HIER vs PIER for IHC: A Comprehensive Guide for Researchers on Antigen Retrieval Methods
This article provides a detailed, evidence-based comparison of Heat-Induced Epitope Retrieval (HIER) and Proteolytic-Induced Epitope Retrieval (PIER) for immunohistochemistry.
HIER vs PIER for IHC: A Comprehensive Guide for Researchers on Antigen Retrieval Methods
Abstract
This article provides a detailed, evidence-based comparison of Heat-Induced Epitope Retrieval (HIER) and Proteolytic-Induced Epitope Retrieval (PIER) for immunohistochemistry. Targeted at researchers and drug development professionals, it explores the fundamental principles, protocols, and applications of each method. It addresses common challenges, optimization strategies, and head-to-head validation data to empower scientists in selecting the optimal antigen retrieval technique for their specific biomarkers, tissue types, and research objectives, ultimately enhancing reproducibility and data quality in biomedical research.
Understanding HIER and PIER: Core Principles and Historical Context in IHC
Antigen Retrieval (AR) is the cornerstone process in immunohistochemistry (IHC) that reverses formaldehyde-induced cross-links, thereby recovering antigenicity and enabling specific antibody binding. Its development transformed IHC from a capricious technique into a robust, reproducible pillar of diagnostic and research pathology. The efficacy of AR is the primary determinant of staining success in formalin-fixed, paraffin-embedded (FFPE) tissues. Within this field, the debate between Heat-Induced Epitope Retrieval (HIER) and Proteolytic-Induced Epitope Retrieval (PIER) constitutes a core methodological thesis, directly impacting data accuracy and biological interpretation.
The Chemical Basis of Epitope Masking and Retrieval
Formalin fixation creates methylene bridges between proteins, obscuring epitopes. AR breaks these cross-links.
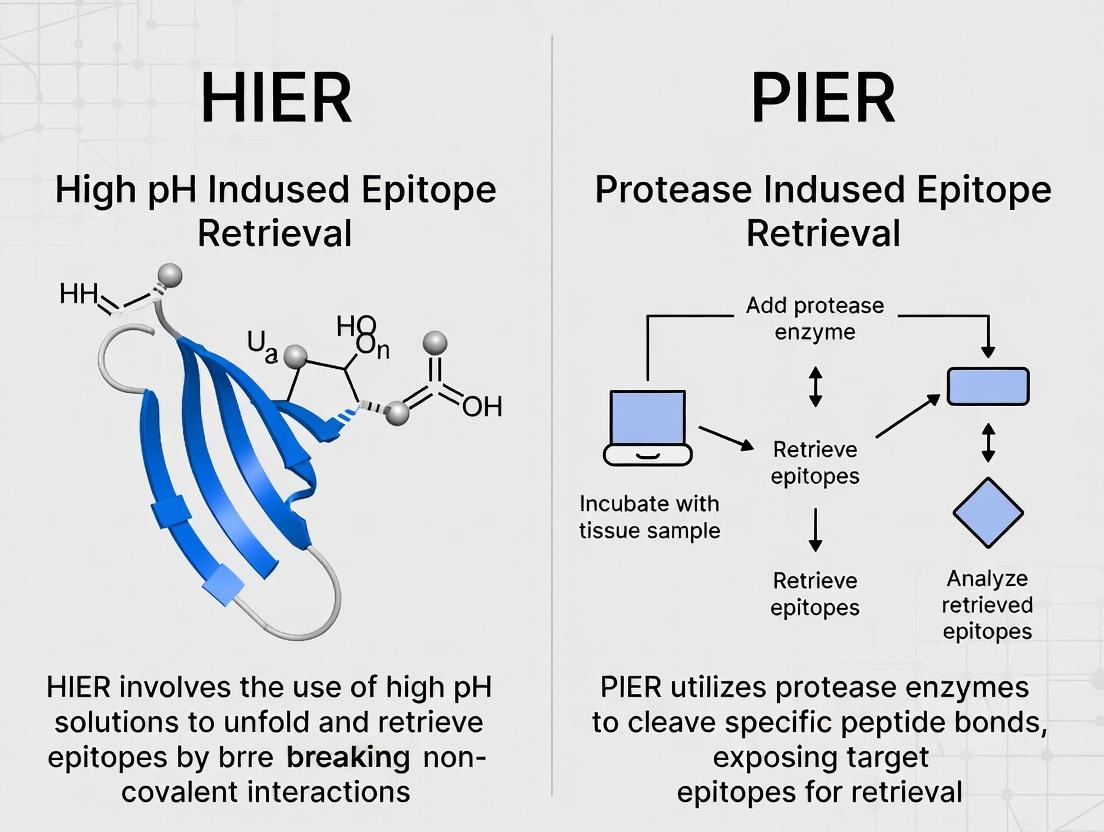
- HIER: Uses heat (95-120°C) and a chemical buffer (e.g., citrate, EDTA, Tris-EDTA) to hydrolyze cross-links.
- PIER: Uses proteolytic enzymes (e.g., trypsin, pepsin, proteinase K) to cleave proteins and expose epitopes.
The choice between HIER and PIER is antigen-specific and depends on the nature of the cross-linking and the epitope's vulnerability to enzymatic digestion.
HIER vs. PIER: A Quantitative Comparative Analysis
Table 1: Core Methodological Comparison of HIER and PIER
| Parameter | Heat-Induced Epitope Retrieval (HIER) | Proteolytic-Induced Epitope Retrieval (PIER) |
|---|---|---|
| Primary Mechanism | Hydrolytic cleavage of methylene cross-links via heat & buffer. | Enzymatic digestion of protein sequences surrounding epitope. |
| Typical Agents | Citrate buffer (pH 6.0), Tris-EDTA (pH 9.0), EDTA (pH 8.0). | Trypsin, Pepsin, Proteinase K. |
| Typical Conditions | 95-120°C for 10-40 minutes. | 37°C for 5-30 minutes. |
| Key Advantage | Broad applicability; superior for most nuclear & many cytoplasmic antigens. | Effective for some antigens resistant to HIER (e.g., collagen, tight junctions). |
| Key Limitation | Can destroy fragile epitopes; requires precise pH optimization. | Over-digestion risks destroying epitope & tissue morphology. |
| Optimal For | Phospho-epitopes, nuclear antigens (ER, PR, p53), membrane antigens. | Extracellular matrix antigens, some tightly cross-linked epitopes. |
Table 2: Experimental Outcomes from a Representative Comparative Study (Staining Intensity & Clarity Score, 0-3 scale)
| Target Antigen (Localization) | HIER (Citrate, pH 6.0) | HIER (Tris-EDTA, pH 9.0) | PIER (Trypsin) | Recommended Method |
|---|---|---|---|---|
| ER (Nuclear) | 3.0 | 2.8 | 0.5 | HIER (Citrate) |
| Ki-67 (Nuclear) | 2.9 | 3.0 | 0.7 | HIER (Tris-EDTA) |
| Her2/neu (Membrane) | 2.5 | 2.7 | 1.2 | HIER (Tris-EDTA) |
| Collagen IV (ECM) | 1.0 | 1.1 | 2.9 | PIER (Trypsin) |
| Cytokeratin (Cytoplasmic) | 2.8 | 2.6 | 1.8 | HIER (Citrate) |
Experimental Protocol: Direct Comparison of HIER vs. PIER
Objective: To determine the optimal AR method for a novel nuclear antigen (Target X) in FFPE human tonsil tissue.
Protocol:
- Sectioning: Cut 4μm serial sections from FFPE block.
- Deparaffinization & Rehydration: Standard xylene and graded ethanol series.
- Antigen Retrieval (Parallel Methods):
- Group A (HIER-Citrate): Place slides in 10mM Sodium Citrate buffer (pH 6.0). Heat in decloaking chamber or pressure cooker at 95-100°C for 20 minutes. Cool at room temp for 30 mins.
- Group B (HIER-Tris): Place slides in 10mM Tris-EDTA buffer (pH 9.0). Process as in Group A.
- Group C (PIER): Incubate slides with 0.1% Trypsin solution in 0.1% CaCl2 (pH 7.8) at 37°C for 10 minutes. Rinse in PBS.
- Peroxidase Blocking: 3% H₂O₂ for 10 minutes.
- Primary Antibody: Incubate with anti-Target X antibody (optimized dilution) for 1 hour at room temp.
- Detection: Apply polymer-based HRP-conjugated secondary antibody for 30 mins, followed by DAB chromogen.
- Counterstaining & Analysis: Hematoxylin counterstain, dehydrate, mount. Score staining intensity (0-3) and proportion of positive cells by two blinded pathologists.
Signaling Pathway & Experimental Workflow
Title: Decision Workflow: HIER vs. PIER in IHC
Title: Core IHC Protocol with Antigen Retrieval Step
The Scientist's Toolkit: Key Reagent Solutions
Table 3: Essential Research Reagents for Antigen Retrieval Optimization
| Reagent / Material | Primary Function & Rationale |
|---|---|
| Sodium Citrate Buffer (10mM, pH 6.0) | Standard HIER buffer for hydrolytic retrieval; ideal for many nuclear antigens. |
| Tris-EDTA Buffer (10mM, pH 9.0) | High-pHIER buffer; effective for phospho-epitopes and many membrane targets. |
| Trypsin, Pepsin, or Proteinase K | Enzymes for PIER; cleave specific peptide bonds to expose resistant epitopes. |
| Decloaking Chamber or Pressure Cooker | Provides consistent, high-temperature heating for HIER protocols. |
| pH Meter & Calibrated Buffers | Critical for accurate AR buffer preparation; pH is a key variable. |
| Positive Control FFPE Tissue | Tissue known to express target antigen; mandatory for validating AR conditions. |
| Polymer-based HRP Detection System | High-sensitivity, low-background detection post-AR, replacing older ABC methods. |
| Antibody Diluent (with Carrier Protein) | Stabilizes primary antibody incubation after aggressive AR treatments. |
Within the ongoing research discourse comparing Heat-Induced Epitope Retrieval (HIER) and Proteolytic-Induced Epitope Retrieval (PIER) for immunohistochemistry (IHC), HIER has emerged as the predominant and often superior method for recovering antigenicity in formalin-fixed, paraffin-embedded (FFPE) tissues. This whitepaper provides an in-depth technical analysis of the core scientific mechanism underpinning HIER. Understanding this mechanism is critical for researchers and drug development professionals optimizing IHC protocols for biomarker validation and diagnostic assay development.
Core Mechanism: Reversal of Formalin-Induced Crosslinks
The primary action of HIER is the hydrolysis of methylene bridges formed during formalin fixation. Formaldehyde reacts with amino groups of proteins, creating crosslinks that mask epitopes. Heat application (typically 95-100°C or higher in pressure cookers) provides kinetic energy that accelerates the breakage of these crosslinks.
Chemical Reactions During HIER
The process involves:
- Rehydration: The tissue section is rehydrated, allowing the retrieval buffer to penetrate.
- Heat-Driven Denaturation: Applied heat disrupts hydrogen bonds and van der Waals forces, partially denaturing proteins and increasing solvent accessibility.
- Hydrolysis of Crosslinks: The critical step is the hydrolysis of the methylene bridges (-CH2-) between proteins. This is significantly accelerated by high temperature and the pH of the retrieval buffer.
- Protein Unfolding: The breakage of crosslinks allows tightly folded proteins to partially relax, exposing previously buried epitopes for antibody binding.
Role of Buffer pH
The pH of the retrieval solution is a key variable that dictates which types of crosslinks are most effectively reversed:
- High-pH buffers (e.g., Tris-EDTA, pH 9.0): Promote ionization of amino acid side chains, increasing electrostatic repulsion which aids protein unfolding. Particularly effective for nuclear antigens (e.g., Ki-67, ER, PR) and many phosphorylated epitopes.
- Low-pH buffers (e.g., citrate, pH 6.0): Effective for a wide range of cytoplasmic and membrane antigens. The mechanism may involve protonation of specific chemical groups in the crosslinks.
Table 1: Quantitative Comparison of Common HIER Buffers
| Buffer Solution | Typical pH Range | Optimal Antigen Categories | Standard Incubation Time/Temp |
|---|---|---|---|
| Sodium Citrate | 6.0 - 6.2 | Cytoplasmic, membranous, viral antigens | 95-100°C, 20-40 min |
| Tris-EDTA | 8.0 - 9.0 | Nuclear antigens, phosphorylated epitopes | 95-100°C, 20-40 min |
| Target Retrieval Solution (DAKO) | 6.1 or 9.0 | Broad spectrum, vendor-optimized | As per vendor protocol |
| EDTA alone | 8.0 - 9.0 | Very tight crosslinks (e.g., MCM2, FoxP3) | 95-100°C, 30-45 min |
Detailed Experimental Protocol for HIER Optimization
The following protocol is a standard method for comparing HIER conditions in a research setting.
