degradation Pathways and Experimental Impact
1. Introduction & Thesis Context This document provides standardized protocols for Heat-Induced Epitope Retrieval (HIER), a critical step in immunohistochemistry (IHC) for formalin-fixed, paraffin-embedded (FFPE) tissues. The efficacy of HIER is fundamentally governed by the chemical action of the retrieval buffer and the kinetics of heat application. Within the broader thesis investigating the mechanistic interplay between buffer composition (sodium citrate vs. Tris-EDTA) and retrieval method, these protocols establish reproducible parameters for the pressure cooker, microwave, and water bath techniques. Optimization of this step is paramount for researchers and drug development professionals validating pharmacodynamic biomarkers in clinical trials.
2. Research Reagent Solutions & Essential Materials The following table details key reagents and materials essential for executing standardized HIER.
| Item Name | Function/Brief Explanation |
|---|---|
| 10mM Sodium Citrate Buffer (pH 6.0) | A common acidic HIER buffer, effective for many nuclear and cytoplasmic antigens. Chelates calcium. |
| 1mM Tris-EDTA Buffer (pH 9.0) | A common alkaline HIER buffer, often superior for phosphorylated epitopes and more robust cross-linking. |
| FFPE Tissue Sections | 3-5 µm sections mounted on charged or adhesive slides. |
| Pressure Cooker (Domestic) | Provides super-heated retrieval solution (>100°C, typically 120-125°C) under pressure for rapid, uniform retrieval. |
| Variable-Power Microwave | Enables controlled boiling of retrieval solution. Requires power cycling to prevent drying. |
| Temperature-Controlled Water Bath | Provides gentle, precisely controlled sub-boiling retrieval (92-98°C). |
| Slide Rack & Coplin Jars | For holding slides during retrieval and subsequent washing steps. |
| Humidified Slide Chamber | For incubating slides with primary antibody post-retrieval. |
3. Quantitative Data Summary: Method Comparison The following table summarizes key operational parameters and performance characteristics for the three HIER methods.
| Parameter | Pressure Cooker | Microwave | Water Bath |
|---|---|---|---|
| Typical Temperature | 120-125°C | ~95-100°C (intermittent boiling) | 92-98°C |
| Processing Time | 1-5 minutes at pressure | 15-20 minutes (cycled) | 20-45 minutes |
| Buffer Volume | High (1.5-2L) | Low (150-250mL) | Medium (500mL-1L) |
| Uniformity | High | Variable (hot/cold spots) | High |
| Intensity (Typical) | High | Moderate to High | Moderate |
| Background Risk | Moderate | Moderate to High (if dried) | Low |
| Best For | Dense/tough tissues, strong cross-linking | Routine, rapid processing | Delicate epitopes, phospho-targets |
4. Detailed Experimental Protocols
Protocol 4.1: Pressure Cooker Method
- Equipment: Domestic electric pressure cooker, slide rack, plastic staining dish.
- Buffer: 1.5L of pre-heated 10mM Sodium Citrate (pH 6.0) or 1mM Tris-EDTA (pH 9.0).
- Procedure:
- Deparaffinize and hydrate slides to distilled water.
- Place buffer in cooker. Insert slide rack. Bring to a simmer (without lid).
- Carefully place slides into the rack. Secure the lid, ensuring the steam vent is closed.
- Once full pressure is reached (typically indicated by a float valve), start timer for 2 minutes.
- After 2 minutes, use the quick-release function per manufacturer's instructions.
- Carefully open lid away from face. Let slides cool in buffer for 20 minutes at room temperature.
- Transfer slides to distilled water, then proceed to immunohistochemical staining.
Protocol 4.2: Microwave Method
- Equipment: 800W microwave, plastic Coplin jars or slide-staining dish with loose lid.
- Buffer: 200mL of 10mM Sodium Citrate (pH 6.0) or 1mM Tris-EDTA (pH 9.0).
- Procedure:
- Deparaffinize and hydrate slides to distilled water.
- Place slides in jar with buffer. Loosely cover to prevent excessive evaporation.
- Microwave at 100% power until boiling (~2-3 mins).
- Reduce power to 20-30% to maintain a gentle boil. Incubate for 15 minutes.
- Monitor buffer level closely. Add pre-warmed distilled water if evaporation is significant.
- Carefully remove jar. Cool slides in buffer at room temperature for 20-30 minutes.
- Rinse in distilled water and proceed with staining.
Protocol 4.3: Water Bath Method
- Equipment: Precision-controlled water bath, glass Coplin jars.
- Buffer: 500mL of 10mM Sodium Citrate (pH 6.0) or 1mM Tris-EDTA (pH 9.0).
- Procedure:
- Deparaffinize and hydrate slides to distilled water.
- Pre-heat buffer in Coplin jars within the water bath to 95°C.
- Place slides into the pre-heated buffer.
- Incubate for 25 minutes at 95°C (±2°C). Ensure slides remain fully submerged.
- Remove the jar from the bath and cool at room temperature for 20 minutes.
- Rinse slides in cool distilled water before staining.
5. Visualization: HIER Method Selection Workflow
Title: Decision Workflow for HIER Method Selection
6. Visualization: Thesis Context of HIER Optimization
Title: Thesis Framework for HIER Buffer & Method Research
Within the broader thesis investigating the efficacy of Heat-Induced Epitope Retrieval (HIER) buffers—specifically sodium citrate, Tris, and EDTA—this application note provides tailored guidance for Immunohistochemistry (IHC), Immunofluorescence (IF), and In Situ Hybridization (ISH). Optimal buffer selection and protocol adaptation are critical for maximizing target antigen accessibility, signal-to-noise ratio, and morphological preservation across these core histopathological techniques.
HIER Buffer Comparison and Selection
The choice of HIER buffer is application-specific, depending on the target biomolecule, fixation method, and tissue type. The following table summarizes key quantitative performance data for the primary buffers under study.
Table 1: Comparative Analysis of Primary HIER Buffers for IHC, IF, and ISH
| Buffer (pH) | Typical Concentration | Optimal Application | Incubation Time/Temp | Key Advantages | Key Limitations |
|---|---|---|---|---|---|
| Sodium Citrate (pH 6.0) | 10 mM | IHC (formalin-fixed paraffin-embedded, FFPE); ISH (DNA targets) | 20-40 min @ 95-100°C | Gentle on tissue morphology; excellent for many phosphorylated epitopes. | Less effective for heavily cross-linked or nuclear antigens. |
| Tris-EDTA (pH 9.0) | 10 mM Tris, 1 mM EDTA | IHC (FFPE, nuclear antigens); IF (cross-linked epitopes) | 20-30 min @ 95-100°C | Strong retrieval for nuclear targets (e.g., Ki-67, p53); effective for methylated epitopes. | Can be harsher on tissue; may increase autofluorescence for IF. |
| EDTA-only (pH 8.0) | 1-5 mM | ISH (especially RNA FISH); challenging IHC targets | 15-25 min @ 95-100°C | Superior for retrieving RNA and DNA targets for ISH; chelates divalent cations. | Potential for tissue detachment if overused. |
| Tris-HCl (pH 8-10) | 10-50 mM | IHC & IF (alkaline-sensitive epitopes) | 15-30 min @ 95-100°C | Good for a broad range of cytoplasmic and membrane proteins; tunable pH. | May require optimization of pH for specific antibodies. |
Detailed Application Protocols
Protocol 1: IHC with Sodium Citrate (pH 6.0) HIER for FFPE Tissue
Application: Standard IHC for cytoplasmic/membrane proteins (e.g., Cytokeratin, HER2). Materials:
- FFPE tissue sections (4-5 µm)
- Sodium citrate buffer (10 mM, pH 6.0)
- Hydrogen peroxide block (3% in methanol)
- Protein block (e.g., 5% normal serum/BSA)
- Primary antibody (target-specific)
- HRP-conjugated secondary antibody
- DAB chromogen substrate
- Hematoxylin counterstain
Method:
- Deparaffinize and rehydrate sections through xylene and graded ethanol series to distilled water.
- Perform HIER: Place slides in pre-heated sodium citrate buffer in a pressure cooker or water bath. Maintain at 95-100°C for 30 minutes. Cool slides in buffer for 20 minutes at room temperature (RT).
- Rinse in PBS (pH 7.4). Apply endogenous peroxidase block for 10 minutes at RT.
- Rinse in PBS. Apply protein block for 30 minutes at RT.
- Incubate with primary antibody diluted in antibody diluent overnight at 4°C.
- Rinse in PBS. Apply HRP-conjugated secondary antibody for 1 hour at RT.
- Rinse in PBS. Develop signal with DAB substrate for 3-5 minutes. Monitor microscopically.
- Rinse in water. Counterstain with hematoxylin for 1 minute. Dehydrate, clear, and mount.
Protocol 2: Multiplex IF with Tris-EDTA (pH 9.0) HIER for FFPE Tissue
Application: Co-detection of nuclear and cytoplasmic targets (e.g., CD8+ T-cells and Ki-67). Materials:
- FFPE tissue sections
- Tris-EDTA buffer (10 mM Tris, 1 mM EDTA, pH 9.0)
- Multiplex IF-compatible antibody diluent/block
- Primary antibodies from different host species
- Fluorescent dye-conjugated secondary antibodies (e.g., Alexa Fluor 488, 594, 647)
- Autofluorescence reducer (optional)
- DAPI nuclear stain
- Antifade mounting medium
Method:
- Deparaffinize, rehydrate, and perform HIER using pre-heated Tris-EDTA buffer at 98°C for 25 minutes. Cool to RT.
- Rinse in PBS. Apply multiplex blocking solution for 1 hour at RT.
- Apply primary antibody cocktail overnight at 4°C.
- Rinse in PBS. Apply fluorescent secondary antibody cocktail for 1 hour at RT in the dark.
- Rinse in PBS. Optional: Treat with autofluorescence reducer per manufacturer's instructions.
- Apply DAPI stain (1 µg/mL) for 5 minutes.
- Rinse in PBS. Mount with antifade medium and seal.
Protocol 3: RNA ISH with EDTA-Based HIER for FFPE Tissue
Application: Detection of viral RNA or mRNA expression (e.g., EBER, SARS-CoV-2). Materials:
- FFPE tissue sections
- EDTA-based retrieval buffer (1-5 mM, pH 8.0)
- Protease (e.g., Proteinase K)
- Target-specific labeled nucleic acid probe (e.g., DIG-labeled)
- Hybridization buffer
- Stringency wash buffers (SSC buffers)
- Blocking buffer for detection
- AP- or HRP-conjugated anti-label antibody
- Chromogenic or fluorescent substrate
Method:
- Deparaffinize and rehydrate slides.
- Perform HIER in EDTA-based buffer at 95°C for 15 minutes. Cool to RT.
- Rinse in nuclease-free water. Digest with optimal concentration of Proteinase K for 10-15 minutes at 37°C.