Protocol Title: Optimization of HIER for a Novel Epitope in FFPE Tissue.
Objective: To determine the optimal pH and heating time for unmasking a target antigen.
Materials: See "The Scientist's Toolkit" below.
Method:
- Sectioning: Cut 4-5 μm thick sections from the FFPE tissue block of interest and mount on charged slides.
- Baking: Bake slides at 60°C for 1 hour to enhance adhesion.
- Deparaffinization & Rehydration:
- Immerse slides in xylene (or xylene substitute) for 10 minutes. Repeat with fresh xylene for another 10 minutes.
- Rehydrate through a graded ethanol series: 100% ethanol (twice, 5 min each), 95% ethanol (5 min), 70% ethanol (5 min).
- Rinse in distilled water for 5 minutes.
- Antigen Retrieval Setup:
- Prepare separate Coplin jars or a slide rack with 1x retrieval buffers at pH 6.0 (citrate) and pH 9.0 (Tris-EDTA). Fill sufficiently to cover slides.
- Using a microwave, water bath, or commercial decloaking chamber/pressure cooker, pre-heat the buffer to the target temperature (95-100°C).
- Heat Treatment:
- Carefully place slides into the pre-heated buffer.
- For each buffer type, incubate separate slides for 10, 20, and 30 minutes at a maintained temperature just below boiling.
- Critical: Ensure slides remain fully immersed and the buffer does not evaporate to dryness.
- Cooling: After heating, remove the container from the heat source and allow it to cool at room temperature for 20-30 minutes. Do not cool rapidly, as this may promote protein refolding.
- Washing: Rinse slides gently in running distilled water, then transfer to Wash Buffer (1x PBS or TBS) for 5 minutes.
- Proceed to Immunostaining: Continue with standard IHC protocol steps (peroxidase blocking, protein blocking, primary antibody incubation, etc.).
Diagram 1: HIER Optimization Workflow
Title: Workflow for HIER Condition Optimization
Signaling and Molecular Pathways Impacted by HIER
HIER does not activate a biological signaling pathway but rather reverses artifactual chemical modifications. Its efficacy is determined by the chemistry of the crosslink and the local protein microenvironment.
Diagram 2: Molecular Mechanism of HIER
Title: Chemical Mechanism of Heat-Induced Epitope Unmasking
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for HIER Experiments
| Item | Function & Rationale |
|---|---|
| FFPE Tissue Sections | The standard archival material. Antigen masking is uniform, providing a consistent challenge for retrieval. |
| Sodium Citrate Buffer (10x, pH 6.0) | A low-pH, chelating buffer. Citrate ions may chelate calcium, potentially stabilizing proteins while heat breaks crosslinks. |
| Tris-EDTA Buffer (10x, pH 9.0) | A high-pH, chelating buffer. EDTA chelates divalent cations more strongly, which can disrupt additional protein structures. The high pH aids in breaking crosslinks involving tyrosine. |
| Commercial Antigen Retrieval Buffer (e.g., from Agilent, Abcam, Vector Labs) | Proprietary, often optimized blends that may contain denaturants, chelators, and detergents for consistent, high-performance retrieval. |
| Pressure Cooker / Commercial Decloaking Chamber | Provides a consistent, high-temperature environment (~120°C under pressure). Reduces retrieval time and often improves uniformity compared to microwave methods. |
| Microwave or Water Bath | Alternative heat sources. Microwave heating can be uneven, requiring careful protocol standardization. Water baths offer gentle, uniform heating but at lower max temperature (~95-98°C). |
| Charged Microscope Slides (e.g., positively charged) | Prevent tissue detachment during the high-temperature, high-fluid-shear stress of HIER. |
| Heat-Resistant Slide Rack and Container | Must withstand prolonged high temperatures without warping or leaching chemicals. |
| pH Meter | Critical for verifying the pH of prepared retrieval buffers, as minor deviations significantly impact results. |
Comparative Data: HIER vs. PIER in Research
The debate of "HIER vs. PIER which is better" is largely settled in favor of HIER for most applications, as supported by contemporary research.
Table 3: Quantitative and Qualitative Comparison of HIER vs. PIER
| Parameter | Heat-Induced Epitope Retrieval (HIER) | Proteolytic-Induced Epitope Retrieval (PIER) |
|---|---|---|
| Primary Mechanism | Chemical hydrolysis of methylene crosslinks. | Enzymatic cleavage of peptide bonds. |
| Typical Agents | Citrate, Tris-EDTA, proprietary buffers. | Trypsin, pepsin, proteinase K. |
| Treatment Time | 20-40 minutes (longer for some methods). | 5-30 minutes (highly enzyme-dependent). |
| Key Advantage | Superior for most antigens, especially nuclear. Gentle on tissue morphology. Broadly applicable. | Can be effective for some tightly fixed antigens where heat fails. |
| Key Disadvantage | May destroy some delicate epitopes sensitive to heat. | Over-digestion risk: Can destroy the epitope and damage tissue morphology. Less reproducible. |
| Impact on Morphology | Generally excellent preservation. | Often causes tissue fragility, loss of detail, or "honeycomb" artifact. |
| Modern Usage Prevalence | >95% of standard IHC protocols. | <5%, reserved for specific, stubborn antigens when HIER fails. |
| Research Consensus | Method of choice and default starting point. Offers superior consistency, signal-to-noise ratio, and morphology preservation. | Largely obsolete for routine work. Used as a last-resort alternative. |
The science of HIER is grounded in the heat- and pH-accelerated reversal of the formaldehyde chemistry that defines FFPE tissue preservation. Its mechanism—hydrolytic cleavage of methylene crosslinks—is more controllable, reproducible, and gentler on tissue architecture than the proteolytic scission employed by PIER. For the researcher or drug developer, HIER represents the unequivocal standard for epitope retrieval. The critical research task is not choosing between HIER and PIER, but rather systematically optimizing HIER conditions (buffer pH, heating time, temperature) for each novel antigen-antibody pair to achieve maximum specificity and sensitivity in IHC assays.
1. Introduction Proteolytic Induced Epitope Retrieval (PIER) is a critical technique in immunohistochemistry (IHC) used to unmask epitopes in formalin-fixed, paraffin-embedded (FFPE) tissue sections. It operates through the enzymatic cleavage of cross-links formed during fixation, contrasting with Heat-Induced Epitope Retrieval (HIER), which relies on heat and pH to break these bonds. This whitepaper details the biochemical mechanisms, protocols, and applications of PIER, framed within the ongoing scientific debate regarding the comparative efficacy of HIER versus PIER.
2. Core Mechanism of Enzymatic Cleavage in PIER Formalin fixation creates methylene bridges between proteins, obscuring antigenic sites. PIER utilizes specific proteases (e.g., trypsin, proteinase K, pepsin) to hydrolyze peptide bonds, thereby physically cleaving these cross-links and restoring antibody accessibility. The mechanism is a targeted catalytic process:
- Enzyme Binding: The protease binds to specific amino acid sequences on the target protein.
- Catalysis: The enzyme's active site facilitates nucleophilic attack on the carbonyl carbon of the peptide bond, leading to hydrolysis.
- Cleavage & Unmasking: The breakage of peptide bonds liberates the epitope from the cross-linked network.
The choice of enzyme is determined by the target antigen's amino acid composition and the tissue type.
3. Quantitative Comparison of PIER vs. HIER The selection between PIER and HIER is antigen- and tissue-dependent. The following table summarizes key performance characteristics based on recent literature.
Table 1: Comparative Analysis of PIER and HIER Methods
| Parameter | Proteolytic Induced Epitope Retrieval (PIER) | Heat-Induced Epitope Retrieval (HIER) |
|---|---|---|
| Primary Mechanism | Enzymatic hydrolysis of peptide bonds. | Thermo-chemical reversal of cross-links via heat & buffer. |
| Typical Conditions | 37°C, 10-30 mins; [Enzyme] 0.05-0.5% w/v. | 95-100°C, 20-40 mins; or 121°C, 10-15 mins (pressure). |
| Key Advantages | Gentle on tissue morphology; effective for many intracellular and nuclear antigens (e.g., collagen, cytokeratin). | Broad-spectrum efficacy; standardizable; superior for many membrane targets. |
| Key Limitations | Risk of over-digestion & tissue loss; enzyme-specific; less effective for some cross-linked epitopes. | Can damage delicate morphology; may not retrieve highly cross-linked epitopes. |
| Optimal Use Cases | FFPE tissues with extensive cross-linking; antigens sensitive to heat; collagen-rich matrices. | Most general IHC applications; when preserving fine ultrastructure is less critical. |
| Reported Success Rate (Range) | 60-85% for specific antigen classes. | 80-95% for a broad range of antigens. |
4. Experimental Protocol: Standard PIER Workflow
- Reagents: 0.05-0.4% Trypsin or Proteinase K in Tris-HCl or PBS buffer (pH 7.4-8.0); Phosphate Buffered Saline (PBS); Distilled Water.
- Procedure:
- Dewax & Hydrate: Deparaffinize FFPE sections in xylene and rehydrate through a graded ethanol series to distilled water.
- Buffer Rinse: Rinse slides in PBS for 5 minutes.
- Protease Digestion: Pipette pre-warmed enzyme solution to cover tissue section. Incubate in a humidified chamber at 37°C for 10-30 minutes. Optimal time must be determined empirically.
- Enzyme Inactivation: Rinse slides thoroughly in two changes of cold PBS for 5 minutes each to halt proteolytic activity.
- Immunostaining: Proceed with standard blocking, primary/secondary antibody incubation, and detection steps.
5. Visualization of PIER Mechanism and Workflow
Mechanism of Proteolytic Epitope Unmasking
Standard PIER Experimental Workflow
6. The Scientist's Toolkit: Key Research Reagents for PIER
Table 2: Essential Reagent Solutions for PIER Protocols
| Reagent | Typical Concentration/Type | Function in PIER |
|---|---|---|
| Trypsin | 0.05-0.1% in 0.1% CaCl₂, pH 7.8 | Serine protease cleaving after Lys/Arg residues; common for cytoplasmic antigens. |
| Proteinase K | 0.5-20 µg/mL in Tris-EDTA, pH 8.0 | Broad-spectrum serine protease; effective for highly cross-linked nuclear antigens. |
| Pepsin | 0.1-0.5% in 0.01N HCl | Aspartic protease active at low pH; used for extracellular matrix targets. |
| Tris-HCl Buffer | 0.05M, pH 7.4-8.0 | Maintains optimal pH for enzymatic activity and tissue integrity. |
| Phosphate Buffered Saline (PBS) | 0.01M, pH 7.2-7.4 | Used for rinsing slides to stop digestion and maintain physiological pH. |
| Calcium Chloride (CaCl₂) | 0.1% in trypsin solution | Cofactor required for trypsin stability and activity. |
7. Conclusion PIER remains an indispensable, mechanism-driven technique for antigen retrieval, particularly where HIER fails or damages morphology. Its efficacy is predicated on the precise matching of protease specificity to antigen and tissue context. The "HIER vs. PIER" debate is not a quest for a universal winner but a strategic decision tree for the researcher. Optimal IHC requires empirical validation of both methods, with PIER offering a powerful enzymatic solution to the challenge of formalin-induced epitope masking.
Within the ongoing debate on HIER (Heat-Induced Epitope Retrieval) versus PIER (Protease-Induced Epitope Retrieval) for optimal immunohistochemistry (IHC), the historical evolution of tissue fixation and antigen retrieval is paramount. This whitepaper details the technical progression from formalin fixation to modern retrieval breakthroughs, providing a foundation for evaluating the core methodologies at the heart of the HIER vs. PIER thesis.
Part 1: The Formalin Fixation Standard and Its Challenge
Formalin fixation, primarily using 10% Neutral Buffered Formalin (NBF), creates methylene cross-links between proteins, preserving tissue morphology but masking epitopes. This created a fundamental barrier to IHC.
Key Quantitative Data on Formalin Fixation Effects:
| Parameter | Typical Range/Value | Impact on IHC |
|---|---|---|
| Fixation Time (Optimal) | 18-24 hours | Under-fixation: poor morphology; Over-fixation: excessive masking. |
| Formaldehyde Concentration | 3.7-4.0% (in 10% NBF) | Standard for consistent cross-linking. |
| Cross-link Type Formed | Methylene bridges (-CH2-) | Primary cause of epitope masking. |
| pH of Fixative | 7.0-7.4 (Buffered) | Prevents acid-induced artifacts. |
Part 2: The Retrieval Revolution: HIER and PIER Mechanisms
The breakthrough came with the development of methods to reverse formalin-induced cross-linking. HIER and PIER represent two philosophically distinct approaches.