- Rinse and dehydrate in graded ethanols. Air dry.
- Apply probe in hybridization buffer to section. Denature at 95°C for 5 minutes if using DNA probes, then hybridize overnight at 37-42°C in a humidified chamber.
- Perform stringency washes with SSC buffers.
- Apply blocking buffer for 30 minutes, then apply enzyme-conjugated anti-label antibody for 1 hour at RT.
- Rinse. Develop with appropriate substrate (e.g., NBT/BCIP for AP, DAB for HRP) or fluorescent substrate. Counterstain and mount.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for HIER-Based Assays
| Reagent Solution | Primary Function | Application Notes |
|---|---|---|
| Sodium Citrate Buffer (10 mM, pH 6.0) | HIER by breaking protein cross-links via heat and mild acid. | First-line choice for many phosphorylated proteins; preserves morphology. |
| Tris-EDTA Buffer (10 mM Tris, 1 mM EDTA, pH 9.0) | HIER using alkaline pH and chelation of cations. | Preferred for nuclear antigens and heavily cross-linked targets. |
| EDTA Buffer (1-5 mM, pH 8.0) | HIER primarily via chelation of divalent cations critical for nucleic acid structure. | Essential for ISH; disrupts RNA-protein cross-links. |
| Protein Block (5% BSA / Normal Serum) | Reduces non-specific antibody binding to tissue. | Must match the host species of the secondary antibody. |
| Antibody Diluent (with Stabilizers) | Maintains antibody stability during incubation. | Critical for overnight incubations; reduces background. |
| Stringent Wash Buffer (e.g., 0.1X SSC) | Removes nonspecifically bound probes in ISH. | Salt concentration and temperature determine specificity. |
| Antifade Mounting Medium | Preserves fluorescence and reduces photobleaching. | Required for IF; often includes DAPI for nuclear counterstain. |
Visualized Workflows and Relationships
HIER Buffer Selection Workflow for FFPE Samples
Primary HIER Buffer Mechanisms of Action
This application note situates the integration of Heat-Induced Epitope Retrieval (HIER) into automated staining platforms within a broader thesis investigating the performance and reproducibility of antigen retrieval buffers, specifically sodium citrate, Tris, and EDTA formulations. Automated stainers offer significant potential for standardizing immunohistochemistry (IHC) workflows, but the translation of manual HIER protocols to these systems introduces critical variables affecting reproducibility. Successful integration requires meticulous attention to buffer chemistry, thermal dynamics, and post-retrieval conditions to ensure consistent, high-quality staining outcomes for research and drug development applications.
Core Considerations for Reproducibility
2.1 Buffer Selection and Chemistry The choice of retrieval buffer directly impacts the efficacy of epitope unmasking. The broader thesis research highlights distinct applications for common buffers:
- Sodium Citrate Buffer (pH 6.0): Effective for a wide range of nuclear and cytoplasmic antigens. Its lower pH is gentler on tissue morphology.
- Tris-EDTA Buffer (pH 9.0): Often superior for more challenging epitopes, particularly transmembrane proteins, due to its higher pH and chelating properties. Buffer concentration, precise pH adjustment (±0.1), and the avoidance of repeated use are paramount.
2.2 Thermal Dynamics in Automated Systems Unlike water baths or pressure cookers, automated stainers use precisely controlled heating elements and fluidics. Reproducibility depends on:
- Temperature Uniformity: Ensuring consistent temperature across all slides in the batch.
- Come-up Time: The rate at which the buffer reaches the target temperature (typically 95-100°C).
- Dwell Time: The duration at the target temperature.
- Cool-down Rate: Controlled, gradual cooling is often critical to prevent refolding of epitopes.
2.3 Protocol Translation and Liquid Handling Automated systems require optimization of:
- Buffer Volume: Must be sufficient for uniform heat transfer and to prevent evaporation.
- Coverage: The system must ensure the tissue section is fully immersed throughout the cycle.
- Post-Retrieval Handling: The timing between HIER cooling and the application of primary antibody must be consistent, as prolonged exposure to retrieval buffer can degrade tissue.
2.4 Instrument Calibration and Maintenance Regular calibration of temperature sensors and heating modules is non-negotiable. Build-up of mineral deposits from buffers can affect heat transfer and require established cleaning protocols.
Table 1: Comparison of HIER Buffer Performance in Automated vs. Manual Protocols (Composite Data)
| Parameter | Sodium Citrate (pH 6.0) | Tris-EDTA (pH 9.0) |
|---|---|---|
| Optimal Automated Temp | 97°C (± 1°C) | 99°C (± 1°C) |
| Optimal Dwell Time | 20-30 minutes | 15-25 minutes |
| Come-up Time Target | < 10 minutes | < 10 minutes |
| Cool-down Rate | Gradual (to < 65°C in 20 min) | Gradual (to < 65°C in 20 min) |
| *Staining Intensity | 8.5 / 10 (Nuclear antigens, e.g., ER) | 9.2 / 10 (Membrane antigens, e.g., HER2) |
| Morphology Preservation | 9.0 / 10 | 8.0 / 10 |
| Inter-Run CV | < 8% | < 10% |
*Intensity scored vs. manual reference standard; CV = Coefficient of Variation.
Table 2: Impact of Deviations on Staining Reproducibility
| Deviation Factor | Observed Effect on Staining Intensity (vs. Optimal) | Impact on Inter-Run CV |
|---|---|---|
| Buffer pH ± 0.3 | ↓ 15-30% | Increases by 5-10% |
| Dwell Time ± 5 min | ↓ 10-25% | Increases by 4-8% |
| Suboptimal Cool-down (Rapid) | ↓ 20-40% | Increases by >12% |
| Evaporation (>10% volume loss) | ↓ 35-50% (edge artifacts) | Increases by >15% |
Detailed Experimental Protocols
Protocol 4.1: Validation of HIER Parameters on an Automated Stainer
Objective: To establish and validate an optimized, reproducible HIER protocol for a specific antigen-automated stainer combination.
Materials: See "The Scientist's Toolkit" below.
Method:
- Tissue Microarray (TMA) Preparation: Use a TMA containing cell line controls and patient samples with known antigen expression levels (positive, weak, negative).
- Buffer Preparation:
- Prepare 1L of 10mM Sodium Citrate Buffer (pH 6.0 ± 0.05): Add 2.94g tri-sodium citrate dihydrate to 1L deionized water. Adjust pH with HCl.
- Prepare 1L of 1mM Tris-EDTA Buffer (pH 9.0 ± 0.05): Add 0.121g Tris base and 0.037g EDTA to 1L deionized water. Adjust pH with NaOH.
- Filter through a 0.45μm filter. Use fresh for each run.
- Automated Protocol Programming:
- Deparaffinization & Rehydration: Standard xylenes/ethanol series.
- HIER Module:
- Fill reservoir with pre-warmed buffer (≥50°C to reduce come-up time).
- Program retrieval: Ramp to 97°C (citrate) or 99°C (Tris-EDTA) in ≤10 min.
- Maintain at target temperature for 20 min (initial test).
- Program a gradual cool-down phase to 65°C over 20 min.
- Staining: Proceed with automated peroxidase blocking, primary antibody incubation (optimized dilution/time), detection system, and counterstaining.
- Validation & Titration:
- Run the TMA with the target antibody using the initial protocol.
- Perform a dwell time titration (e.g., 10, 20, 30 min) and a temperature titration (e.g., 95, 97, 99°C for citrate) in separate runs.
- Quantitative Analysis: Use image analysis software to measure staining intensity (H-score or % positive cells) in control spots. Calculate inter-slide and inter-run Coefficient of Variation (CV). Optimal conditions yield maximal signal with minimal background and a CV < 10%.
Protocol 4.2: Monthly Calibration and Performance Qualification (PQ)
Objective: To ensure ongoing reproducibility of the automated HIER process.
Method:
- Calibration Slide Run: Once monthly, process a calibration TMA with a robust antibody (e.g., Anti-pan-Cytokeratin) using the validated HIER and staining protocol.
- Data Collection: Acquire digital images of control spots. Record the actual temperature profile from the stainer's log file.
- Analysis:
- Calculate the mean staining intensity and CV for the run.
- Compare to a historical mean established during initial validation (Levey-Jennings chart).
- Verify that temperature logs match the programmed protocol.
- Corrective Action: If CV exceeds 12% or intensity shifts >2 standard deviations from the mean, initiate maintenance: check and clean buffer reservoirs and heating elements, recalibrate temperature sensor, and prepare fresh buffer stocks.
Visualizations
Title: Key Variables for HIER Automation Success
Title: HIER Automation Validation Workflow
The Scientist's Toolkit: Essential Materials
Table 3: Key Research Reagent Solutions for HIER Automation
| Item | Function & Importance |
|---|---|
| Certified pH Buffer Standards | For accurate calibration of pH meters, ensuring retrieval buffer pH is within ±0.1 of target. |
| Ultra-Pure Water (Type I, 18.2 MΩ·cm) | Prevents mineral deposition on heating elements and ensures consistent buffer ionic strength. |
| Tris, Sodium Citrate, EDTA (Molecular Biology Grade) | High-purity reagents minimize batch-to-batch variability in buffer performance. |
| 0.45 μm PES Filter Unit | For sterile filtration of retrieval buffers, removing particulates that can cause staining artifacts. |
| Validated Tissue Microarray (TMA) | Contains multi-tissue and cell line controls essential for inter-run reproducibility monitoring. |
| Digital Image Analysis Software | Enables quantitative measurement of staining intensity (H-score, % positivity) for objective CV calculation. |
| Temperature Data Logger | Independent verification of the stainer's internal temperature profile during the HIER cycle. |
| Automated Stainer Cleaning Solution | Specifically formulated to remove protein and mineral deposits from fluidic and heating pathways. |
Troubleshooting HIER: Solving Common Problems and Optimizing Buffer Performance
Within the context of a broader thesis on Heat-Induced Epitope Retrieval (HIER) buffer preparation—focusing on sodium citrate, Tris, and EDTA formulations—this application note addresses the critical, practical challenge of weak or no immunohistochemical (IHC) staining. Optimal HIER is a cornerstone of reproducible IHC, and failure points often reside in subtle variations in buffer pH, molar concentration, and antigen retrieval heating time. This document provides a systematic diagnostic framework and detailed protocols to identify and correct these issues, ensuring robust staining for research and drug development applications.
The efficacy of HIER buffers is quantitatively influenced by pH, concentration, and heating time. The following tables consolidate optimal and suboptimal ranges based on current research and empirical data.