HIER (Heat-Induced Epitope Retrieval)
HIER uses heat (95-125°C) in a low-pH (citrate, pH ~6.0) or high-pH (Tris-EDTA, pH ~9.0) buffer to hydrolyze and break cross-links.
Detailed Protocol for Standard HIER (Citrate Buffer, pH 6.0):
- Deparaffinization & Rehydration: Slide incubation in xylene (2 changes, 5 min each), followed by graded ethanol (100%, 100%, 95%, 70%; 2 min each) and finally dH₂O.
- Retrieval Buffer Preparation: 10mM Sodium Citrate buffer, pH 6.0. Add 0.5 mL Tween 20 per 1000 mL.
- Heating: Place slides in a pre-filled, pre-heated (95-100°C) retrieval buffer. Maintain at sub-boiling temperature (95-100°C) for 20 minutes in a water bath or commercial decloaking chamber.
- Cooling: Remove container from heat and cool at room temperature for 20-30 minutes.
- Rinsing: Rinse slides in distilled water, then transfer to IHC wash buffer (e.g., PBS).
PIER (Protease-Induced Epitope Retrieval)
PIER uses proteolytic enzymes (e.g., trypsin, proteinase K) to cleave proteins and physically expose epitopes.
Detailed Protocol for Proteinase K Retrieval:
- Deparaffinization & Rehydration: As per HIER protocol steps.
- Enzyme Solution Preparation: Dilute proteinase K in 50mM Tris-HCl, 5mM EDTA, pH 7.5 buffer to a final concentration of 5-20 µg/mL.
- Digestion: Apply sufficient enzyme solution to cover tissue section. Incubate at 37°C for 5-15 minutes in a humidified chamber.
- Enzyme Inhibition: Rinse slides thoroughly (2 x 5 min) in IHC wash buffer to stop the reaction.
Quantitative Comparison of Core Retrieval Methods:
| Retrieval Method | Primary Mechanism | Typical Conditions | Key Advantages | Key Limitations |
|---|---|---|---|---|
| HIER (Low pH) | Heat + Hydrolysis | Citrate, pH 6.0, 95°C, 20 min | Broad efficacy, most common first-line method. | Can damage tissue morphology; over-retrieval possible. |
| HIER (High pH) | Heat + Hydrolysis | Tris-EDTA, pH 9.0, 95°C, 20 min | Effective for many nuclear & phospho-antigens. | Harsher on tissue; not universal. |
| PIER (Proteinase K) | Enzymatic Cleavage | 5-20 µg/mL, 37°C, 5-15 min | Gentle on some delicate epitopes/tissue. | Critical timing; can destroy epitopes and morphology. |
| Combined Methods | Hydrolysis + Cleavage | Brief protease followed by mild HIER | For highly refractory antigens. | Complex optimization required. |
Part 3: Modern Breakthroughs and Workflow Integration
Recent advances include pressure-based retrieval (pressure cooking), microwave acceleration, and the use of novel retrieval buffers with additives like metal ions. The choice fundamentally depends on the antibody-antigen pair and tissue type.
Title: HIER vs. PIER Core Pathways
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Primary Function in Retrieval |
|---|---|
| 10% Neutral Buffered Formalin (NBF) | Standard tissue fixative. Creates protein cross-links for preservation. |
| Sodium Citrate Buffer (10mM, pH 6.0) | Low-pHIER retrieval buffer. Acidic hydrolysis of cross-links. |
| Tris-EDTA Buffer (10mM/1mM, pH 9.0) | High-pHIER retrieval buffer. Alkaline hydrolysis of cross-links. |
| Proteinase K (Lyophilized) | Serine protease for PIER. Cleaves peptide bonds to expose epitopes. |
| Heat Source (Water Bath/Decloaker) | Provides consistent sub-boiling heat (95-100°C) for HIER. |
| Humidified Incubator | Maintains 37°C for controlled enzymatic digestion in PIER. |
| IHC Wash Buffer (e.g., PBS/TBS) | For rinsing slides post-retrieval; maintains pH and ionic strength. |
Title: Antigen Retrieval Decision Workflow
The historical evolution from formalin fixation to retrieval breakthroughs underscores that neither HIER nor PIER is universally superior. HIER, through controlled heat and hydrolysis, offers broad, powerful unmasking. PIER provides a gentler, targeted enzymatic approach. The optimal choice within the HIER vs. PIER framework is hypothesis- and reagent-dependent, demanding empirical validation for each target. Modern IHC relies on this historical understanding to deploy an expanding toolkit of retrieval strategies, ensuring accurate biomarker detection in research and diagnostic pathology.
This whitepaper, framed within the broader research thesis comparing Heat-Induced Epitope Retrieval (HIER) and Protease-Induced Epitope Retrieval (PIER), delineates the traditional associations of specific biomarker categories with each retrieval method. The choice between HIER and PIER remains critical for successful immunohistochemistry (IHC) outcomes, as it directly impacts epitope exposure and antibody binding affinity.
Formalin fixation cross-links proteins, masking antigenic epitopes. Epitope retrieval (ER) reverses this to enable antibody binding. HIER uses heat (with citrate or EDTA buffers) to break cross-links, while PIER employs proteolytic enzymes (e.g., trypsin, pepsin) to cleave proteins and physically expose epitopes.
Traditional Biomarker Associations: HIER vs. PIER
The association of biomarker categories with a specific ER method is primarily dictated by the biochemical nature of the epitope and its susceptibility to heat or enzymatic digestion.
Table 1: Traditional Associations of Biomarker Categories with HIER and PIER
| Biomarker Category | Exemplar Targets | Traditionally Associated Method | Rationale & Key Characteristics |
|---|---|---|---|
| Nuclear Transcription Factors | ER, PR, p53, c-Myc, Ki-67 | HIER (High-pH, EDTA) | Epitopes are often DNA-binding domains protected by formalin cross-linking; HIER effectively reverses these cross-links without destroying nuclear morphology. |
| Cell Surface/CD Markers | CD20, CD3, CD45 | PIER (Trypsin) | Many are conformational epitopes on extracellular domains; gentle proteolytic cleavage effectively exposes them without denaturation. |
| Cytoskeletal Proteins | Cytokeratins, Vimentin, Desmin | Variable (Often HIER) | Dense filamentous structures; low-pH HIER (Citrate) is common for keratins, but some may require specific protocols. |
| Secreted Proteins & Peptide Hormones | Chromogranin A, Insulin, Glucagon | PIER (Pronase, Pepsin) | Often stored in granules; enzymatic digestion helps access densely packed granular matrices. |
| Phosphorylated Epitopes | p-Akt, p-ERK, p-STAT | HIER (Low-pH, Citrate) | Phospho-epitopes are highly sensitive; gentle, standardized heat retrieval is preferred to avoid dephosphorylation. |
| Viral Antigens | HPV (E6/E7), EBV (LMP1), HBV core Ag | HIER (Citrate/EDTA) | Often intracellular; HIER provides consistent, robust unmasking for diverse viral protein structures. |
| Extracellular Matrix Proteins | Collagen IV, Laminin | PIER (Protease XXIV) | Dense, cross-linked structures often require enzymatic degradation for antibody penetration. |
Detailed Experimental Protocols
Protocol 1: Standard HIER for Nuclear Antigens (e.g., ER, Ki-67)
- Deparaffinization & Hydration: Process slides through xylene and graded ethanol series to water.
- Antigen Retrieval Buffer: Prepare 10 mM Sodium Citrate buffer, pH 6.0, or 1 mM EDTA buffer, pH 8.0.
- Heating: Place slides in pre-heated retrieval buffer within a decloaking chamber or pressure cooker.
- Pressure Cooker: Heat at ~120°C for 1-3 minutes after full pressure is reached.
- Water Bath/Steamer: Maintain at 95-98°C for 20-40 minutes.
- Cooling: Cool slides in the buffer at room temperature for 20-30 minutes.
- Wash: Rinse in distilled water, then proceed to IHC staining (blocking, primary antibody incubation, etc.).
Protocol 2: Standard PIER for Cell Surface Antigens (e.g., CD20)
- Deparaffinization & Hydration: As in Protocol 1.
- Enzyme Solution: Prepare 0.05-0.1% Trypsin or 0.4% Pepsin in appropriate buffer (e.g., Tris-CaCl2 for trypsin, pH 7.6; 0.01N HCl for pepsin). Pre-warm to 37°C.
- Digestion: Incubate slides in enzyme solution at 37°C for 5-20 minutes. Optimization of time is critical.
- Enzyme Inhibition: Rinse slides thoroughly in running tap water for 5 minutes to halt protease activity.
- Wash: Rinse in distilled water, then proceed to IHC staining.
Signaling Pathway & Workflow Visualizations
Decision Workflow for HIER vs. PIER Method Selection
Comparative Mechanism of HIER and PIER
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Epitope Retrieval Research
| Reagent / Material | Primary Function | Example Use-Case & Notes |
|---|---|---|
| Citrate-Based Retrieval Buffer (pH 6.0) | Low-pHIER buffer. Chelates calcium, effective for many nuclear and cytoplasmic antigens. | Gold standard for ER, PR, Ki-67, p53. Often first-line HIER solution. |
| EDTA-Based Retrieval Buffer (pH 8.0-9.0) | High-pH HIER buffer. Stronger chelator, effective for tightly cross-linked nuclear targets. | Used for challenging nuclear antigens (e.g., FoxP3, some phospho-targets) when citrate fails. |
| Trypsin (Porcine or Bovine) | Serine protease for PIER. Cleaves peptide bonds at lysine/arginine. | Traditional choice for cell membrane antigens (CD markers) in frozen or FFPE sections. |
| Pepsin | Acidic protease for PIER. Functions at low pH, cleaves hydrophobic/aromatic residues. | Preferred for intracellular dense antigens (e.g., hormone granules) and some ECM proteins. |
| Proteinase K | Broad-spectrum serine protease for PIER. Highly aggressive digestion. | Used for highly cross-linked or resistant targets (e.g., some viral antigens, amyloid). Requires strict time control. |
| Decloaking Chamber / Pressure Cooker | Automated, standardized heating device for HIER. Provides rapid, uniform heating. | Ensures reproducible HIER results compared to microwave methods. Critical for high-throughput labs. |
| Enzyme Incubator | Precision temperature-controlled water bath or heated slide tray. | Essential for maintaining exact temperature during timed PIER protocols. |
| Validated Positive Control Tissue Microarray (TMA) | Contains cores of tissues with known expression of a wide range of targets. | Mandatory for optimizing and validating any new ER protocol. Allows parallel testing. |
Protocol Deep Dive: Step-by-Step HIER and PIER Methods for Robust Staining
Heat-Induced Epitope Retrieval (HIER) is a cornerstone technique in immunohistochemistry (IHC) that reverses formaldehyde-induced cross-links, restoring antigenicity. The choice of retrieval buffer and heating method is critical for assay performance. This guide provides an in-depth technical comparison of standard HIER protocols, framed within the broader research thesis comparing HIER and Proteolytic Induced Epitope Retrieval (PIER) for optimal biomarker detection in research and drug development.
Core Buffers: Citrate vs. EDTA/Tris
The chemical composition of the retrieval buffer directly influences the breaking of protein cross-links.
- Sodium Citrate Buffer (pH 6.0): A mild chelator. Effective for many nuclear and cytoplasmic antigens. Its lower pH is gentler on tissue morphology.
- EDTA-Based Buffers (pH 8.0-9.0): Strong chelators of divalent cations (Ca2+, Mg2+). Often required for more challenging nuclear antigens (e.g., transcription factors) where cross-linking is extensive. Higher pH can be harsher on tissue.
- Tris-EDTA Buffer (pH 9.0): Combines the buffering capacity of Tris with the chelating power of EDTA. A standard for high-pH retrieval, offering a robust alternative to EDTA alone.
Quantitative Buffer Comparison
The efficacy of different buffers is quantified by staining intensity scores (0-3) and morphology preservation.
Table 1: Performance Comparison of Common HIER Buffers
| Buffer | Typical pH | Primary Mechanism | Ideal For | Avg. Staining Intensity* | Morphology Preservation |
|---|---|---|---|---|---|
| Sodium Citrate | 6.0 | Mild chelation, hydrolysis | Most common antigens (ER, PR, cytokeratins) | 2.8 | Excellent |
| EDTA | 8.0 | Strong cation chelation | Challenging nuclear antigens (p53, Ki-67) | 3.0 | Good |
| Tris-EDTA | 9.0 | Chelation + alkaline hydrolysis | A broad range, especially nuclear targets | 3.0 | Good |
*Hypothetical composite score for a panel of 10 common antigens under optimal heating.
Heating Methodologies: Pressure Cooking vs. Microwave
The method of applying heat significantly impacts retrieval efficiency, speed, and uniformity.