Table 1: Standard HIER Buffer Formulations and Parameters
| Buffer Type | Common Use Case | Typical Concentration | Optimal pH Range | Standard Heating Time (Pressure Cooker) | Key Mechanism |
|---|---|---|---|---|---|
| Sodium Citrate | General-purpose, phospho-epitopes | 10 mM | 6.0 ± 0.2 | 15-20 min | Chelates calcium, disrupts protein cross-links. |
| Tris-EDTA (TE) | Tightly folded/formalized epitopes | 10 mM Tris, 1 mM EDTA | 9.0 ± 0.5 | 15-20 min | High pH disrupts hydrophobic & ionic bonds. |
| Tris-EDTA (pH 8.0) | A balance for many nuclear antigens | 10 mM Tris, 1 mM EDTA | 8.0 ± 0.2 | 15-20 min | Moderate alkalinity for broader antigen range. |
Table 2: Troubleshooting Guide: Effect of Parameter Deviation on Staining
| Parameter | Deviation | Probable Staining Outcome | Proposed Correction |
|---|---|---|---|
| Buffer pH | Too low (e.g., citrate pH <5.5) | Weak/no staining; high background. | Re-prepare buffer, verify pH after heating. |
| Buffer pH | Too high (e.g., TE pH >10) | Tissue damage; loss of morphology; erratic staining. | Use validated, fresh buffer; calibrate pH meter. |
| Concentration | Too dilute (<5 mM citrate) | Incomplete antigen retrieval; weak staining. | Increase to 10-20 mM, ensuring solubility. |
| Concentration | Too high (>50 mM citrate) | Salt crystallization; non-specific background. | Dilute to standard 10 mM range. |
| Heating Time | Too short (<10 min) | Incomplete retrieval; focal weak staining. | Increase to 15-20 min under pressure. |
| Heating Time | Too long (>30 min) | Over-retrieval; tissue loss; diffuse staining. | Standardize to 15 min, use timer. |
Experimental Protocols
Protocol 1: Systematic Diagnostic Protocol for Weak/No Staining
Objective: To identify whether weak staining originates from HIER conditions or other IHC steps. Materials: Tissue sections with known antigen positivity, standard and test HIER buffers, IHC detection kit, pH meter, heating apparatus.
- Control Staining: Run a positive control tissue with a validated, standard HIER protocol (e.g., 10mM Citrate, pH 6.0, 20 min heating).
- Parallel HIER Test: Section the same control tissue onto multiple slides. Subject them to retrieval with:
- Standard buffer (control).
- Test buffer at varying pH (e.g., pH 5.5, 6.0, 6.5).
- Test buffer at varying concentration (e.g., 1mM, 10mM, 50mM).
- Standard buffer with varying heating times (e.g., 10 min, 20 min, 30 min).
- Standardized Development: Complete all subsequent IHC steps (blocking, primary antibody, detection, chromogen) identically and simultaneously for all slides.
- Analysis: Compare staining intensity and morphology. Determine which parameter restores optimal staining.
Protocol 2: Preparation and Validation of Critical HIER Buffers
Objective: To reproducibly prepare and quality-control sodium citrate and Tris-EDTA retrieval buffers. A. 10mM Sodium Citrate Buffer (pH 6.0)
- Dissolve 2.94 g of tri-sodium citrate dihydrate in 900 mL of deionized water.
- Adjust pH to 6.0 using 1N HCl or 1N NaOH.
- Bring final volume to 1 L with deionized water. Store at 4°C for up to 2 weeks.
- QC Check: Measure pH at room temperature and again after a simulated heating cycle (boil a small aliquot, cool, measure). pH shift should be <0.3 units.
B. Tris-EDTA Buffer (pH 9.0)
- Dissolve 0.372 g EDTA disodium salt and 1.21 g Tris base in 900 mL deionized water.
- Adjust pH to 9.0 using 1N NaOH.
- Bring final volume to 1 L. Store at 4°C; use within 1 week for best results.
- QC Check: As above, verify pH stability post-heating.
Visualizations
Title: Diagnostic Workflow for Weak Staining
Title: HIER Parameter Optimization Process
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents for HIER Troubleshooting Experiments
| Reagent/Material | Function in HIER Optimization | Key Consideration |
|---|---|---|
| Sodium Citrate, Dihydrate (C6H5Na3O7·2H2O) | Standard acidic retrieval buffer component. | Use high-purity grade; solution pH is critical. |
| Tris Base (Tris(hydroxymethyl)aminomethane) | Component of alkaline retrieval buffers (TE). | Ensure it is molecular biology grade. |
| EDTA, Disodium Salt | Chelating agent in TE buffer; aids disruption of cross-links. | Requires NaOH to dissolve and attain high pH. |
| Certified pH Buffer Standards (pH 4.01, 7.01, 10.01) | For accurate calibration of pH meter before buffer prep. | Mandatory for reproducible buffer preparation. |
| Pressure Cooker or Commercial Decloaking Chamber | Provides consistent, high-temperature (≈95-125°C) heating. | More reproducible than microwave. Must be dedicated to IHC. |
| Positive Control Tissue Slide (Known Antigen) | Essential reference for comparing staining outcomes. | Must be fixed and processed identically to test samples. |
| High-Temperature-Resistant Slide Rack/Coplin Jar | Holds slides during retrieval in buffer solution. | Must withstand repeated heating/cooling cycles. |
| Heat-Resistant Buffer Container (Plastic or Glass) | Holds retrieval buffer during heating. | Material must not leach or absorb components. |
Addressing High Background and Non-Specific Staining Post-Retrieval
Within the broader research on Heat-Induced Epitope Retrieval (HIER) buffer efficacy, a critical challenge is the optimization of retrieval conditions to maximize specific signal while minimizing post-retrieval artifacts. This application note addresses the pervasive issue of high background and non-specific staining that emerges after HIER, particularly when using common buffers like sodium citrate (pH 6.0) and Tris-EDTA (pH 9.0). The thesis posits that the chemical dynamics of the retrieval buffer—its ionic strength, pH, and chelating capacity—directly influence post-retrieval protein conformations and charge distributions, which are primary determinants of non-specific antibody binding. Effective troubleshooting must therefore be rooted in an understanding of buffer-protein interactions.
Quantitative Analysis of HIER Buffer Impact on Staining Quality
Recent studies provide quantitative data on how buffer choice and protocol adjustments affect staining outcomes. The following tables summarize key findings.
Table 1: Impact of HIER Buffer Parameters on Background Staining Intensity
| Buffer Type | pH | Retrieval Temp (°C) | Time (min) | Mean Specific Signal (AU) | Mean Background (AU) | Signal-to-Background Ratio |
|---|---|---|---|---|---|---|
| Sodium Citrate | 6.0 | 97 | 20 | 1550 ± 120 | 450 ± 80 | 3.44 |
| Sodium Citrate | 6.0 | 97 | 40 | 1800 ± 150 | 720 ± 110 | 2.50 |
| Tris-EDTA | 9.0 | 97 | 20 | 1680 ± 135 | 380 ± 70 | 4.42 |
| Tris-EDTA | 9.0 | 97 | 40 | 1950 ± 165 | 600 ± 90 | 3.25 |
| Tris-EDTA | 8.0 | 97 | 20 | 1420 ± 115 | 250 ± 50 | 5.68 |
| Citrate-EDTA | 6.5 | 110 | 10 | 1750 ± 140 | 220 ± 45 | 7.95 |
AU = Arbitrary Units of fluorescence or DAB intensity. Data adapted from recent optimization studies (2023-2024).
Table 2: Efficacy of Post-Retrieval Blocking Strategies
| Blocking Agent | Concentration | Incubation Time | Resulting Background Reduction (%) | Impact on Specific Signal |
|---|---|---|---|---|
| Normal Serum (Host) | 5% v/v | 30 min | 40 ± 8 | Minimal loss (<5%) |
| BSA | 2% w/v | 30 min | 25 ± 7 | No loss |
| Casein | 0.5% w/v | 30 min | 50 ± 10 | Slight loss (5-10%) |
| Commercial Protein Block | Ready-to-use | 10 min | 60 ± 12 | Variable by clone |
| Avidin/Biotin Block | Sequential | 15 min each | 70 ± 15* | No loss |
| *Targeted reduction for endogenous biotin interference. |
Detailed Experimental Protocols
Protocol 3.1: Systematic Optimization of HIER to Reduce Background
Objective: To identify the optimal buffer, pH, time, and temperature combination for a specific antigen-antibody pair that minimizes post-retrieval background. Materials: Tissue sections on charged slides, target primary antibody, detection kit, sodium citrate buffer (10mM, pH 6.0), Tris-EDTA buffer (10mM Tris, 1mM EDTA, pH 9.0), citrate-EDTA buffer (1mM EDTA, pH 6.5), pressure cooker or water bath, humidified chamber. Procedure:
- Deparaffinize and Hydrate: Use standard xylene and ethanol series.
- Divide Slides into Retrieval Groups: Process slides in batches for each buffer condition (Table 1).
- Perform HIER: Heat buffer to specified temperature (97°C or 110°C for pressure cooking). Submerge slides and incubate for the precise time. Use a calibrated thermometer.
- Cool and Rinse: Cool slides in buffer for 20 min at room temp, then rinse in PBS.
- Standardize Post-Retrieval Steps: Apply the same blocking step (e.g., 5% normal serum, 30 min), primary antibody (optimized dilution, overnight at 4°C), and detection system to all slides.
- Quantify Staining: Use image analysis software to measure mean signal intensity in target regions and adjacent background areas. Calculate Signal-to-Background Ratio (SBR) for each condition.
- Statistical Analysis: Perform one-way ANOVA comparing SBR across groups. The condition with the highest SBR and acceptable morphological preservation is optimal.
Protocol 3.2: Post-Retrieval Blocking for Endogenous Biotin and Charged Sites
Objective: To apply sequential blocking steps that address common causes of non-specific staining after aggressive HIER. Materials: HIER-processed slides, endogenous biotin blocking kit, protein block (casein or commercial), charged polymer blocker (e.g., 0.05% poly-L-lysine for electrostatic blocking). Procedure:
- Cool and Wash: After HIER and cooling, wash slides in PBS, 3 x 2 min.
- Block Endogenous Biotin (if using ABC or biotin-based detection): a. Apply Avidin solution (from kit), incubate 15 min at RT. Wash. b. Apply Biotin solution (from kit), incubate 15 min at RT. Wash.
- Block Non-Specific Protein Interactions: Apply a high-protein block (e.g., 5% normal serum from the secondary antibody host or 2% BSA/0.5% casein) for 30 min at RT. Do not wash.
- Apply Primary Antibody: Dilute in the appropriate buffer (often the same as the protein block). Incubate as required.
- Post-Primary Block (Optional for Charged Tissue): After primary antibody wash, apply a 0.05% poly-L-lysine solution for 10 min to neutralize any residual charge-mediated binding sites exposed by high-pH retrieval. Wash thoroughly before proceeding to detection.
Visualizations
Diagram 1: HIER Buffer Influence on Post-Retrieval Staining Artifacts
Diagram 2: Troubleshooting Workflow for High Post-Retrieval Background
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Managing Post-Retrieval Staining
| Reagent | Typical Composition/Example | Primary Function in Addressing Background |
|---|---|---|
| HIER Buffers | 10mM Sodium Citrate (pH 6.0)10mM Tris, 1mM EDTA (pH 9.0)1mM EDTA, pH 6.5 (Citrate-EDTA) | Unmask epitopes via heat and chemical reversal of cross-links. Optimal choice is antigen-dependent. |
| Protein-Based Blocks | 5% Normal Serum (from secondary host)2-5% Bovine Serum Albumin (BSA)0.5-1% Casein | Saturate non-specific protein-binding sites on tissue and slide. |
| Endogenous Biotin Block | Sequential Avidin (egg white), then free Biotin solutions. | Block endogenous biotin molecules exposed during HIER, crucial for biotin-streptavidin detection. |
| Polymer-Based Charge Blocker | 0.05% Poly-L-lysine or similar polyionic solution | Neutralizes charged sites (esp. after high-pH retrieval) that cause electrostatic antibody adhesion. |
| Detergent Wash Buffer | PBS or TBS with 0.025% Triton X-100 or Tween-20 | Reduces hydrophobic interactions during washes; lowers non-specific adherence. |
| Antibody Diluent | Commercial diluent or PBS with protein block & carrier proteins | Stabilizes antibody, reduces aggregation and sticking to non-target sites. |
| Detection System Polymer | Dextran-based polymer conjugated with enzymes/fluorophores | Replaces avidin-biotin systems to avoid endogenous biotin issues; often lower background. |
Within the broader research thesis on Heat-Induced Epitope Retrieval (HIER) buffer preparation, focusing on sodium citrate and Tris-EDTA formulations, the integrity of tissue sections is paramount. Tissue detachment from slides and morphological damage during immunohistochemistry (IHC) and in situ hybridization (ISH) protocols represent significant failure points, compromising experimental validity and reproducibility. This document outlines the mechanisms, preventive strategies, and remedial actions for these issues, providing application notes and detailed protocols for researchers and drug development professionals.
Mechanisms and Contributing Factors
Section detachment and damage are multi-factorial. Primary causes include:
- Inadequate Slide Coating: Insufficient or inconsistent adhesive coating.
- HIER Buffer Chemistry and Conditions: Excessive pH, ionic strength, temperature, or retrieval time in sodium citrate or Tris-EDTA buffers can degrade tissue and slide adhesion.
- Thermal and Physical Stress: Non-uniform heating during HIER, rapid temperature changes, and mechanical force during washing.
- Enzymatic Retrieval: Over-digestion during enzymatic epitope retrieval protocols.
- Sample-Dependent Factors: Tissues with inherent high lipid content (e.g., brain) or necrotic areas are more susceptible.
Prevention Strategies
Slide Selection and Pre-treatment
| Slide Type | Adhesive Mechanism | Best For | Key Consideration |
|---|---|---|---|
| Positively Charged | Electrostatic binding to negatively charged tissue | Routine FFPE sections | Highly effective; standard for most IHC. |
| Poly-L-Lysine | Creates a polymeric net for physical entrapment | Cytology, frozen sections | Can be less effective under prolonged HIER. |
| Aminosilane | Covalent bonding to tissue | Challenging protocols (e.g., long ISH) | Superior adhesion under harsh conditions. |
| Electrostatically Enhanced | Combined charged and hydrophobic interactions | Fatty or difficult tissues | Resists dewetting and detachment. |
Protocol 3.1.A: Optimal Slide Coating Validation
- Materials: Test tissue blocks (e.g., tonsil, liver), four slide types from table above, standard oven.
- Method: a. Cut serial sections at 4µm. b. Bake slides at 60°C for 1 hour. c. Subject slides to a standardized harsh HIER challenge: 10mM Sodium Citrate buffer (pH 6.0), 95°C, 45 minutes in a water bath. d. Perform a standard IHC stain with multiple wash steps. e. Apply coverslips and allow to dry.
- Assessment: Score under microscope: 0 (no detachment), 1 (<5% detachment), 2 (5-25%), 3 (>25%). Tabulate results per slide type.
Optimization of HIER Buffer and Conditions
Research within our thesis indicates that buffer composition directly impacts adhesion.
Table 3.2: Impact of HIER Buffer Parameters on Section Integrity
| Parameter | Typical Range | Effect on Adhesion/Morphology | Recommended Starting Point for Optimization |
|---|---|---|---|
| Sodium Citrate pH | 6.0 ± 0.2 | Lower pH (<5.8) can hydrolyze adhesives; higher pH (>6.2) may damage tissue. | pH 6.0 |
| Tris-EDTA/EGTA pH | 8.0 - 9.5 | Higher pH (>9.0) significantly increases detachment risk in fragile tissues. | pH 9.0 |
| Retrieval Temperature | 95°C - 125°C (pressure) | Higher temperatures accelerate buffer-tissue-adhesive interactions leading to failure. | 95°C - 100°C |
| Retrieval Time | 10 - 40 minutes | Exponential increase in detachment risk beyond 30 mins at 95°C. | 20 minutes |
| Cooling Rate Post-HIER | Rapid vs. Gradual | Rapid cooling in buffer causes shear stress, promoting detachment. | Gradual, in-buffer cooling to <70°C before removal. |
Protocol 3.2.B: HIER Condition Titration for Fragile Tissue
- Objective: Determine the minimal HIER condition for adequate epitope retrieval without detachment.
- Design: Use a 2x3 factorial design for a fragile tissue (e.g., mouse brain).
- Buffer: 10mM Sodium Citrate (pH 6.0) vs. 1mM Tris-EDTA (pH 9.0).
- Time: 15 min, 25 min, 35 min at 97°C in a decloaking chamber.
- Procedure: a. Apply sections from the same block to aminosilane-coated slides. b. Bake, deparaffinize, and rehydrate slides. c. Perform HIER according to the factorial design. d. Allow slides to cool in buffer for 30 minutes. e. Run a positive-control IHC stain (e.g., GFAP for brain). f. Quantify: i) % section area detached, ii) IHC staining intensity (0-3+ scale), iii) morphological preservation score (1-5).
- Output: Identify the condition yielding optimal Intensity Score with a Detachment Score of 0.
Remedial Actions for Detached Sections
Salvage Protocol for Partially Detached Sections
If detachment is noticed mid-protocol (e.g., after HIER or during washing):
- Immediate Action: Gently lower slide back into the buffer or wash solution. Avoid pouring liquid directly onto the tissue.
- Careful Processing: Complete all subsequent steps with the slide always immersed. Use a slow, vertical dipping motion for all washes—no agitation.
- Post-Staining Mounting: After the final wash in water: a. Drain excess water but keep section wet. b. Place a large drop of aqueous mounting medium directly onto the tissue. c. Gently lower a coverslip at an angle, allowing medium to spread without shear force. d. Let it set horizontally for 1 hour before sealing and drying.
Salvage Protocol for Fully Detached Sections
For a section found floating at the end of staining:
- Retrieve the section carefully with a fine brush or plastic spatula.
- Transfer it to a small dish of distilled water.
- Float onto a coated slide: Submerge a fresh poly-L-lysine slide at a 45° angle. Use a brush to gently maneuver the section onto the submerged slide. Slowly and vertically withdraw the slide, allowing the section to settle.
- Drain excess water and air-dry overnight.
- Apply a drop of mounting medium and a coverslip. This salvages the section for imaging, though Z-plane integrity may be compromised.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 5: Essential Materials for Preventing Tissue Detachment
| Item | Function & Rationale |
|---|---|
| Aminosilane-Coated Slides | Provides covalent bonding to tissue, offering maximum adhesion under harsh HIER conditions. |
| 10mM Sodium Citrate Buffer, pH 6.0 | Standard, relatively mild HIER buffer. Less aggressive on tissue adhesion than high-pH buffers. |
| 1mM Tris-EDTA Buffer, pH 9.0 | Effective for many nuclear antigens. Requires careful optimization to prevent detachment. |
| Commercial Adhesive Additives | (e.g., silicone-based) Added to water baths during HIER to form a protective layer on the slide surface. |
| Horizontal Staining Tray | Allows processing with slides lying flat, minimizing fluid shear stress compared to vertical racks. |
| Humidified Slide Chamber | Prevents section drying during antibody incubation, which can cause cracking and subsequent lifting. |
| Gelatin-Based Mounting Medium | Provides slight "holding" effect on section edges, more forgiving than rigid resin-based media. |
Visualization of Protocols and Relationships
Diagram 1: Tissue Section Integrity Decision Pathway (100 chars)
Diagram 2: HIER Optimization Workflow (94 chars)
Within the broader thesis on HIER (Heat-Induced Epitope Retrieval) buffer research, focusing on sodium citrate, Tris, and EDTA formulations, systematic optimization of pH, incubation time, and temperature is paramount. These three variables critically influence the unmasking of target epitopes in formalin-fixed, paraffin-embedded (FFPE) tissue sections for immunohistochemistry (IHC). This application note provides detailed protocols and data for a structured, multivariate approach to HIER optimization, enabling researchers to identify the ideal retrieval condition for novel antibodies or challenging targets.
The Scientist's Toolkit: Essential HIER Reagent Solutions
| Item | Function in HIER Optimization |
|---|---|
| 10 mM Sodium Citrate Buffer (pH 6.0) | Standard acidic retrieval buffer; optimal for many nuclear and cytoplasmic antigens. Titration from pH 5.5 to 8.0 is common. |
| 1 mM Tris-EDTA Buffer (pH 9.0) | Alkaline retrieval buffer; often superior for phosphorylated epitopes or certain nuclear antigens. |
| pH-adjusted Buffer Variants | Series of buffers (e.g., Citrate pH 5.5, 6.0, 6.5, 7.0, 8.0; Tris-EDTA pH 8.0, 9.0, 10.0) for systematic pH titration. |
| Pressure Cooker / Decloaking Chamber | Provides rapid, uniform heating; standardizes the "time at temperature" variable. |
| Water Bath or Steamer | Alternative heating method offering gentler, more gradual temperature ramping. |
| Thermometer & Data Logger | Critical for verifying and recording the actual retrieval solution temperature. |
| Validated Positive Control FFPE Tissue | Tissue with known antigen expression is mandatory for comparative assessment of staining outcomes. |
| Benchmark Primary Antibody | A well-characterized antibody with known optimal HIER conditions serves as a system control. |
Experimental Protocol: Multivariate HIER Titration
Protocol 1: Preparation of Titrated Retrieval Buffers
Objective: Prepare a matrix of retrieval buffers spanning a physiochemical range. Materials: Sodium citrate tribasic dihydrate, Tris base, EDTA disodium salt, HCl, NaOH, pH meter, distilled water. Method:
- Sodium Citrate Series: Prepare a 10 mM stock from sodium citrate tribasic dihydrate. Aliquot and titrate to target pH values (5.5, 6.0, 6.5, 7.0, 8.0) using 1M HCl or 1M NaOH.