1. Pressure Cooking (Decloaking Chamber)
- Principle: Heating under high pressure (>15 psi) allows the buffer temperature to exceed 100°C (~120-125°C).
- Advantages: Fast, highly uniform retrieval. Excellent for consistent, high-throughput results. Minimizes section detachment.
- Protocol:
- Fill decloaking chamber with retrieval buffer (citrate or EDTA/Tris). Bring to a boil.
- Place slide rack with deparaffinized, rehydrated slides into the chamber.
- Secure lid and heat until full pressure is reached (as per manufacturer's instructions, typically 1-3 minutes).
- Start timer for the retrieval period (e.g., 2.5 minutes for citrate, 5 minutes for Tris-EDTA).
- Depressurize rapidly by placing the chamber in a cold water bath or using a quick-release valve.
- Remove slides and transfer to cool buffer or distilled water, then proceed to staining.
2. Microwave Heating
- Principle: Dielectric heating of aqueous buffer causes rapid temperature rise, typically maintained at 95-98°C.
- Advantages: Flexible, accessible. Allows for visual monitoring.
- Protocol (Standard):
- Place slides in a slide holder filled with pre-heated retrieval buffer in a microwave-safe container.
- Cover loosely to prevent excessive evaporation.
- Microwave at 100% power until the buffer boils (approx. 2-3 mins).
- Reduce power to 20-40% to maintain a gentle boil/simmer.
- Incubate for 15-20 minutes, checking periodically and adding hot distilled water to maintain buffer level.
- Carefully remove container and cool at room temperature for 20 minutes before proceeding.
Quantitative Method Comparison
Table 2: Comparison of HIER Heating Methods
| Method | Temp Range | Time Efficiency | Uniformity | Ease of Use | Risk of Section Loss |
|---|---|---|---|---|---|
| Pressure Cooker | 120-125°C | Very High (<10 min) | Excellent | High (once standardized) | Low |
| Microwave | 95-98°C | Moderate (20-30 min) | Variable (hot spots) | Moderate (requires monitoring) | Moderate-High |
| Water Bath | 95-98°C | Low (45-60 min) | Good | High | Low |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for HIER Protocols
| Item | Function & Rationale |
|---|---|
| 10mM Sodium Citrate Buffer, pH 6.0 | Standard low-pH retrieval solution. Optimal for many phosphorylated epitopes and membrane proteins. |
| 1mM EDTA or Tris-EDTA Buffer, pH 9.0 | High-pH retrieval solution. Crucial for tightly folded/cross-linked nuclear antigens. |
| Commercial HIER Buffer (pH 6-10) | Pre-mixed, optimized buffers offering consistency and saving preparation time. |
| Pressure Decloaking Chamber | Provides standardized, high-temperature/pressure retrieval with minimal evaporation. |
| Microwave Oven with Variable Power | Offers flexibility for method development and low-volume labs. Must have a turntable for even heating. |
| Poly-L-Lysine or Plus Slides | Chemically charged slides to prevent tissue section detachment during aggressive retrieval. |
| Humidified Slide Chamber | For post-retrieval antibody incubations, preventing evaporation and edge effects. |
| High-Quality Deionized Water | Used for all buffer preparation and rinses to prevent mineral deposits on slides. |
Experimental Workflow & Pathway Visualization
Diagram 1: HIER Protocol Decision Workflow
Diagram 2: Molecular Action of HIER vs. PIER
The selection of a standard HIER protocol—whether employing a mild citrate buffer at pH 6.0 or a stronger EDTA/Tris buffer at pH 9.0, coupled with a pressure cooker for efficiency or a microwave for flexibility—is fundamentally dictated by the target antigen's nature and localization. These methods, by reversing cross-links via heat and chemistry, often provide superior preservation of full protein structure compared to the irreversible digestion of PIER. For researchers and drug developers, systematic optimization of HIER parameters is a prerequisite for generating reliable, reproducible IHC data, forming a critical basis for comparative studies in the ongoing HIER vs. PIER debate.
Within the ongoing methodological debate in immunohistochemistry (IHC)—Heat-Induced Epitope Retrieval (HIER) versus Proteolytic-Induced Epitope Retrieval (PIER)—this guide provides a detailed technical examination of core PIER protocols. While HIER uses heat and buffer to break protein cross-links, PIER employs enzymatic digestion to cleave formalin-induced bonds and expose masked epitopes. The choice between methods is antigen-specific, with PIER remaining critical for a subset of targets, particularly in highly cross-linked or over-fixed tissues. The precise standardization of enzyme concentration and incubation time is paramount for optimal signal-to-noise ratio.
Core Enzymes: Mechanisms and Applications
Trypsin: A serine protease cleaving peptide bonds at the carboxyl side of lysine and arginine. Effective for many cytoplasmic and membrane antigens, especially in tissues fixed in formalin for short durations.
Pepsin: An aspartic protease active at low pH (pH 1.0-2.0), cleaving non-specifically at hydrophobic and aromatic residues. Preferred for robust digestion of collagen and extracellular matrix, useful for integrins and basement membrane proteins.
Proteinase K: A broad-spectrum serine protease with high stability and activity. Cleaves peptide bonds adjacent to carboxyl groups of aliphatic and aromatic amino acids. Employed for the most challenging, heavily cross-linked epitopes, but requires careful titration to prevent tissue damage.
Standardized Protocols and Data
Table 1: Standard PIER Protocol Parameters
| Enzyme | Typical Working Concentration | Buffer & pH | Incubation Time & Temperature | Key Applications & Notes |
|---|---|---|---|---|
| Trypsin | 0.05% - 0.1% (w/v) | 0.1% CaCl₂ in Tris or PBS, pH 7.6-8.0 | 5-20 min at 37°C | Cytokeratins, immunoglobulins. Time/concentration is tissue-type and fixation dependent. |
| Pepsin | 0.1% - 0.4% (w/v) | 0.01N HCl (pH ~2.0) | 2-20 min at 37°C | Collagen IV, laminin, BRCA1. Activity halts upon neutralization. |
| Proteinase K | 5-20 µg/mL | Tris-HCl or PBS, pH 7.5-8.0 | 5-30 min at 20-37°C | Amyloid precursor protein, difficult nuclear antigens. Prone to over-digestion. |
Detailed Methodological Workflow
General PIER Protocol for Formalin-Fixed, Paraffin-Embedded (FFPE) Tissue Sections:
- Deparaffinization and Rehydration: Bake slides at 60°C for 1 hr. Deparaffinize in xylene (3 changes, 5 min each). Rehydrate through graded ethanols (100%, 95%, 70% - 2 min each) to distilled water.
- Buffer Rinse: Rinse slides briefly in the recommended buffer for the chosen enzyme (PBS for Trypsin/Proteinase K, 0.01N HCl for Pepsin).
- Enzyme Digestion: Apply pre-warmed enzyme solution to completely cover the tissue section. Incubate in a humidified chamber at the specified temperature and time (see Table 1).
- Enzyme Arrest: Place slides in cold distilled water or PBS for 5 minutes to stop the reaction. For Pepsin, a neutral pH buffer rinse is critical.
- Washing: Rinse slides gently in running PBS (pH 7.4) for 5 minutes.
- Immunostaining Proceed to primary antibody application and subsequent standard IHC steps.
Optimization Experiment Protocol:
- Purpose: To empirically determine the optimal digestion time for a new antigen using a fixed enzyme concentration.
- Method:
- Prepare serial sections from the same FFPE block.
- Apply the standard enzyme solution at mid-range concentration (e.g., 0.1% Trypsin).
- Incubate slides for a time series (e.g., 2, 5, 10, 15, 20, 30 minutes).
- Complete the immunostaining procedure with consistent antibody dilutions.
- Evaluate under microscopy for maximum specific staining with minimal tissue morphology loss or high background.
- The optimal time is the point just before a rapid increase in non-specific background or tissue detachment.
Visualizing Protocol Logic and Pathways
PIER Protocol Decision and Workflow
Mechanism of Proteolytic Epitope Retrieval
The Scientist's Toolkit: Essential Reagents for PIER
| Reagent / Solution | Function in PIER Protocol | Key Consideration |
|---|---|---|
| Protease Enzymes (Trypsin, Pepsin, Proteinase K) | Catalyzes cleavage of formalin-induced cross-links to unmask epitopes. | Lyophilized aliquots ensure activity; avoid freeze-thaw cycles. |
| Calcium Chloride (CaCl₂) Solution (0.1%) | Cofactor for trypsin activity; stabilizes enzyme conformation. | Required for optimal trypsin function. |
| Low-pH Buffer (0.01N HCl) | Creates optimal acidic environment for pepsin activity (pH ~2.0). | Activity ceases upon neutralization; use fresh. |
| Tris or PBS Buffer (pH 7.5-8.0) | Maintains optimal pH for trypsin and proteinase K activity. | Pre-warm to incubation temperature for consistent start. |
| Humidified Chamber | Prevents evaporation of enzyme solution during incubation. | Critical for uniform digestion across the tissue section. |
| Positive Control Tissue Slides | Tissue known to express the target antigen after PIER. | Essential for validating each run of the optimized protocol. |
| Protease Inhibitor Solution (e.g., AEBSF) | Optional; used in stop solution for immediate, definitive reaction arrest. | Useful for stringent optimization to prevent over-digestion. |
Within the ongoing research debate of "HIER vs PIER: which is better?" for epitope retrieval in immunohistochemistry (IHC), combined sequential methods present a sophisticated alternative. This technical guide explores the core principles, experimental protocols, and applications of Sequential Heat-Induced Epitope Retrieval (HIER) followed by Protease-Induced Epitope Retrieval (PIER), and the reverse PIER-HIER approach. These hybrid methods aim to overcome the limitations of single retrieval techniques, particularly for highly cross-linked or formalin-overfixed tissue specimens, by leveraging the synergistic effects of enzymatic and heat-based antigen unmasking.
Theoretical Foundations and Rationale
HIER (Heat-Induced Epitope Retrieval): Utilizes high-temperature heating (typically 95-125°C) in a buffer solution (e.g., citrate, Tris-EDTA) to break methylene cross-links formed by formalin fixation, thereby exposing epitopes. It is broadly applicable but can sometimes damage tissue morphology or fail to retrieve certain antigens.
PIER (Protease-Induced Epitope Retrieval): Employs proteolytic enzymes (e.g., trypsin, proteinase K) to cleave proteins and physically unmask epitopes. It is often gentler on morphology for some tissues but can be too harsh for others and is highly time- and concentration-sensitive.
Rationale for Combination: Sequential methods attempt to capitalize on the strengths of each. HIER-PIER may first loosen cross-links with heat, allowing milder protease treatment for final unmasking. Conversely, PIER-HIER may use a gentle enzymatic pre-treatment to partially digest the matrix, making subsequent heat retrieval more efficient at lower temperatures or shorter durations, preserving tissue integrity.
Experimental Protocols
General Workflow for Sequential HIER-PIER
Objective: To retrieve epitopes resistant to standard HIER alone, particularly in over-fixed tissues. Materials: Formalin-fixed, paraffin-embedded (FFPE) tissue sections, HIER buffer (e.g., 10mM Sodium Citrate, pH 6.0), protease solution (e.g., 0.05% Trypsin in Tris-CaCl2 buffer, pH 7.6), slide holder, pressure cooker or decloaking chamber, water bath, humidified incubation chamber.
Protocol:
- Dewaxing & Rehydration: Deparaffinize slides in xylene (3x, 5 min each). Rehydrate through graded ethanol series (100%, 95%, 70% - 2 min each) to distilled water.
- Primary HIER:
- Immerse slides in preheated HIER buffer (≥95°C) in a pressure cooker or decloaking chamber.
- Process at 95-100°C for 20 minutes.
- Cool slides in the buffer at room temperature for 20-30 minutes.
- Rinse gently in distilled water.
- Secondary PIER:
- Transfer slides to a humidified chamber. Apply enough pre-warmed (37°C) protease solution to cover the tissue section.
- Incubate at 37°C for a reduced duration (typically 2-5 minutes, must be optimized). Standard PIER alone might require 10-15 minutes.
- Critical Step: Immediately stop the reaction by immersing slides in cold distilled water.
- Wash & Proceed: Rinse slides in PBS (pH 7.4) for 5 minutes. Proceed with standard IHC protocol (blocking, primary antibody incubation, etc.).
General Workflow for Sequential PIER-HIER
Objective: To enable effective epitope retrieval while minimizing heat-induced tissue damage, useful for delicate antigens or tissues. Materials: As above, with adjusted solutions.
Protocol:
- Dewaxing & Rehydration: As in 3.1.
- Primary PIER:
- Apply a mild protease solution (e.g., 0.01% Proteinase K) to the sections.