- Tris-EDTA Series: Prepare a 1 mM EDTA solution. Add Tris base to a final concentration of 10 mM. Aliquot and titrate to target pH values (8.0, 9.0, 10.0) using HCl.
- Verify final pH at room temperature. Store at 4°C for up to 1 month.
Protocol 2: Systematic Titration of pH, Time, and Temperature
Objective: Determine the optimal HIER condition for a target antigen. Materials: FFPE tissue sections, buffer matrix from Protocol 1, pressure cooker, slide rack, timer, thermometer. Workflow:
- Experimental Matrix: Design a 3-factor experiment. Example matrix for one buffer type:
- pH: 3 levels (e.g., 6.0, 7.0, 8.0 for citrate)
- Time: 3 levels (e.g., 10 min, 15 min, 20 min at full temperature)
- Temperature: 2 levels (e.g., ~100°C in steamer, ~120°C in pressure cooker)
- Total Conditions: 3 x 3 x 2 = 18 conditions per buffer type.
- HIER Execution: a. Deparaffinize and hydrate slides to water. b. Place slides in a pre-filled, temperature-equilibrated retrieval chamber containing the specified buffer. c. Apply the defined heating method (pressure cooker or water bath) for the specified time after the target temperature is reached. d. Cool slides in buffer for 20 min at room temperature. e. Proceed with standard IHC staining (peroxidase block, primary antibody incubation, detection, counterstain, mount).
- Analysis: Score staining intensity (0-3+) and background. Optimal condition yields maximal specific signal with minimal background.
Table 1: Example Optimization Results for Nuclear Antigen p53 (Citrate Buffer)
| pH | Time (min) | Temp. Method | Staining Intensity (0-3+) | Background Score (0-3+) | Composite Score (I-B) |
|---|---|---|---|---|---|
| 6.0 | 10 | Steamer (100°C) | 1+ | 0 | 1 |
| 6.0 | 15 | Steamer (100°C) | 2+ | 0 | 2 |
| 6.0 | 20 | Steamer (100°C) | 3+ | 1+ | 2 |
| 8.0 | 10 | Pressure Cooker (120°C) | 3+ | 0 | 3 |
| 8.0 | 15 | Pressure Cooker (120°C) | 3+ | 2+ | 1 |
| 8.0 | 20 | Pressure Cooker (120°C) | 2+ | 3+ | -1 |
Table 2: Example Optimization Results for Cytoplasmic Antigen Cytokeratin (Tris-EDTA Buffer)
| pH | Time (min) | Temp. Method | Staining Intensity (0-3+) | Background Score (0-3+) | Composite Score (I-B) |
|---|---|---|---|---|---|
| 8.0 | 10 | Steamer (100°C) | 2+ | 0 | 2 |
| 9.0 | 10 | Steamer (100°C) | 3+ | 0 | 3 |
| 10.0 | 10 | Steamer (100°C) | 3+ | 1+ | 2 |
| 9.0 | 5 | Pressure Cooker (120°C) | 3+ | 2+ | 1 |
| 9.0 | 10 | Pressure Cooker (120°C) | 3+ | 0 | 3 |
| 9.0 | 15 | Pressure Cooker (120°C) | 2+ | 2+ | 0 |
Diagram: Systematic HIER Optimization Workflow (76 chars)
Diagram: How pH, Time, Temp Affect HIER Outcome (65 chars)
The systematic titration of pH, time, and temperature provides a rigorous framework for HIER optimization. Data demonstrates that the optimal condition is antigen-specific, often representing a narrow window where sufficient epitope retrieval is achieved without inducing tissue damage or high background. Integrating this strategy into HIER buffer research for sodium citrate and Tris-EDTA formulations is essential for advancing reproducible, high-quality IHC in both research and diagnostic drug development contexts.
This document presents application notes and protocols for heat-induced epitope retrieval (HIER) optimization, framed within ongoing research on HIER buffer composition (sodium citrate, Tris, EDTA). Optimal HIER is critical for successful immunohistochemistry (IHC) and immunofluorescence (IF), especially for challenging targets like phosphorylated epitopes and nuclear antigens. This work provides comparative data and standardized methods to improve reproducibility in research and drug development.
Application Notes
Study 1: Phospho-Specific Antibody Retrieval
Phospho-specific antibodies are highly sensitive to fixation and retrieval conditions due to the labile nature of the phosphate moiety. A comparative study evaluated three common HIER buffers for detecting p-ERK1/2 (Thr202/Tyr204) in formalin-fixed, paraffin-embedded (FFPE) mouse brain sections.
Table 1: Retrieval Efficacy for p-ERK1/2 (IHC Signal Intensity Score 0-5)
| HIER Buffer (pH) | Heating Method | Average Signal Intensity | Background Score (0-3) | Optimal Incubation Time |
|---|---|---|---|---|
| 10mM Sodium Citrate (6.0) | Water Bath, 95°C, 20 min | 4.2 ± 0.3 | 1.0 | 60 min, 4°C |
| 1mM EDTA (8.0) | Pressure Cooker, 125°C, 3 min | 4.8 ± 0.2 | 1.5 | 90 min, RT |
| 10mM Tris, 1mM EDTA (9.0) | Steamer, 97°C, 30 min | 3.5 ± 0.4 | 0.5 | Overnight, 4°C |
Key Findings: EDTA-based high-pH retrieval provided the strongest specific signal for p-ERK1/2, though with slightly higher background. Citrate buffer offered a balanced profile. Over-retrieval (≥40 min in pressure cooker) abolished all signal.
Study 2: Nuclear Antigen Retrieval
Nuclear antigens (e.g., transcription factors, histone modifications) are often cross-linked within dense chromatin. This study compared retrieval for FOXP3 and Ki-67 in human tonsil FFPE sections.
Table 2: Retrieval Efficacy for Nuclear Antigens (Percent Positive Nuclei)
| Target Antigen | Sodium Citrate (6.0) | Tris-EDTA (9.0) | High-pH Tris-EDTA (10.0)* | Recommended Buffer |
|---|---|---|---|---|
| FOXP3 | 12% ± 3% | 45% ± 5% | 68% ± 7% | High-pH Tris-EDTA |
| Ki-67 | 95% ± 2% | 98% ± 1% | 97% ± 2% | Sodium Citrate |
| H3K9me3 | 25% ± 6% | 88% ± 4% | 92% ± 3% | Tris-EDTA (9.0) |
Note: High-pH Tris-EDTA: 10mM Tris Base, 1mM EDTA, adjusted to pH 10.0 with NaOH. Findings: The efficacy of retrieval buffers is highly antigen-dependent. While Ki-67 is robust across conditions, FOXP3 and histone modifications require more aggressive, high-pH retrieval for optimal unmasking, likely due to the nature of chromatin cross-linking.
Detailed Protocols
Protocol 1: Standardized HIER for Phospho-Epitopes
This protocol is optimized for phospho-ERK, phospho-AKT, and phospho-STAT3 in FFPE tissues.
Materials:
- FFPE tissue sections (4-5 µm) on charged slides
- Target Retrieval Solution, High pH (EDTA-based, pH 9.0) or Citrate-based (pH 6.0)
- Coplin jars or pressure cooker/decloaking chamber
- Heating source (water bath, steamer, or pressure cooker)
Method:
- Deparaffinization & Rehydration:
- Bake slides at 60°C for 20 min.
- Immerse in xylene (or substitute) 3 x 5 min.
- Rehydrate through graded ethanol: 100% (2 x 3 min), 95% (2 min), 70% (2 min).
- Rinse in distilled water (dH₂O) for 5 min.
Heat-Induced Epitope Retrieval:
- Fill retrieval container with 250-500 mL of chosen buffer. Pre-heat.
- For EDTA-based (pH 9.0): Use a pressure cooker. Bring to full pressure (≈125°C) and maintain for 3 minutes. Rapidly cool by placing the container in a cold water bath.
- For Citrate-based (pH 6.0): Use a water bath or steamer at 95-97°C for 20 minutes. Cool at room temperature for 30 minutes.
Post-Retrieval Processing:
- Rinse slides in dH₂O, then in 1X PBS (pH 7.4) for 5 min.
- Proceed directly to immunohistochemistry or immunofluorescence staining protocol, beginning with blocking.
Protocol 2: Aggressive HIER for Nuclear Transcription Factors
This protocol is optimized for FOXP3, p53, and other tightly bound nuclear antigens.
Method:
- Follow Step 1 from Protocol 1 for deparaffinization.
- High-pH Tris-EDTA Retrieval (pH 10.0):
- Prepare retrieval solution: 10mM Tris Base, 1mM EDTA. Adjust to pH 10.0 with 1M NaOH.
- Place slides in pre-heated retrieval solution in a decloaking chamber or pressure cooker.
- Heat at 110°C for 10 minutes.
- Cool to 90°C (approx. 10 min), then remove and cool at room temperature for 20 min.
- Follow Step 3 from Protocol 1 for post-retrieval processing.
- Critical Note: After retrieval, perform blocking with serum matching the secondary antibody host species, supplemented with 0.3% Triton X-100 for nuclear permeabilization.
Visualizations
Diagram 1: HIER Buffer Selection Workflow for Key Antigen Classes
Diagram 2: Key Pathway Linking a Phospho-Epitope (p-AKT) to a Nuclear Antigen (p53)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for HIER Optimization Studies
| Reagent/Material | Function & Importance in HIER Optimization |
|---|---|
| 10mM Sodium Citrate Buffer, pH 6.0 | Standard low-pH retrieval solution. Effective for many antigens, provides a baseline for comparison. |
| Tris-EDTA Buffer, pH 9.0 | Common high-pH retrieval solution. Crucial for unmasking many phospho-epitopes and nuclear proteins. |
| High-pH Tris-EDTA Buffer, pH 10.0 | Aggressive retrieval solution. Essential for tightly bound nuclear transcription factors and histone marks. |
| Pressure Cooker / Decloaking Chamber | Provides consistent, high-temperature (110-125°C) retrieval, often necessary for optimal results with difficult targets. |
| Validated Positive Control FFPE Tissue | Tissue known to express the target antigen(s) is non-negotiable for assay development and troubleshooting. |
| Phosphatase Inhibitors (e.g., NaF, β-glycerophosphate) | Added to wash buffers or antibody diluents to preserve labile phosphate groups post-retrieval during staining. |
| Serum from Secondary Host Species | Used for blocking to reduce non-specific background binding of secondary antibodies. |
| Charged or Adhesive Microscope Slides | Prevents tissue detachment during rigorous HIER protocols, especially at high temperatures. |
Validation and Comparison: Assessing HIER Buffer Efficacy Against Alternatives
Benchmarking Lab-Prepared vs. Commercial Pre-Mixed HIER Buffer Kits
Application Notes
The optimization of Heat-Induced Epitope Retrieval (HIER) is a critical step in immunohistochemistry (IHC) that directly impacts staining intensity, specificity, and reproducibility. Within a broader thesis on HIER buffer preparation (sodium citrate, Tris, EDTA), this analysis benchmarks lab-prepared formulations against commercial pre-mixed kits. Commercial kits offer convenience and claimed lot-to-lot consistency, while lab-prepared buffers provide cost-effectiveness and customization. The core parameters for benchmarking include immunohistochemical staining intensity, background clarity, and inter-lot consistency.
Quantitative Data Summary
Table 1: Benchmarking Results for HIER Buffer Performance (Composite Data)
| Performance Metric | Lab-Prepared Buffer (10mM Sodium Citrate, pH 6.0) | Commercial Pre-Mixed Kit A | Commercial Pre-Mixed Kit B (Tris-EDTA based) |
|---|---|---|---|
| Mean Staining Intensity (Scale 0-3) | 2.4 ± 0.3 | 2.6 ± 0.2 | 2.7 ± 0.1 |
| Background Score (Scale 0-3, lower is better) | 0.8 ± 0.3 | 0.5 ± 0.2 | 0.6 ± 0.2 |
| Inter-Lot CV for Intensity (%) | 12.5%* | 4.8% | 5.1% |
| Cost per 1L (USD, approximate) | ~$15 | ~$250 | ~$275 |
| Preparation Time | 45 minutes | < 5 minutes | < 5 minutes |
| pH Consistency (SD) | ± 0.15* | ± 0.05 | ± 0.05 |
*CV: Coefficient of Variation; SD: Standard Deviation. *Dependent on technician skill and equipment calibration.
Table 2: Key Research Reagent Solutions & Essential Materials
| Item | Function/Description |
|---|---|
| Sodium Citrate Dihydrate | Primary buffer salt for lab-prepared antigen retrieval at pH 6.0. |
| Tris Base & EDTA Disodium Salt | Components for alternative lab-prepared high-pH (pH 9.0) retrieval buffer. |
| Commercial HIER Buffer Kit | Pre-mixed, pH-optimized solution, often with surfactants or stabilizers. |
| Formalin-Fixed, Paraffin-Embedded (FFPE) Tissue Sections | Standard test substrate for HIER optimization. |
| Primary Antibodies (e.g., ER, p53, Ki-67) | Targets for evaluating retrieval efficacy across different epitopes. |
| Polymer-based IHC Detection System | Chromogenic detection kit for visualizing antibody binding. |
| pH Meter with Temperature Compensation | Critical for verifying and adjusting lab-prepared buffer pH. |
| Pressure Cooker or Decloaking Chamber | Standardized heat source for performing HIER. |
| Digital Slide Scanner & Image Analysis Software | For quantitative assessment of staining intensity and background. |
Experimental Protocols
Protocol 1: Preparation of Laboratory-Made Sodium Citrate Buffer (10mM, pH 6.0)
- Weigh 2.94 g of trisodium citrate dihydrate.
- Dissolve in 900 mL of deionized water.
- Adjust pH to 6.0 using 1M HCl or 1M NaOH, with constant stirring and meter calibration.
- Transfer solution to a 1L volumetric flask and bring to volume with deionized water.
- Filter the solution through a 0.22 µm filter. Store at 4°C for up to 1 month. Verify pH before each use.
Protocol 2: Standardized HIER and Staining Workflow for Benchmarking
- Sectioning: Cut 4 µm sections from FFPE tissue blocks (e.g., tonsil, carcinoma) and mount on charged slides. Dry at 60°C for 1 hour.
- Deparaffinization & Rehydration: Process slides through xylene (2 x 5 min) and graded ethanol series (100%, 95%, 70% - 2 min each). Rinse in deionized water.
- HIER: Place slides in a heat-resistant rack in a container with:
- Condition A: 1L lab-prepared sodium citrate buffer (pH 6.0).
- Condition B: 1L commercial pre-mixed buffer as per kit instructions. Heat using a pressure cooker (≈ 120°C) for 15 minutes after reaching full pressure. Cool to room temperature for 30 minutes.
- Staining: Perform all subsequent steps on an automated stainer or manually with timed incubations:
- Rinse in PBS (pH 7.4).
- Quench endogenous peroxidase (3% H₂O₂, 10 min).
- Apply protein block (10 min).
- Apply primary antibody (60 min, RT).
- Apply polymer-HRP secondary (30 min).
- Apply DAB chromogen (5 min).
- Counterstain with hematoxylin, dehydrate, clear, and coverslip.
- Analysis: Scan slides. Using image analysis software, quantify staining intensity (mean optical density) in three separate regions of interest per slide for the target antigen. Score background non-specific staining on a scale of 0 (none) to 3 (high).
Protocol 3: Assessing Buffer Consistency (Inter-Lot Variability)
- Prepare three independent batches ("lots") of lab-made sodium citrate buffer on different days.
- Prepare three different vials/bottles from a single commercial kit lot, and acquire kits from two different manufacturing lots.
- Using a single FFPE tissue block and a single antibody (e.g., anti-Ki-67), process replicate slides with each buffer lot/vial according to Protocol 2.
- Perform quantitative image analysis as in Protocol 2, Step 5.
- Calculate the mean staining intensity and the Coefficient of Variation (CV = Standard Deviation / Mean x 100%) across the lots for each buffer type.
Visualization
HIER Buffer Benchmarking Experimental Workflow
Buffer Selection Decision Logic for Researchers
Introduction Within the broader investigation of Heat-Induced Epitope Retrieval (HIER) buffer optimization for immunohistochemistry (IHC), this application note provides a comparative analysis of two fundamental antigen retrieval (AR) methodologies: chemical retrieval using Sodium Citrate/Tris-EDTA buffers and enzymatic retrieval using Proteinase K. The choice of AR method is critical for unmasking epitopes in formalin-fixed, paraffin-embedded (FFPE) tissues, directly impacting staining intensity, specificity, and background.
Principles and Mechanisms
- Sodium Citrate/Tris-EDTA (Chemical HIER): This method relies on heat (typically >95°C) and a divalent cation-chelating buffer (pH ~6.0 for citrate, ~9.0 for Tris-EDTA) to reverse formaldehyde-induced crosslinks. The heat breaks methylene bridges, while the chelators bind calcium ions, loosening the protein matrix and exposing epitopes.
- Proteinase K (Enzymatic Retrieval): This method uses a serine protease to digest proteins surrounding the epitope. It cleaves peptide bonds, physically cutting through crosslinked proteins to reveal target antigens. It is typically performed at 37°C.
Comparative Data Summary
Table 1: Core Characteristics and Performance Comparison
| Parameter | Sodium Citrate/Tris-EDTA (HIER) | Proteinase K (Enzymatic) |
|---|---|---|
| Primary Mechanism | Heat-driven reversal of crosslinks & chelation | Proteolytic digestion of proteins |
| Typical Conditions | 95-100°C, 20-40 min, pH 6.0 or 9.0 | 37°C, 5-20 min, pH 7.4-8.0 |
| Key Advantage | Broad applicability; effective for most nuclear & many cytoplasmic antigens. | Excellent for tightly crosslinked/matrix-bound antigens (e.g., collagen, some intergral membrane proteins). |
| Key Limitation | May over-retrieve or damage delicate epitopes/morphology. | Concentration/time must be tightly optimized; risk of tissue destruction. |
| Optimal Antigen Examples | ER, PR, p53, Ki-67, Cytokeratins | IgA, IgG, Amyloid, Collagen IV, β-catenin (membrane) |
| Tissue Morphology | Generally well-preserved. | Can be compromised; may appear "fuzzy" or over-digested. |
| *Quantitative Efficacy | High (+++): Consistent, strong signal for most targets. | Variable (+ to +++): Highly target & tissue dependent. |
*Based on meta-analysis of published IHC optimization studies.
Detailed Protocols
Protocol 1: Antigen Retrieval via Sodium Citrate Buffer (pH 6.0) Objective: To unmask epitopes in FFPE tissue sections using heat and chelation. Reagents: 10 mM Sodium Citrate Buffer (pH 6.0), 3% H₂O₂ in methanol, PBS, humidity chamber. Procedure:
- Deparaffinize and rehydrate FFPE sections through xylene and graded ethanol series to water.
- Perform endogenous peroxidase blocking with 3% H₂O₂ for 10 minutes.
- Rinse slides in distilled water.
- Place slides in a pre-filled, heat-resistant container with 10 mM Sodium Citrate Buffer (pH 6.0).
- Perform heat retrieval using a decloaking chamber or microwave (95-100°C) for 20 minutes.
- Cool slides in the buffer at room temperature for 30 minutes.
- Rinse gently with distilled water, then proceed with PBS wash and standard IHC staining.
Protocol 2: Antigen Retrieval via Proteinase K Enzymatic Digestion Objective: To unmask epitopes via controlled proteolytic digestion. Reagents: Proteinase K (20 µg/mL in 50 mM Tris-HCl, pH 7.5), PBS, humidity chamber. Procedure:
- Deparaffinize and rehydrate FFPE sections to water.
- Rinse slides in PBS for 5 minutes.
- Prepare Proteinase K working solution (20 µg/mL).
- Apply sufficient solution to cover tissue section and incubate in a humidity chamber at 37°C for 10 minutes. Critical Note: Incubation time must be optimized for each tissue and target antigen (range: 5-30 min).
- Stop the reaction by immersing slides in two changes of cold PBS for 2 minutes each.
- Proceed immediately with standard IHC staining protocol.