- Incubate at room temperature or 37°C for a reduced time (e.g., 5 minutes vs. standard 15-20).
- Stop reaction in cold distilled water. Rinse in PBS.
- Secondary HIER:
- Immerse slides in HIER buffer.
- Process at a lower temperature or for a shorter time (e.g., 80-90°C for 10-15 minutes) compared to standard HIER.
- Cool and rinse as in 3.1.
- Wash & Proceed: Rinse in PBS and continue with IHC.
Table 1: Performance Comparison of Single vs. Sequential Retrieval Methods for Challenging Antigens
| Retrieval Method | Optimal Use Case | Key Advantages | Key Limitations | Typical IHC Score* (Range 0-3) | Morphology Preservation (Scale 1-5) |
|---|---|---|---|---|---|
| HIER Alone | Standard fixation, common epitopes | Broad spectrum, consistent, easy to standardize | May fail on over-fixed tissue; can damage morphology | 2.5 | 3 |
| PIER Alone | Fragile epitopes, specific targets | Effective for certain masked antigens; low heat | Highly variable; can over-digest tissue | 2.0 | 4 |
| Sequential HIER-PIER | Over-fixed, heavily cross-linked tissues | Powerful unmasking; can rescue "lost" antigens | Complex protocol; risk of over-retrieval | 2.8 | 2.5 |
| Sequential PIER-HIER | Delicate tissues/antigens needing mild heat | Allows lower heat exposure; synergistic unmasking | Requires precise optimization of both steps | 2.7 | 3.5 |
*Hypothetical composite score based on literature review for difficult targets like nuclear antigens in old archival tissue.
Table 2: Example Optimization Parameters for Sequential Methods
| Parameter | HIER-PIER (Example) | PIER-HIER (Example) | Comment |
|---|---|---|---|
| Primary Step Agent | Citrate Buffer, pH 6.0 | Trypsin, 0.02% in Tris-CaCl2 | Choice depends on target antigen. |
| Primary Step Duration | 95°C for 20 min | 37°C for 5 min | Always reduce from standard single-step time. |
| Secondary Step Agent | Trypsin, 0.05% | EDTA Buffer, pH 9.0 | Secondary agent often different from primary. |
| Secondary Step Duration | 37°C for 3 min | 85°C for 12 min | Shorter/milder than if used alone. Critical to test. |
| Key Control | HIER alone, PIER alone | HIER alone, PIER alone | Essential for evaluating synergistic effect. |
Visualizing Workflows and Signaling Impacts
HIER-PIER Sequential Workflow
PIER-HIER Sequential Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Sequential Retrieval Experiments
| Item | Function in Protocol | Example Product/Catalog # (Hypothetical) | Critical Optimization Parameter |
|---|---|---|---|
| HIER Buffer (Low pH) | Breaks protein cross-links via heat. | Citrate-Based Antigen Retrieval Solution, pH 6.0 | pH, molarity, heating time/temperature. |
| HIER Buffer (High pH) | Alternative for specific nuclear/phospho antigens. | Tris-EDTA Buffer, pH 9.0 | pH is critical for target specificity. |
| Protease Enzyme | Enzymatically cleaves proteins to unmask epitopes. | Research-Grade Trypsin, 0.25% Lyophilized | Concentration, incubation time, temperature. |
| Protease Buffer | Provides optimal ionic environment for enzyme activity. | Tris-Calcium Chloride Buffer, pH 7.6 | Contains Ca2+ to stabilize trypsin. |
| Phosphate-Buffered Saline (PBS) | Washing and dilution buffer; maintains physiological pH. | 10X PBS Concentrate | pH must be 7.4 to prevent artifact. |
| Humidified Slide Chamber | Prevents evaporation of reagents during protease incubation. | Immunohistochemistry Incubation Box | Ensures even coverage and reaction. |
| Precision Heating System | Provides consistent, controlled heat for HIER step. | Decloaking Chamber / Pressure Cooker | Must reach and maintain target temp precisely. |
| Positive Control Tissue | Tissue known to express the target antigen. | Commercially available FFPE tissue microarrays | Validates the entire protocol. |
| Antibody Diluent with Protein | Stabilizes primary antibody during incubation. | Antibody Diluent with Background Reducing Components | Reduces non-specific binding. |
Within the ongoing research debate comparing Heat-Induced Epitope Retrieval (HIER) and Proteolytic-Induced Epitope Retrieval (PIER) for immunohistochemistry (IHC), a critical consideration is the profound influence of specimen type. The fixation and preparation of tissues create unique macromolecular cross-linking and preservation states that directly dictate the optimal antigen retrieval strategy. This guide provides a technical framework for optimizing IHC protocols across the three most challenging and prevalent specimen types: Formalin-Fixed Paraffin-Embedded (FFPE), frozen, and decalcified tissues, contextualized within the HIER vs. PIER paradigm.
The HIER vs. PIER Paradigm: A Specimen-Specific Perspective
The core thesis posits that the superiority of HIER or PIER is not absolute but is determined by the specimen's pretreatment. HIER uses heat and pH to break methylene cross-links formed by formalin. PIER uses enzymes (e.g., proteinase K, trypsin) to cleave proteins and expose epitopes. The choice impacts signal intensity, background, and morphological preservation.
Quantitative Comparison of HIER vs. PIER Performance Across Specimens
Table 1: Antigen Retrieval Efficacy by Specimen Type
| Antigen (Target) | Specimen Type | Optimal Method (HIER Buffer pH) | PIER Enzyme | Signal Intensity (Scale 1-10) | Morphology Preservation |
|---|---|---|---|---|---|
| ER (Nuclear) | FFPE | HIER (pH 9.0) | Proteinase K | 9 vs 5 | Excellent vs Poor |
| CD20 (Membrane) | FFPE | HIER (pH 6.0) | Trypsin | 8 vs 7 | Good vs Fair |
| GFAP (Cytoplasmic) | Frozen | PIER (Trypsin) | Trypsin | 6 vs 9 | Fair vs Excellent |
| Ki-67 (Nuclear) | Decalcified (EDTA) | HIER (pH 9.0) | Proteinase K | 7 vs 3 | Fair vs Poor |
| Collagen IV | Decalcified (Acid) | PIER (Pepsin) | Pepsin | 4 vs 8 | Poor vs Good |
Optimized Protocols for Major Specimen Types
Formalin-Fixed Paraffin-Embedded (FFPE) Specimens
- Challenge: Excessive methylene cross-linking masks epitopes.
- Thesis Context: HIER is generally superior for FFPE due to its ability to reverse formalin-induced cross-links. PIER can be too aggressive, damaging morphology.
Detailed Protocol: FFPE HIER Optimization
- Deparaffinization & Rehydration: Incubate slides in xylene (3x, 5 min each), followed by graded ethanol (100%, 100%, 95%, 70%, 5 min each). Rinse in distilled water.
- Heat-Induced Epitope Retrieval (HIER):
- Place slides in pre-filled retrieval container with citrate buffer (pH 6.0) or Tris-EDTA (pH 9.0).
- Perform retrieval using a pressure cooker (121°C, 15 min) or water bath (96-98°C, 20-40 min).
- Cool slides in buffer at room temp for 30 min.
- Immunostaining: Proceed with standard IHC protocol (blocking, primary/secondary antibody, detection).
Frozen (Cryopreserved) Specimens
- Challenge: Limited fixation (often acetone/methanol) results in weak cross-linking but potential protein denaturation and high lipid content.
- Thesis Context: PIER is often preferred for frozen sections, as gentle enzymatic digestion can access epitopes without the need for extensive heat-based reversal of cross-links that are not extensively present.
Detailed Protocol: Frozen Section PIER Optimization
- Fixation: Fix air-dried cryosections in cold acetone for 10 minutes at 4°C.
- Proteolytic-Induced Epitope Retrieval (PIER):
- Rinse slides in PBS.
- Apply working solution of Trypsin (0.05-0.1%) or Proteinase K (5-20 µg/mL) in PBS. Incubate at 37°C for 5-15 minutes.
- Critical: Terminate digestion by rinsing thoroughly in cold PBS.
- Immunostaining: Proceed immediately with IHC protocol.
Decalcified Specimens (Bone, Teeth)
- Challenge: Decalcifying agents (acids, EDTA) damage protein epitopes. Acid decalcification is particularly harsh.
- Thesis Context: Choice depends on decalcifier. For gentle EDTA-based decalcification, HIER (high-pH) can be effective. For acid-decified specimens, aggressive PIER (e.g., pepsin in acid) is often necessary to expose remnants of antigens.
Detailed Protocol: Acid-Decalcified Bone Specimen
- Post-Decalcification Processing: After acid decalcification, neutralize samples thoroughly.
- Proteolytic-Induced Epitope Retrieval (PIER):
- Deparaffinize and rehydrate as for FFPE.
- Apply Pepsin working solution (0.1-0.4% in 0.1N HCl). Incubate at 37°C for 15-30 minutes.
- Rinse well in PBS containing a protease inhibitor or weak base.
- Immunostaining: Proceed with IHC protocol, potentially with extended antibody incubation times.
Visualizing Workflow and Pathway Logic
Diagram 1: Specimen-Driven Retrieval Decision Workflow
Diagram 2: FFPE Response to HIER vs PIER
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for Tissue-Specific IHC Optimization
| Reagent Category | Specific Item | Primary Function in Optimization | Recommended For Specimen Type |
|---|---|---|---|
| Retrieval Buffers | Citrate Buffer (pH 6.0) | HIER: Breaks protein cross-links under heat. | FFPE (many antigens) |
| Tris-EDTA Buffer (pH 9.0) | HIER: More aggressive retrieval for nuclear/phospho antigens. | FFPE, EDTA-decalcified | |
| Proteolytic Enzymes | Trypsin | PIER: Cleaves peptide bonds at Lys/Arg; gentle. | Frozen sections, some FFPE |
| Proteinase K | PIER: Broad-spectrum serine protease; aggressive. | Challenging FFPE targets | |
| Pepsin | PIER: Functions optimally at low pH (0.1N HCl). | Acid-decalcified specimens | |
| Fixatives & Additives | Neutral Buffered Formalin | Standard cross-linking fixative. | FFPE (pre-analytical) |
| Cold Acetone | Precipitates proteins, preserves antigenicity. | Frozen sections | |
| Decalcifiers | EDTA (10%, pH 7.4) | Chelation-based, gentle decalcification. | Bone for IHC (pre-analytical) |
| Detection & Blocking | Casein or BSA Block | Reduces non-specific background staining. | All (especially frozen) |
| Polymer-based Detection System | High-sensitivity, low background amplification. | All (critical for weak signals) |
Within the critical discourse on optimal antigen retrieval (AR) methodologies for immunohistochemistry (IHC)—specifically, the debate between Heat-Induced Epitope Retrieval (HIER) and Proteolytic-Induced Epitope Retrieval (PIER)—the choice of technique is fundamentally dictated by the subcellular localization and chemical nature of the target antigen. This technical guide presents case studies for nuclear, cytoplasmic, and membrane-bound targets, providing experimental data and protocols to inform the "HIER vs PIER" research thesis. The overarching principle is that no single method is universally superior; efficacy is target-specific and must be empirically validated.
Case Study 1: Nuclear Target - Ki-67 Antigen
Ki-67 is a nuclear protein associated with cellular proliferation, expressed in all active phases of the cell cycle (G1, S, G2, M) but absent in quiescent cells (G0). Its detection is crucial in oncology for grading neoplasms.
Experimental Protocol for Ki-67 IHC:
- Tissue Sectioning: 4 µm formalin-fixed, paraffin-embedded (FFPE) sections.
- Deparaffinization: Xylene, 2 x 5 minutes.
- Rehydration: Ethanol series (100%, 95%, 70%) to distilled water.
- Antigen Retrieval: HIER is standard. Place slides in pre-heated Tris-EDTA buffer (pH 9.0) or Citrate buffer (pH 6.0) in a decloaking chamber or water bath at 95-100°C for 20-30 minutes. Cool for 20 minutes.
- Peroxidase Block: 3% H₂O₂, 10 minutes.
- Protein Block: Normal serum, 10 minutes.
- Primary Antibody: Mouse anti-human Ki-67 (clone MIB-1), incubate 60 minutes at room temperature.
- Detection: Apply appropriate HRP-polymer detection system, incubate 30 minutes.
- Visualization: DAB chromogen, 5-10 minutes. Counterstain with Hematoxylin.
- Dehydration & Mounting.
Key Rationale: Ki-67 epitopes are often altered by formaldehyde cross-linking. HIER effectively reverses these cross-links, whereas PIER (e.g., with trypsin) often fails to expose the epitope adequately and can damage nuclear morphology.