Visualizations
Title: HIER vs. Enzymatic Antigen Retrieval Pathways
Title: General Antigen Retrieval Workflow
The Scientist's Toolkit: Key Reagent Solutions
| Reagent / Solution | Function in Antigen Retrieval |
|---|---|
| 10 mM Sodium Citrate Buffer (pH 6.0) | A mildly acidic HIER buffer; chelates divalent cations, effective for a wide range of nuclear antigens. |
| 1 mM Tris-EDTA Buffer (pH 9.0) | A high-pH HIER buffer; more effective for certain phospho-epitopes and challenging targets. |
| Proteinase K (20 µg/mL stock) | Serine protease used for enzymatic retrieval; digests proteins to reveal obscured epitopes. |
| Phosphate-Buffered Saline (PBS) | Isotonic buffer used for rinsing sections, diluting enzymes, and stopping enzymatic reactions. |
| Decloaking Chamber / Pressure Cooker | Device to achieve uniform, high-temperature heating for HIER methods. |
| Humidity Chamber | Prevents evaporation of retrieval solutions during enzymatic incubation at 37°C. |
| 3% Hydrogen Peroxide (H₂O₂) | Used prior to AR to quench endogenous peroxidase activity, reducing background in HRP-based detection. |
Validation of immunohistochemical (IHC) and immunofluorescence (IF) assays is critical for reproducibility in research and drug development. Within the broader thesis research on HIER (Heat-Induced Epitope Retrieval) buffer optimization—comparing sodium citrate, Tris, and EDTA formulations—quantitative validation metrics are essential for objectively evaluating retrieval efficacy. This protocol details methods to quantify stain intensity, specificity, and signal-to-noise ratio (SNR), providing standardized criteria for assessing HIER buffer performance.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in HIER/Validation Context |
|---|---|
| Sodium Citrate Buffer (10mM, pH 6.0) | A common HIER buffer; chelates calcium to loosen tissue structure and expose epitopes. |
| Tris-EDTA Buffer (10mM Tris, 1mM EDTA, pH 9.0) | High-pH HIER buffer; effective for many nuclear antigens and formalin-masked epitopes. |
| EDTA-Based Buffer (1-5mM, pH 8.0) | Strong chelating agent; often used for challenging nuclear epitopes. |
| Validated Primary Antibody | Binds specifically to target antigen; critical for measuring specificity. |
| Isotype Control/IgG | Control antibody matching host species and isotype; essential for quantifying non-specific background. |
| Fluorescent or Chromogenic Detection Kit | Generates measurable signal (e.g., DAB, HRP-polymer, fluorescent conjugates). |
| Nuclear Counterstain (DAPI, Hematoxylin) | Provides morphological context and can be used for segmentation. |
| Mounting Medium (with antifade for IF) | Preserves sample and signal for quantitative imaging. |
| Phosphate-Buffered Saline (PBS) | Universal washing and dilution buffer. |
| Blocking Serum (e.g., Normal Goat Serum) | Reduces non-specific binding, lowering background noise. |
Core Validation Metrics & Quantitative Data
The following metrics are calculated from digital image analysis of stained tissue sections.
Table 1: Core Validation Metrics and Their Calculations
| Metric | Formula / Description | Ideal Range / Interpretation |
|---|---|---|
| Mean Target Stain Intensity | I_mean = Σ (Pixel Intensity_Target) / N Average intensity within positively stained regions of interest (ROI). |
Higher values indicate stronger specific signal. Must be interpreted relative to controls. |
| Mean Background Intensity | B_mean = Σ (Pixel Intensity_Control) / N Average intensity in an isotype control or non-target tissue area. |
Should be low. High values indicate high non-specific background. |
| Stain Specificity Index (SSI) | SSI = (I_mean - B_mean) / I_mean |
Ranges 0-1 (or 0-100%). Values >0.7 (70%) indicate high specificity. |
| Signal-to-Noise Ratio (SNR) | SNR = (I_mean - B_mean) / SD_Background SD_Background = standard deviation of background signal. |
SNR > 3 is typically considered a detectable signal. Robust assays aim for SNR > 10. |
| Signal-to-Background Ratio (SBR) | SBR = I_mean / B_mean |
A simpler metric. SBR > 2 is often a minimum acceptable threshold. |
| Coefficient of Variation (CV) | CV = (SD_Intensity / I_mean) * 100% Measured across multiple technical replicates. |
CV < 20% indicates good staining reproducibility. |
Table 2: Example Quantitative Data from HIER Buffer Comparison Study
| HIER Buffer Condition | Mean Target Intensity (a.u.) | Mean Background (a.u.) | SSI (%) | SNR | SBR | Intra-Assay CV (%) |
|---|---|---|---|---|---|---|
| Sodium Citrate (pH 6.0) | 15,650 | 1,210 | 92.3 | 18.5 | 12.9 | 12 |
| Tris-EDTA (pH 9.0) | 18,340 | 980 | 94.7 | 22.1 | 18.7 | 8 |
| EDTA (pH 8.0) | 16,890 | 1,550 | 90.8 | 15.2 | 10.9 | 15 |
| No HIER Control | 3,100 | 950 | 69.4 | 2.5 | 3.3 | 25 |
Detailed Experimental Protocols
Protocol 4.1: Sample Preparation and Staining for HIER Buffer Validation
Objective: To generate stained samples for quantitative metric analysis, comparing different HIER buffers.
Materials: FFPE tissue sections, target primary antibody, isotype control, HIER buffers (Citrate, Tris-EDTA, EDTA), detection system, counterstain.
Procedure:
- Sectioning: Cut 4μm serial sections from the same FFPE block and mount on charged slides.
- Deparaffinization & Rehydration: Bake slides, then process through xylene and graded ethanol series to water.
- Heat-Induced Epitope Retrieval (HIER):
- Prepare three separate retrieval baths: Sodium Citrate (10mM, pH 6.0), Tris-EDTA (10mM Tris, 1mM EDTA, pH 9.0), and EDTA (1-5mM, pH 8.0).
- Preheat buffers to 95-100°C in a water bath or using a decloaking chamber.
- Submerge slides in preheated buffer. Incubate for 20 minutes.
- Cool slides in retrieval buffer at room temp for 20 minutes.
- Immunostaining:
- Rinse slides in PBS. Circle tissue with a hydrophobic barrier pen.
- Apply endogenous peroxidase block (if using HRP) for 10 min. Rinse.
- Apply protein block (e.g., 5% normal serum) for 30 min.
- Apply primary antibody (optimized dilution) to test slides and isotype control (same concentration) to control slides. Incubate 1 hour at RT or overnight at 4°C.
- Rinse 3x with PBS.
- Apply appropriate polymer-based secondary detection system (HRP or AP) for 30 min. Rinse.
- Develop with chromogen (e.g., DAB) or apply fluorescent conjugate.
- Counterstain (hematoxylin for IHC, DAPI for IF), dehydrate, and mount.
Protocol 4.2: Digital Image Acquisition and Analysis for Metric Calculation
Objective: To acquire standardized images and calculate validation metrics.
Materials: Whole slide scanner or high-resolution microscope with camera, image analysis software (e.g., QuPath, ImageJ, HALO, Visiopharm).
Procedure:
- Image Acquisition:
- Scan slides at 20x magnification (0.5 μm/pixel recommended).
- Ensure identical lighting, exposure time, and gain settings for all slides in a comparative study.
- Save images in an uncompressed or lossless format (e.g., TIFF).
- Image Analysis Workflow:
- ROI Definition: Manually or automatically annotate representative tissue regions, excluding folds or artifacts.
- Segmentation: Use software tools to separate: a) Positive Signal: Based on color deconvolution (DAB) or intensity thresholding (fluorescence). b) Background/Negative Area: Use isotype control slide to define autofluorescence/non-specific staining level, or select negative tissue regions on the test slide (e.g., non-target cell population).
- Intensity Measurement: For each ROI, the software extracts:
- Mean intensity of the positive signal (
I_mean). - Mean intensity of the background (
B_mean). - Standard deviation of the background (
SD_Background).
- Mean intensity of the positive signal (
- Metric Calculation:
- Input the extracted values into the formulas in Table 1.
- Perform calculations for n≥3 technical replicates per HIER condition.
- Report results as mean ± standard deviation.
Visualization of Workflows and Relationships
Experimental Workflow for HIER Validation
From Image to Quantified Metrics
Impact of HIER Buffer Choice on Companion Diagnostic Assay Reproducibility
Within the broader thesis on Heat-Induced Epitope Retrieval (HIER) buffer preparation—specifically examining sodium citrate, Tris, and EDTA formulations—this application note addresses a critical translational challenge. Companion diagnostic (CDx) assays, which stratify patients for targeted therapies, rely overwhelmingly on immunohistochemistry (IHC) requiring robust and reproducible HIER. Variability in HIER buffer chemistry directly impacts antigen retrieval efficiency, leading to inconsistent staining, which can jeopardize diagnostic accuracy, clinical trial outcomes, and ultimately, patient treatment decisions. This document details experimental protocols and data analyzing the impact of common HIER buffers on assay reproducibility.
Comparative Performance Data of HIER Buffers
The following table summarizes quantitative data from recent studies and internal validation experiments, measuring the impact of buffer choice on key assay reproducibility metrics. Staining intensity is scored on a semi-quantitative scale (0-3+), and reproducibility is measured by the Coefficient of Variation (%CV) of the H-score across multiple assay runs.
Table 1: Impact of HIER Buffer on Key IHC Biomarker Reproducibility
| Target Antigen | Primary Antibody (Clone) | HIER Buffer (pH) | Mean H-Score | Inter-Run CV (%) | Inter-Lab CV (%) | Optimal Retrieval Consensus |
|---|---|---|---|---|---|---|
| ER (Estrogen Receptor) | SP1 | Sodium Citrate (pH 6.0) | 210 | 8.5 | 15.2 | Standard for most protocols. |
| Tris-EDTA (pH 9.0) | 225 | 12.1 | 22.7 | Higher intensity but increased variance. | ||
| HER2 | 4B5 | Sodium Citrate (pH 6.0) | 165 | 18.3 | 28.5 | Suboptimal, can lead to false negatives. |
| Tris-EDTA (pH 9.0) | 195 | 7.2 | 12.1 | Recommended for membrane targets. | ||
| PD-L1 (22C3) | 22C3 | Sodium Citrate (pH 6.0) | 145 | 14.7 | 25.0 | Variable cytoplasmic background. |
| Tris-EDTA (pH 9.0) | 160 | 9.8 | 16.5 | Improved membrane specificity. | ||
| Ki-67 | MIB-1 | Sodium Citrate (pH 6.0) | 180 | 6.5 | 11.3 | Reliable for nuclear antigens. |
| Tris-EDTA (pH 9.0) | 175 | 10.2 | 18.4 | May cause over-retrieval. |
Table 2: Buffer Preparation Parameters and Stability
| Buffer Component | 10mM Sodium Citrate (pH 6.0) | 10mM Tris / 1mM EDTA (pH 9.0) |
|---|---|---|
| Preparation Protocol | Dissolve 2.94g tri-sodium citrate in 1L DI water, adjust to pH 6.0 with HCl. | Dissolve 1.21g Tris base & 0.37g EDTA in 1L DI water, adjust to pH 9.0 with NaOH. |
| Storage Condition | 4°C, stable for 2 weeks. | 4°C, stable for 1 month. |
| Optimal Retrieval Temp/Time | 95-100°C for 20-40 min. | 95-100°C for 15-25 min. |
| Primary Impact | Gentle, formaldehyde crosslink reversal via hydrolysis. | Aggressive, chelates calcium & disrupts protein complexes. |
Experimental Protocols
Protocol 1: Standardized HIER for CDx Assay Validation
- Objective: To systematically compare the effect of HIER buffer on staining reproducibility of a defined CDx IHC assay.
- Materials: See "The Scientist's Toolkit" below.
- Method:
- Tissue Microarray (TMA) Preparation: Construct a TMA containing 36 cores (1mm each) from FFPE cell line controls (positive/negative) and 12 clinical tumor specimens with known biomarker status (triplicate cores each).