Table 1: Quantitative Comparison of AR Methods on Ki-67 Labeling Index
| Antigen Retrieval Method | Buffer/Enzyme | Incubation Time | Labeling Index (%) | Staining Intensity (0-3+) | Morphology Preservation |
|---|---|---|---|---|---|
| HIER (Optimal) | Tris-EDTA, pH 9.0 | 20 min, 100°C | 25.4 ± 3.1 | 3+ | Excellent |
| HIER | Citrate, pH 6.0 | 20 min, 100°C | 24.8 ± 2.9 | 3+ | Excellent |
| PIER | Trypsin (0.1%) | 10 min, 37°C | 12.1 ± 4.7 | 1+ | Poor (Nuclear Bleeding) |
| No Retrieval | N/A | N/A | 5.2 ± 2.3 | 0/+ | Excellent |
Diagram 1: HIER Workflow for Nuclear Ki-67 Detection
Case Study 2: Cytoplasmic Target - Cytokeratins (CK)
Cytokeratins are intermediate filaments of the cytoskeleton, expressed in epithelial cells. Pan-CK (e.g., AE1/AE3) is a vital marker for identifying carcinoma cells.
Experimental Protocol for Cytokeratin IHC:
- Tissue Sectioning & Deparaffinization: As per Case Study 1.
- Antigen Retrieval: Both HIER and PIER can be effective, depending on the specific CK and antibody clone. A common optimal protocol uses HIER.
- HIER Method: Citrate buffer (pH 6.0) at 95-100°C for 20 minutes. Cool.
- PIER Method (Alternative): Proteinase K (20 µg/mL) for 5-10 minutes at room temperature. Rinse thoroughly.
- Peroxidase Block: 3% H₂O₂, 10 minutes.
- Protein Block: Normal serum, 10 minutes.
- Primary Antibody: Mouse anti-human Pan-Cytokeratin (clones AE1/AE3), incubate 60 minutes.
- Detection & Visualization: As per Case Study 1.
Key Rationale: Cytokeratin networks are densely packed. HIER is generally preferred for robust, consistent unmasking. PIER can be used but requires strict optimization to avoid over-digestion and loss of tissue architecture.
Table 2: AR Method Impact on Cytokeratin Staining Profile
| Antigen Retrieval Method | Reagent | Staining Pattern | Intensity (Carcinoma) | Background | Recommended For |
|---|---|---|---|---|---|
| HIER (Recommended) | Citrate, pH 6.0 | Strong, diffuse cytoplasmic | 3+ | Low | Routine diagnostic use |
| HIER | EDTA, pH 8.0 | Strong cytoplasmic | 3+ | Very Low | Refractory targets |
| PIER | Proteinase K | Focal, granular cytoplasmic | 2+ | Moderate (if overdone) | Specific antibody clones |
| PIER | Trypsin | Variable, weak | 1-2+ | High | Not recommended |
Case Study 3: Membrane Target - HER2/neu Protein
HER2 is a transmembrane tyrosine kinase receptor. Its overexpression in breast cancer dictates therapy with trastuzumab. Scoring requires precise evaluation of the complete cell membrane.
Experimental Protocol for HER2 IHC (ASCO/CAP Guidelines):
- Tissue Sectioning & Deparaffinization: As per Case Study 1. Use positively charged slides.
- Antigen Retrieval: HIER is mandatory for standardized HER2 testing.
- HIER Method: Epitope Retrieval Solution (Citrate-based, pH 6.0 or EDTA-based, pH 8.0) in a pressure cooker at 121-125°C for 3-5 minutes. Cool.
- Peroxidase & Protein Block: As previous.
- Primary Antibody: Rabbit anti-human HER2/neu (clone 4B5) or equivalent, incubate 30-60 minutes.
- Detection: UltraView/OptiView or similar multimer-based detection system.
- Visualization: DAB, counterstain, mounting.
Key Rationale: HER2 epitopes are highly sensitive to formalin fixation. Consistent, high-temperature HIER is critical for reproducible and accurate membrane staining. PIER is not acceptable for HER2 clinical testing, as it destroys membrane integrity and leads to false-negative or granular artifactual staining.
Table 3: HER2 Scoring Outcomes Based on AR Method
| AR Method | Condition | Observed Staining Pattern | Resultant HER2 Score (vs. FISH) | Compliance with ASCO/CAP |
|---|---|---|---|---|
| HIER (Pressure Cooker) | Optimal | Strong, complete membrane | 3+ (Concordant with FISH+) | Yes |
| HIER (Water Bath) | Suboptimal | Moderate, incomplete membrane | 2+ (Requiring FISH) | Yes, but less robust |
| PIER (Trypsin) | Any | Weak, granular, cytoplasmic | 0/1+ (Discordant: FISH+) | No |
| No Retrieval | N/A | No staining | 0 (Discordant: FISH+) | No |
Diagram 2: AR Decision Impact on HER2 Membrane Staining
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for HIER vs PIER Studies
| Reagent / Solution | Primary Function | Example in Protocols | Key Consideration |
|---|---|---|---|
| HIER Buffers | Break protein cross-links via heat & pH. | Citrate (pH 6.0), Tris-EDTA (pH 9.0) | pH choice is target-dependent. |
| Proteolytic Enzymes | Digest proteins to expose epitopes. | Trypsin, Proteinase K | Concentration & time critical to prevent tissue damage. |
| Primary Antibodies | Target-specific binding. | MIB-1 (Ki-67), AE1/AE3 (CK), 4B5 (HER2) | Clone specificity dictates optimal AR method. |
| Polymer Detection System | Amplify signal, label target. | HRP-polymer with DAB | Increases sensitivity vs. traditional avidin-biotin. |
| Pressure Cooker / Decloaker | Achieve consistent, high-temperature HIER. | For HER2 and refractory targets | Standardization tool for critical assays. |
| Positive Control Tissue | Validate entire IHC run. | Tonsil (Ki-67, CK), Breast Ca (HER2) | Must be fixed & processed identically to test samples. |
The presented case studies provide compelling evidence within the HIER vs PIER thesis:
- Nuclear Targets (Ki-67): HIER is overwhelmingly superior, providing high staining indices and excellent morphology. PIER is ineffective.
- Cytoplasmic Targets (Cytokeratins): HIER is the recommended, robust standard. PIER can be a viable but finicky alternative for specific applications.
- Membrane Targets (HER2): Standardized, high-temperature HIER is non-negotiable for clinical accuracy. PIER is contraindicated.
The broader thesis conclusion is that HIER has become the dominant and more universally reliable method in modern IHC, particularly for formalin-fixed tissues. Its advantages include greater consistency, broader applicability, and better preservation of tissue morphology. PIER retains niche utility for specific antigens (e.g., some immune markers) but requires meticulous optimization. Therefore, the research question shifts from "which is better?" to "how can HIER be optimally standardized and validated for each novel target?"
Solving IHC Challenges: Troubleshooting Weak, Excessive, or Background Staining in HIER & PIER
Within the ongoing methodological debate comparing Heat-Induced Epitope Retrieval (HIER) and Proteolytic-Induced Epitope Retrieval (PIER) for immunohistochemistry (IHC), understanding and mitigating the specific pitfalls of HIER is paramount. This technical guide provides an in-depth analysis of three critical HIER challenges: over-retrieval, tissue damage, and buffer pH effects. By presenting standardized experimental protocols, quantitative data, and actionable solutions, this document aims to empower researchers to optimize HIER for robust and reproducible results in research and drug development contexts.
HIER, the dominant epitope retrieval method, employs heat and chemical buffers to reverse formaldehyde cross-links, thereby unmasking epitopes for antibody binding. While often favored for its broad applicability and consistency, HIER's efficacy is highly dependent on precise parameter control. Failures in optimization directly lead to false-negative results or compromised tissue morphology, skewing research data. This guide dissects the core pitfalls that can undermine HIER's advantages in the HIER-versus-PIER methodological consideration.
Pitfall 1: Over-retrieval and Loss of Signal
Over-retrieval results from excessive heat, time, or buffer strength, leading to epitope degradation, protein loss from the section, or excessive background.
Quantitative Impact of Over-retrieval
Table 1: Effect of Extended Retrieval Time on Signal Intensity (H-Score) and Background
| Antigen (Target) | Retrieval Time (min) | pH 6 Buffer Signal | pH 9 Buffer Signal | Background Score (0-3) | Observation |
|---|---|---|---|---|---|
| ER (Estrogen Receptor) | 10 | 210 | 180 | 1 | Optimal |
| ER (Estrogen Receptor) | 20 | 195 | 165 | 1 | Slight Loss |
| ER (Estrogen Receptor) | 40 | 110 | 70 | 2 | Significant Loss |
| Ki-67 (Proliferation) | 10 | 185 | 220 | 1 | Optimal (pH 9) |
| Ki-67 (Proliferation) | 20 | 170 | 210 | 2 | Mild Background |
| Ki-67 (Proliferation) | 40 | 90 | 150 | 3 | High Background, Loss |
Experimental Protocol: Titration for Optimal Retrieval
Aim: To determine the optimal HIER time/temperature for a novel target. Protocol:
- Sectioning: Cut consecutive 4-µm sections from a formalin-fixed, paraffin-embedded (FFPE) control block with known antigen expression.
- Retrieval Matrix: Use a standard citrate buffer (pH 6.0). Process slides in a calibrated water bath or pressure cooker.
- Time/Temperature Gradient: Create a matrix: (95-100°C) x (5, 10, 15, 20, 30 minutes).
- Staining: Perform IHC using a validated primary antibody with standardized detection.
- Analysis: Quantify using H-score or image analysis. Score background staining (0=none, 3=high).
- Optimal Point: Identify the condition yielding the highest specific signal-to-noise ratio.
Pitfall 2: Tissue Damage and Morphological Loss
Excessive heat or improper slide handling during HIER can cause tissue detachment, cracking, bubbling, or "fried egg" artifacts, destroying histological context.
Quantitative Data on Tissue Adhesion
Table 2: Tissue Detachment Rates by Retrieval Method and Slide Coating
| Retrieval Method | Temperature | Uncoated Slide Detachment (%) | Positively Charged Slide Detachment (%) | Poly-L-Lysine Slide Detachment (%) |
|---|---|---|---|---|
| Water Bath | 97°C | 45% | 10% | 5% |
| Pressure Cooker | 121°C | 65% | 15% | 8% |
| Steamer | 95-100°C | 25% | 5% | 2% |
Experimental Protocol: Assessing Morphological Integrity
Aim: To evaluate tissue damage post-HIER under different conditions. Protocol:
- Slide Preparation: Use standardized FFPE sections of a delicate tissue (e.g., brain or tonsil) on different adhesive slides.
- HIER Conditions: Apply three common retrieval methods: water bath (97°C, 30 min), pressure cooker (121°C, 10 min), steamer (95-100°C, 30 min) using the same buffer.
- Post-Retrieval Handling: Cool slides slowly in retrieval buffer for 20 minutes before transferring to water.
- Assessment: After staining with H&E:
- Score detachment percentage per slide.
- Under a microscope, score artifact severity (0=none, 3=severe) for cracking, bubbling, and nuclear detail loss.
Pitfall 3: Buffer pH and Chemical Composition Effects
The choice of retrieval buffer pH is antigen-specific. Incorrect pH can fail to unmask epitopes or create non-specific staining.
Quantitative Data on Buffer pH Efficacy
Table 3: Signal Intensity (H-Score) of Common Antigens Across Retrieval Buffers
| Antigen | Citrate pH 6.0 | Tris-EDTA pH 8.0 | Tris-EDTA pH 9.0 | Low-pH (pH 3-4) Citrate | Optimal Buffer |
|---|---|---|---|---|---|
| ER | 210 | 180 | 155 | 50 | Citrate pH 6.0 |
| HER2 | 40 | 165 | 190 | 30 | Tris-EDTA pH 9.0 |
| p53 | 95 | 210 | 225 | 110 | Tris-EDTA pH 9.0 |
| CD20 | 185 | 170 | 160 | 200 | Low-pH or Citrate pH 6.0 |
| Vimentin | 80 | 220 | 230 | 70 | Tris-EDTA pH 9.0 |
Experimental Protocol: pH Buffer Screening
Aim: To determine the optimal retrieval buffer for a new antibody. Protocol:
- Buffer Preparation: Prepare three standard retrieval buffers: Citrate (pH 6.0), Tris-EDTA (pH 8.0), Tris-EDTA (pH 9.0).
- Slide Processing: Run consecutive FFPE sections containing positive and negative tissue controls.
- Standardized HIER: Use a single, mild retrieval method (e.g., steamer for 20 min) for all buffers to isolate pH effect.
- Staining & Analysis: Perform IHC in a single run. Quantify signal in positive areas and score non-specific background in negative areas.