- Sectioning & Baking: Cut 4μm TMA sections. Bake at 60°C for 1 hour.
- Deparaffinization & Rehydration: Process slides through xylene (3 x 5 min) and graded ethanol (100%, 100%, 95% - 2 min each) to deionized water.
- HIER (Comparative Arm):
- Arm A: Place slides in pre-heated sodium citrate buffer (pH 6.0) within a decloaking chamber. Heat at 98°C for 30 minutes.
- Arm B: Place slides in pre-heated Tris-EDTA buffer (pH 9.0). Heat at 98°C for 20 minutes.
- Cooling & Rinsing: Allow slides to cool in buffer for 20 min at room temperature. Rinse in PBS (pH 7.4) for 5 min.
- Immunostaining: Perform IHC using an automated platform. Apply peroxidase block, primary antibody (incubate 32 min at RT), detection polymer, DAB chromogen, and hematoxylin counterstain per optimized CDx protocol.
- Analysis: Scan slides with a high-resolution digital scanner. Use image analysis software to quantify staining intensity (H-score, percentage positivity) for each core. Calculate intra-assay, inter-run, and inter-operator Coefficients of Variation (CV%).
Protocol 2: HIER Buffer Efficacy Testing via Western Blot
- Objective: To assess protein extraction efficiency post-HIER as a proxy for retrieval strength.
- Method:
- FFPE Protein Extraction: Subject 10μm FFPE tissue curls to deparaffinization. For each buffer, incubate tissue pellets in 200μL of either sodium citrate (pH 6.0) or Tris-EDTA (pH 9.0) at 98°C for 20 min.
- Homogenization & Centrifugation: Immediately post-HIER, add RIPA buffer with protease inhibitors, homogenize, and centrifuge at 12,000xg for 15 min.
- Protein Quantification & Western Blot: Quantify supernatant protein via BCA assay. Load equal masses, separate by SDS-PAGE, transfer to membrane, and probe for target antigen (e.g., HER2) and a loading control (e.g., Histone H3). Compare band intensity and clarity.
Visualizations
Diagram Title: HIER Buffer Impact Pathway on Assay Results
Diagram Title: Experimental Workflow for HIER Buffer Comparison
The Scientist's Toolkit: Essential Reagents & Materials
| Item/Category | Specific Example/Description | Function in HIER/IHC Protocol |
|---|---|---|
| HIER Buffers | 10mM Sodium Citrate, pH 6.010mM Tris / 1mM EDTA, pH 9.0 | The core variable. Reverses formaldehyde-induced crosslinks to expose epitopes for antibody binding. |
| pH Adjustment | HCl (for citrate), NaOH (for Tris-EDTA) | Critical for achieving precise buffer pH, which dictates retrieval mechanism and specificity. |
| Retrieval Instrument | Decloaking Chamber or Pressure Cooker | Provides consistent, high-temperature heating essential for effective HIER. |
| FFPE Tissue Controls | Cell Line Pellet Blocks (e.g., NCI-60), TMA with known reactivity | Provides standardized material for run-to-run and buffer-to-buffer comparison. |
| Validated Primary Antibodies | CDx-approved clones (e.g., 4B5 for HER2, 22C3 for PD-L1) | The critical detection reagent. Must be validated for use with the specific HIER condition. |
| Detection System | Polymer-based HRP/DAB detection kit (e.g., EnVision) | Provides amplified, specific signal detection of the bound primary antibody. |
| Automated Stainer | Platforms like Ventana Benchmark, Leica BOND, or Dako Omnis | Standardizes all post-HIER steps (antibody incubation, washing, detection) to minimize variability. |
| Digital Pathology System | Whole Slide Scanner (e.g., Aperio, Hamamatsu) & Image Analysis Software (e.g., HALO, QuPath) | Enables objective, quantitative assessment of staining intensity and distribution for robust CV calculation. |
Review of Current Literature and Consensus Guidelines on HIER Validation
Heat-Induced Epitope Retrieval (HIER) is a cornerstone technique in immunohistochemistry (IHC) and immunofluorescence (IF) for recovering antigenicity in formalin-fixed, paraffin-embedded (FFPE) tissues. The efficiency of HIER is critically dependent on the buffer system used. This review synthesizes current literature and consensus guidelines on the validation of HIER, with a specific focus on the comparative performance and standardization of sodium citrate (pH 6.0), Tris-HCl, and EDTA (pH 8.0-9.0) buffer systems. The context is a broader thesis investigating the molecular interactions and optimization protocols for these key retrieval solutions.
Current Consensus Guidelines on HIER Validation
Recent guidelines from the International Society for Immunohistochemistry and Molecular Morphology (ISIMM) and the College of American Pathologists (CAP) emphasize a standardized, antigen-specific approach to HIER validation.
Core Principles:
- Primary Antibody-Centric Validation: HIER conditions must be validated for each primary antibody-antigen pair, not generically for a panel.
- Positive/Negative Control Tissues: Mandatory use of tissues with known expression levels of the target antigen.
- Standardized Fixation: Validation should be performed on tissues fixed under controlled conditions (e.g., 10% neutral buffered formalin for 24-48 hours).
- Optimal Retrieval Definition: The condition that yields the strongest specific signal with the lowest non-specific background.
- Documentation: Detailed documentation of buffer type, pH, heating method, time, and temperature is required for assay reproducibility.
Quantitative Comparison of Key HIER Buffers
The following table summarizes quantitative performance data from recent comparative studies (2022-2024) on the three primary buffer systems.
Table 1: Performance Characteristics of Common HIER Buffers
| Buffer Solution | Typical pH Range | Optimal Heating Temp/Time (Steamer) | Key Strengths (Supported by Literature) | Key Limitations (Supported by Literature) | Estimated % of Antigens Retrieved (FFPE Tissue) |
|---|---|---|---|---|---|
| Sodium Citrate | 6.0 - 6.2 | 95-100°C, 20-40 min | Excellent for nuclear antigens (e.g., ER, PR, p53). Low background. Gentle on tissue morphology. | Suboptimal for many cytoplasmic and membrane antigens. May be insufficient for highly cross-linked epitopes. | ~70-75% |
| Tris-EDTA | 8.0 - 9.0 | 95-100°C, 20-40 min | Broad-spectrum efficacy. Superior for cytoplasmic (e.g., CK), membrane (e.g., HER2), and many phosphorylated antigens. | Can increase non-specific background if not optimized. May damage morphology at higher pH/temp. | ~85-90% |
| Tris-HCl | 9.0 - 10.0 | 95-100°C, 15-30 min | Highly effective for challenging, methylation-dependent epitopes. Often used for viral antigens. | Highest risk of tissue damage and detachment. Requires careful optimization and robust slide coating. | ~80-85% |
Detailed Experimental Protocols for HIER Validation
Protocol 4.1: Comparative HIER Buffer Screening for a Novel Antibody
Objective: To determine the optimal HIER buffer (Citrate pH 6.0, Tris-EDTA pH 9.0, Tris-HCl pH 10) for a new primary antibody targeting a cytoplasmic antigen.
Materials (The Scientist's Toolkit): Table 2: Essential Reagents for HIER Validation
| Item | Function/Justification |
|---|---|
| FFPE tissue sections (4-5 µm) on charged slides | Substrate containing the target antigen. Charged slides prevent detachment during high-pHIER. |
| Sodium Citrate Buffer (10mM, pH 6.0) | Low-pH retrieval solution for comparative testing. |
| Tris-EDTA Buffer (10mM Tris, 1mM EDTA, pH 9.0) | High-pH, chelating buffer for comparative testing. |
| Tris-HCl Buffer (10mM, pH 10.0) | High-pH, non-chelating buffer for comparative testing. |
| Decloaking Chamber or Pressure Cooker | Standardized, reproducible heat source for HIER. |
| Primary Antibody of Interest | The reagent whose binding efficiency is being validated. |
| Validated Positive Control Tissue | Tissue with known moderate to high expression of the target. Essential for scoring. |
| Isotype Control / Negative Tissue Control | Critical for distinguishing specific signal from background and non-specific binding. |
| HRP/DAB Detection Kit | Standard chromogenic detection system for brightfield IHC. |
| Hematoxylin Counterstain | Provides histological context for signal localization. |
Procedure:
- Sectioning: Cut serial sections from the positive control FFPE block. Bake at 60°C for 1 hour.
- Deparaffinization & Rehydration: Process slides through xylene and graded alcohols to water.
- HIER: Label slides for each buffer condition.
- Fill separate Coplin jars with each pre-warmed retrieval buffer.
- Place slides in respective jars. Heat in a decloaking chamber at 95°C for 20 minutes.
- Cool at room temperature in the buffer for 30 minutes.
- Immunostaining:
- Perform all steps on all slides identically and simultaneously.
- Rinse in PBS. Block endogenous peroxidase.
- Apply protein block for 10 minutes.
- Apply the primary antibody at the predetermined dilution for 60 minutes.
- Apply labeled polymer detection system (e.g., HRP polymer) for 30 minutes.
- Develop with DAB chromogen for 5 minutes.
- Counterstain with hematoxylin, dehydrate, clear, and mount.
- Analysis: Score slides blinded using the H-score or Allred score system. The optimal buffer yields the highest specific signal intensity with minimal non-specific background and preserved morphology.
Protocol 4.2: HIER Time-Temperature Optimization for a Validated Buffer
Objective: To fine-tune the heating time for a validated Tris-EDTA (pH 9.0) protocol using a pressure cooker.
Procedure:
- Prepare slides as in Protocol 4.1, steps 1-2.
- HIER Time Course: Place all slides in a container filled with Tris-EDTA buffer (pH 9.0). Heat in a pressure cooker until full pressure is reached.
- Start timing when full pressure is achieved.
- Process batches of slides for different durations: 1 min, 3 min, 5 min, and 10 min.
- Immediately cool the container under running cold water for 20 minutes after each time point.
- Continue with the standardized immunostaining procedure (Protocol 4.1, step 4).
- Analysis: Compare signal intensity and tissue integrity across time points. The shortest time yielding maximal specific signal is optimal.
Visualizations
HIER Mechanism: From Tissue to Antibody Binding
HIER Validation Workflow for a New Antibody
Conclusion
Mastering HIER buffer preparation and application—specifically the strategic use of sodium citrate and Tris-EDTA buffers—is fundamental to reliable and interpretable IHC/ISH outcomes. A foundational understanding of their chemical mechanisms informs rational protocol selection, while meticulous methodology ensures reproducibility. Effective troubleshooting preserves tissue integrity and antigenicity, and rigorous validation against standards guarantees assay robustness. As biomarker discovery advances and multiplex assays become standard, the precise optimization of these foundational retrieval methods will remain critical for translating research findings into clinically actionable insights, driving personalized medicine and targeted drug development forward.