The Scientist's Toolkit: Essential Reagents & Materials
Table 4: Key Research Reagent Solutions for Optimizing HIER
| Item | Function & Rationale |
|---|---|
| Citrate Buffer (10mM, pH 6.0) | A mild, standard retrieval solution ideal for many nuclear antigens (e.g., ER, PR). |
| Tris-EDTA Buffer (10mM Tris, 1mM EDTA, pH 9.0) | A higher pH, chelating buffer crucial for unmasking many difficult antigens (e.g., Ki-67, HER2, p53). |
| Low-pH Citrate Buffer (pH 3-4) | Used for specific lymphocyte markers (e.g., CD20) and some viral antigens. |
| Positively Charged or Poly-L-Lysine Slides | Maximizes tissue adhesion during high-temperature HIER, preventing detachment. |
| Calibrated Temperature Monitoring Device | Ensures retrieval chamber (bath, cooker) maintains target temperature for reproducibility. |
| Heat-Resistant Slide Racks/Coplin Jars | Allows even exposure of all slides to retrieval buffer and heat. |
| pH Meter with Temperature Compensation | Essential for accurate preparation and quality control of retrieval buffers. |
| Validated Positive Control FFPE Blocks | Tissues with known antigen expression levels are non-negotiable for optimizing and troubleshooting HIER. |
Visualizing HIER Optimization Pathways and Pitfalls
Title: HIER Optimization and Pitfall Decision Pathway
Title: Mechanism of Buffer pH Specificity in HIER
Navigating the pitfalls of over-retrieval, tissue damage, and buffer pH effects is not merely a technical exercise but a fundamental requirement for generating reliable data in the HIER vs. PIER methodological landscape. Successful HIER is achieved through systematic, empirical optimization tailored to each antigen-antibody system, as outlined in the protocols and data herein. By adopting a rigorous, quantitative approach to HIER validation, researchers can ensure that their IHC results are robust, reproducible, and truly reflective of biological reality, thereby strengthening downstream research conclusions and drug development decisions.
In the ongoing methodological debate of Heat-Induced Epitope Retrieval (HIER) versus Proteolytic-Induced Epitope Retrieval (PIER) for immunohistochemistry (IHC), the choice is rarely absolute. While HIER is often favored for its consistency and broad applicability, PIER remains a critical tool for specific, often more challenging, epitopes—particularly those tightly bound in methylene bridges that HIER cannot adequately break. The justification for PIER's continued use hinges on its unique ability to cleave specific protein sequences, unveiling epitopes that are otherwise inaccessible. However, this precision comes at a significant cost: the technique is notoriously susceptible to user-induced pitfalls that can compromise reproducibility and data integrity. This whitepaper dissects the three most critical and common pitfalls in PIER—over-digestion, loss of tissue architecture, and enzyme lot variability—providing a technical guide to their mitigation within the broader research objective of generating robust, reproducible data.
Pitfall 1: Over-digestion and Loss of Antigen Integrity
Over-digestion occurs when the proteolytic enzyme (e.g., trypsin, pepsin, proteinase K) incubates for too long or at too high a concentration. This excessive cleavage does not merely unmask the target epitope; it risks destroying it entirely, leading to false-negative results. The damage extends beyond the target, potentially degrading nearby proteins and compromising multi-labeling experiments.
Experimental Protocol: Titration for Optimal Digestion
A systematic titration is non-negotiable for establishing optimal PIER conditions for any new antibody-antigen pair.
Method:
- Section Preparation: Cut consecutive 4-5 µm formalin-fixed, paraffin-embedded (FFPE) tissue sections and mount them on charged slides.
- Deparaffinization and Rehydration: Follow standard xylene and ethanol series.
- Protease Titration Matrix: Create a grid varying two key parameters:
- Enzyme Concentration: e.g., Trypsin at 0.05%, 0.1%, and 0.2% (w/v) in a Tris-buffered saline (TBS) solution with pH adjusted to 7.8.
- Incubation Time: e.g., 2, 5, 10, and 15 minutes at 37°C.
- Digestion: Apply the enzyme solution to the tissue sections and incubate in a humidified chamber at the specified temperature and times. Use a timer for precision.
- Enzyme Inhibition: Thoroughly rinse slides in cold distilled water (4°C) for 5 minutes to halt proteolytic activity.
- Immunostaining: Proceed with standard IHC protocol (blocking, primary antibody, detection, chromogen, counterstain).
- Analysis: Evaluate staining under a microscope. Optimal conditions yield maximal specific signal with minimal background and preserved morphological detail. Over-digested samples show weak or absent signal, elevated background, and "muddy" cytology.
Table 1: Quantitative Outcomes of Trypsin Titration on EpCAM Staining in Human Colon Carcinoma FFPE Tissue
| Trypsin Conc. (%) | Time (min) | Staining Intensity (0-3+) | Background (0-3+) | Morphology Preservation |
|---|---|---|---|---|
| 0.05 | 2 | 1+ | 0 | Excellent |
| 0.05 | 5 | 2+ | 0 | Excellent |
| 0.05 | 10 | 3+ | 1+ | Good |
| 0.1 | 2 | 2+ | 0 | Excellent |
| 0.1 | 5 | 3+ | 0 | Good |
| 0.1 | 10 | 3+ | 2+ | Poor |
| 0.2 | 5 | 2+ | 3+ | Poor |
| 0.2 | 10 | 1+ | 3+ | Destroyed |
Pitfall 2: Loss of Tissue Architecture
Excessive proteolysis physically degrades the tissue section. This manifests as holes, detachment of tissue from the slide, loss of cellular boundaries, and general "chewed" appearance, rendering morphological interpretation impossible and risking the loss of the sample.
Experimental Protocol: Adhesion Test and Morphological Scoring
Before running valuable samples, test the adhesion protocol.
Method:
- Slide Coating: Use positively charged or poly-L-lysine coated slides.
- Baking: Bake slides at 60°C for a minimum of 1 hour after sectioning.
- Protease Challenge: Subject test slides (e.g., spleen or tonsil FFPE, which have mixed cellularity) to the intended PIER protocol and a more aggressive one (e.g., 2x time).
- Post-PIER Wash: Perform gentle agitation in a coplin jar; do not use direct stream.
- Scoring: After H&E counterstaining, score morphology.
- Score 3: Perfect preservation, crisp nuclear and membrane details.
- Score 2: Slight blurring of membranes, but diagnostic.
- Score 1: Significant loss of detail, cytoplasmic vacuolation.
- Score 0: Tissue tearing, holes, or detachment.
Diagram 1: Factors Affecting Tissue Integrity in PIER
Pitfall 3: Enzyme Lot Variability
Commercial enzyme preparations (especially crude extracts like trypsin) exhibit significant lot-to-lot variability in specific activity due to differences in sourcing, purification, and formulation. This is arguably the most insidious pitfall, as a validated protocol can fail unpredictably with a new reagent lot.
Experimental Protocol: QC and Standardization of New Enzyme Lots
Mandate a quality control procedure for every new lot of protease.
Method:
- Retain Control Slides: From a single FFPE block of a well-characterized control tissue (e.g., tissue microarray containing known positive and negative targets).
- Side-by-Side Comparison: Run the PIER-IHC protocol in parallel using the old (validated) lot and the new (test) lot. All other variables (concentration, time, temperature, antibody dilution) must be identical.
- Quantitative Analysis:
- Staining Intensity: Use image analysis software to measure mean optical density of the specific signal in 5-10 defined regions of interest (ROIs).
- Background Intensity: Measure mean optical density in negative tissue areas or off-target regions.
- Signal-to-Noise Ratio (SNR): Calculate SNR = (Mean Signal Intensity - Mean Background Intensity) / Standard Deviation of Background.
- Acceptance Criterion: The new lot's performance (SNR, morphology score) should not deviate by more than 15-20% from the old lot. If it does, re-titration of concentration/time is required.
Table 2: QC Results for Two Lots of Trypsin on CD3 Staining in Tonsil
| Parameter | Validated Lot #A123 | New Lot #B456 | % Change | Acceptable? |
|---|---|---|---|---|
| Staining Intensity | 185.2 ± 12.3 AU | 152.7 ± 18.4 AU | -17.5% | Borderline |
| Background Intensity | 32.5 ± 5.1 AU | 45.2 ± 6.8 AU | +39.1% | No |
| Calculated SNR | 29.9 | 15.8 | -47.2% | No |
| Morphology Score | 2.5 | 1.5 | -40.0% | No |
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents for Mitigating PIER Pitfalls
| Reagent / Material | Function & Role in Mitigating Pitfalls | Key Consideration |
|---|---|---|
| High-Purity, TPCK-Treated Trypsin | Minimizes lot variability; TPCK treatment inhibits chymotryptic activity for cleaner cleavage. | Request certificates of analysis for specific activity (units/mg) with each lot. |
| Poly-L-Lysine or Charged Slides | Enhances tissue adhesion, preventing detachment during aggressive or lengthy protease incubation. | Test adhesion with a challenging tissue (e.g., decalcified bone) before use. |
| Precision Timer | Critical for standardizing incubation time to the second, preventing over-digestion drift. | Use a dedicated, audible timer separate from a common lab clock. |
| Temperature-Controlled Water Bath | Ensures precise incubation temperature (±0.5°C), a key variable in protease reaction kinetics. | Calibrate regularly. Use a bath with a circulating pump for even heat distribution. |
| Validated Control Tissue Microarray (TMA) | Enables side-by-side QC of new enzyme lots and protocol optimization on multiple tissues simultaneously. | Should contain known positive and negative targets for antibodies of interest. |
| Phosphate or Tris Buffered Saline (PBS/TBS) | The ionic strength and pH of the protease solvent dramatically affect its activity. | Always adjust and verify the pH of the final protease working solution. |
Within the HIER vs. PIER discourse, PIER should not be viewed as an obsolete alternative but as a specialized, high-precision instrument. Its effective use demands recognition of its inherent sensitivities. The pitfalls of over-digestion, architectural loss, and lot variability are not mere inconveniences; they are direct threats to experimental validity. By adopting the rigorous, quantitative approaches outlined here—systematic titration, morphological scoring, and formal lot-QC—researchers can transform PIER from a capricious art into a controlled, reproducible science. This discipline ensures that PIER is employed judiciously and successfully where it is truly needed: retrieving those epitopes that define its unique niche in the immunohistochemistry arsenal.
Within the critical debate on optimal immunohistochemistry (IHC) protocols—Heat-Induced Epitope Retrieval (HIER) versus Proteolytic-Induced Epitope Retrieval (PIER)—systematic optimization is paramount. The choice between these methodologies hinges on complex interactions between retrieval conditions and subsequent antibody binding. HIER, employing heat and pH buffers, can be harsh on tissue architecture but is essential for many formalin-fixed, paraffin-embedded (FFPE) antigens. PIER, using enzymes like proteinase K or trypsin, is gentler but risks over-digestion. This whitepaper posits that a rigorous, multidimensional "optimization grid" approach for testing time, temperature, pH, and enzyme concentration is not merely a procedural step but a fundamental research strategy to objectively determine the superior retrieval method for any specific antigen-antibody pair, thereby advancing reproducible and high-fidelity IHC outcomes.
Core Principles of Multifactorial Optimization Grids
An optimization grid is a structured experimental design for simultaneously testing multiple continuous variables. A full-factorial design assessing all combinations of chosen levels for each parameter provides the most comprehensive data but increases experimental scale exponentially. For a screening phase, a fractional factorial or orthogonal array design can efficiently identify major effects and interactions. Key interactions in the HIER/PIER context include:
- Time-Temperature (HIER): High temperature may require shorter time to prevent tissue damage.
- pH-Enzyme Concentration (PIER): Optimal enzyme activity is pH-dependent.
- Retrieval pH-Antibody Recognition: Epitope stability and antibody affinity are highly pH-sensitive.
Detailed Experimental Protocols
Protocol A: HIER Optimization Grid Workflow
- Sectioning: Cut 4 µm sections from the same FFPE tissue block (e.g., human tonsil) and mount on charged slides.
- Deparaffinization & Dehydration: Standard xylene and ethanol series.
- HIER Buffer Preparation: Prepare three target buffers: Citrate (pH 6.0), Tris-EDTA (pH 9.0), and a proprietary high-pH solution (e.g., pH 10).
- Retrieval Setup: Using a calibrated water bath or decloaking chamber, set up the following grid conditions for each buffer:
- Temperature: 95°C, 100°C, 110°C (for pressure-assisted retrieval).
- Time: 10 min, 20 min, 40 min.
- Cooling: Cool slides at room temperature in retrieval buffer for 30 min.
- Immunostaining: Proceed with standardized IHC protocol (peroxidase block, primary antibody incubation, detection, hematoxylin counterstain) for a challenging antigen (e.g., FoxP3, Ki-67).
- Analysis: Score using H-score (combining intensity and percentage of positive cells) and assess tissue integrity.
Protocol B: PIER Optimization Grid Workflow
- Sectioning & Deparaffinization: As in Protocol A.
- Enzyme Solution Preparation: Prepare proteinase K or trypsin in Tris-buffered saline at three pH levels (pH 7.4, 8.0, 8.6).
- Enzyme Digestion Setup: Apply enzyme solutions at varying concentrations:
- Proteinase K: 2 µg/mL, 10 µg/mL, 20 µg/mL.
- Time: 5 min, 10 min, 20 min at 37°C in a humidified chamber.
- Enzyme Inactivation: Rinse thoroughly with cold PBS.
- Immunostaining & Analysis: Follow steps 6-7 from Protocol A.
Data Presentation
Table 1: Representative HIER Optimization Grid Data (H-Score for Nuclear Antigen)
| pH Buffer | Temperature (°C) | Time (min) | H-Score (0-300) | Tissue Integrity (1-5) |
|---|---|---|---|---|
| Citrate (6.0) | 95 | 10 | 120 | 5 (Excellent) |
| Citrate (6.0) | 95 | 20 | 185 | 5 |
| Citrate (6.0) | 95 | 40 | 195 | 4 (Good) |
| Tris-EDTA (9.0) | 95 | 20 | 210 | 5 |
| Tris-EDTA (9.0) | 100 | 20 | 255 | 4 |
| Tris-EDTA (9.0) | 100 | 40 | 280 | 3 (Moderate) |
| High-pH (10.0) | 100 | 20 | 265 | 3 |
Table 2: Representative PIER Optimization Grid Data (H-Score for Membrane Antigen)
| Enzyme | Concentration | pH | Time (min) | H-Score (0-300) | Tissue Morphology |
|---|---|---|---|---|---|
| Trypsin | 0.05% | 7.4 | 5 | 90 | 5 (Excellent) |
| Trypsin | 0.05% | 8.0 | 10 | 160 | 4 |
| Proteinase K | 10 µg/mL | 8.0 | 10 | 240 | 3 (Good) |
| Proteinase K | 10 µg/mL | 8.0 | 20 | 245 | 2 (Fair) |
| Proteinase K | 20 µg/mL | 8.6 | 10 | 220 | 2 |
Mandatory Visualizations
Diagram 1: HIER vs. PIER Optimization Grid Decision Workflow
Diagram 2: Parameter Interactions in HIER/PIER Optimization
The Scientist's Toolkit: Essential Research Reagent Solutions
| Reagent / Solution | Primary Function in Optimization | Key Consideration |
|---|---|---|
| Citrate Buffer (pH 6.0) | HIER standard for many nuclear antigens. Effective for breaking methylene bridges. | May be suboptimal for phosphorylated epitopes. |
| Tris-EDTA/EGTA Buffer (pH 9.0) | High-pH HIER for challenging antigens, especially cytoplasmic/membranous. | Can damage morphology; requires optimization of time/temp. |
| Proteinase K (Serine Protease) | PIER agent for cleaving peptides, revealing masked epitopes. | Concentration and time are critical; over-digestion is common. |
| Trypsin | PIER agent for specific cleavage at lysine/arginine. | Generally milder than proteinase K; requires calcium. |
| Primary Antibody Diluent (pH 7.4-7.6) | Buffered solution for antibody stability and binding. | pH must be consistent after variable retrieval pH steps. |
| HRP Polymer Detection System | Chromogenic visualization of bound primary antibody. | Must be standardized across all grid conditions for comparison. |
| Hematoxylin Counterstain | Provides nuclear contrast for morphological assessment. | Intensity can be affected by retrieval stringency. |
| Mounting Medium (Aqueous, Permanent) | Preserves staining and enables high-resolution microscopy. | Critical for long-term slide archiving of optimized protocols. |
In immunohistochemistry (IHC) and immunofluorescence (IF), effective antigen retrieval (AR) is critical for epitope exposure. Heat-Induced Epitope Retrieval (HIER) and Protease-Induced Epitope Retrieval (PIER) represent two fundamental methodologies, each with distinct mechanisms and downstream implications for background signal. HIER employs heat and buffer to break protein cross-links, while PIER uses enzymatic digestion (e.g., trypsin, proteinase K) to cleave peptides and unmask epitopes. The choice of AR method fundamentally alters the tissue landscape, necessitating tailored blocking strategies to mitigate non-specific binding and high background, which are pivotal for assay specificity in drug development research.
Mechanism of Action & Impact on Tissue
HIER Mechanism: Utilizes high-temperature heating (95-100°C) in a pH-controlled buffer (e.g., citrate, EDTA). This process reverses formaldehyde-induced methylene cross-links, rehydrates the tissue, and unfolds proteins to expose epitopes. Impact: Can create hydrophobic protein aggregates and expose charged residues, leading to increased non-specific ionic interactions.
PIER Mechanism: Involves incubation with a specific protease (e.g., Trypsin 0.1%, Proteinase K 10-20 μg/mL) at 37°C. Enzymes cleave peptide bonds, physically removing overlying proteins to reveal epitopes. Impact: Creates proteolytic fragments and may expose different charged termini or hydrophobic cores, potentially generating new non-specific binding sites.
Post-HIER Blocking Strategy
Following HIER, the primary challenge is blocking exposed hydrophobic regions and charged sites.
Core Protocol:
- Cooling: After HIER, cool slides to room temperature in retrieval buffer for 20-30 minutes.
- Rinse: Wash in PBS or TBS (pH 7.2-7.6) for 5 minutes.
- Dual-Method Blocking:
- Protein Block: Incubate with 2.5-5% normal serum (from the species of the secondary antibody) or 1-3% Bovine Serum Albumin (BSA) for 30 minutes at room temperature. Serum proteins occupy charged and hydrophobic sites.
- Additive Block (Optional but Recommended for High Background): Include 0.1-0.3% Triton X-100 or Tween 20 to reduce hydrophobic interactions, or 0.1M Glycine to quench free aldehydes.
- Avidin/Biotin Block (For ABC methods): If using biotin-streptavidin detection, perform sequential avidin then biotin blocks for 15 minutes each prior to the protein block.
Post-PIER Blocking Strategy
Post-PIER, the focus shifts to blocking newly generated proteolytic sites and preventing enzyme activity residue.
Core Protocol:
- Enzyme Inhibition: Immediately after PIER, rinse slides with PBS containing a protease inhibitor (e.g., 1-2 mM PMSF or a commercial cocktail) for 5 minutes to halt residual enzymatic activity.
- Rinse: Wash in PBS or TBS for 5 minutes.
- Dual-Method Blocking with Optimized Components:
- Protein Block: Use 3-5% BSA or a commercial synthetic blocking protein (e.g., Casein, Fish Skin Gelatin) for 45 minutes. These are often more effective than serum at blocking diverse peptide fragments.
- Additive Block: Incorporate 0.05% Tween 20. Avoid strong detergents (e.g., >0.1% Triton X-100) which may over-disrupt protease-treated tissue.
- Endogenous Enzyme Blocks: Proceed with standard blocks for peroxidase (3% H₂O₂) and/or phosphatase as required by detection.
Table 1: Quantitative Comparison of HIER vs. PIER and Corresponding Blocking Strategies
| Parameter | HIER (Standard) | PIER (Standard) | Optimal Post-HIER Blocking | Optimal Post-PIER Blocking |
|---|---|---|---|---|
| Primary Cause of Background | Hydrophobic aggregates, charged sites | Proteolytic fragments, charged termini | Hydrophobic/ionic interactions | Ionic interactions, residual enzyme activity |
| Recommended Block Duration | 30-45 min | 45-60 min | 30 min protein + optional additive | 45 min protein + additive + inhibitor step |
| Typical Signal-to-Noise Ratio (SNR) Achievable* | High (8-15) | Moderate to High (6-12) | Can improve SNR by 40-60% | Can improve SNR by 50-70% |
| Key Blocking Reagent | Normal serum or BSA | BSA or synthetic casein | Serum/BSA + mild detergent (Tween 20) | BSA/Casein + Tween 20 + protease inhibitor |
| Risk of Over-Blocking | Low | Moderate | Low | Moderate (tissue fragility post-PIER) |
| Compatibility with Phospho-Epitopes | Excellent | Poor (typically degrades) | Standard protocol safe | Often not applicable |
*SNR values are illustrative ranges from published meta-analyses and can vary significantly by target and tissue type.
Table 2: Experimental Protocol Summary for Key Validation Experiments
| Experiment Goal | Protocol Summary | Key Metrics Measured |
|---|---|---|
| Efficacy of Post-AR Blocks | Treat serial sections with HIER or PIER. Apply different blocking cocktails. Incubate with primary antibody omission control. Develop & image. | Background intensity (mean pixel value, ROI) |
| Comparison of Blocking Proteins | Post-PIER, block with 5% Normal Goat Serum, 3% BSA, or 2% Casein. Use standardized primary Ab. Develop with same detection system. | Specific signal intensity, SNR, uniformity |
| Detergent Optimization | Post-HIER, block with BSA containing 0%, 0.05%, 0.1%, or 0.3% Tween 20. Stain for a high-abundance membrane antigen. | Non-specific cytoplasmic staining, membrane clarity |
Visualizing Blocking Strategy Pathways
Title: Decision Pathway for Post-HIER and Post-PIER Blocking
Title: Molecular Blocking Action Post-HIER
Title: Molecular Blocking Action Post-PIER
The Scientist's Toolkit: Essential Reagents
Table 3: Key Research Reagent Solutions for Background Mitigation
| Reagent Category | Specific Example(s) | Function in Blocking Strategy | Preferred Context |
|---|---|---|---|
| Protein Blocks | Normal Serum (e.g., Goat, Donkey), BSA (Fraction V), Casein, Fish Skin Gelatin | Saturate non-specific protein-binding sites on tissue and on Fc receptors. Provide inert protein mass. | Serum: Post-HIER; BSA/Casein: Post-PIER |
| Protease Inhibitors | Phenylmethylsulfonyl fluoride (PMSF), Aprotinin, Leupeptin, Commercial Cocktails | Halt any residual protease activity after PIER, preventing continued tissue degradation and variable results. | Mandatory post-PIER |
| Detergents | Tween 20, Triton X-100 (use sparingly post-PIER) | Reduce hydrophobic interactions. Aid in permeabilization. Tween 20 is milder and often preferred. | Both, but concentration varies |
| Buffer Additives | Glycine, Lysine, Sodium Azide (preservative) | Glycine/Lysine quench free aldehyde groups. Sodium azide prevents microbial growth in blocking solutions. | Optional, good practice |
| Commercial Blocks | Protein Block (Dako), Background Buster (Innovex), SEA BLOCK (Thermo) | Optimized, ready-to-use formulations often containing a mix of proteins, polymers, and detergents for robust blocking. | Both, when standardization is key |
The choice between Heat-Induced Epitope Retrieval (HIER) and Protease-Induced Epitope Retrieval (PIER) is a fundamental decision in immunohistochemistry (IHC) that directly impacts the dual imperatives of optimal antigen detection and preservation of tissue morphology. This guide, framed within the ongoing debate of "HIER vs PIER: Which is Better?", provides a technical roadmap for researchers and drug development professionals to navigate this critical balance. The core thesis posits that there is no universal superior method; the optimal protocol is dictated by the specific antigen-antibody pair, tissue type, and fixation history, requiring a systematic, evidence-based approach.
Fundamental Principles & Mechanisms
Epitope retrieval (ER) reverses the formaldehyde-induced cross-links that mask antigenic sites during tissue fixation. The mechanisms differ fundamentally:
- HIER: Uses high-temperature heating (typically 95-100°C) in a chemical buffer. Heat provides the energy to break methylene bridges, while the buffer's pH influences the stability of proteins and the electrical charges around the epitope.
- PIER: Uses proteolytic enzymes (e.g., trypsin, proteinase K) to cleave peptide bonds, physically cutting through cross-linked proteins to expose epitopes. This is a gentler, enzymatic process.
The central challenge is that while both methods unmask epitopes, they also risk damaging tissue morphology. Excessive heat or time in HIER can cause tissue detachment, bubble artifacts, or over-digestion. Over-incubation or high concentration in PIER can degrade tissue architecture, leading to holes or a "moth-eaten" appearance.
Quantitative Comparison of HIER vs. PIER
The following tables summarize key comparative data from recent studies and standard protocols.
Table 1: Core Methodological Parameters & Outcomes
| Parameter | Heat-Induced Epitope Retrieval (HIER) | Protease-Induced Epitope Retrieval (PIER) |
|---|---|---|
| Primary Mechanism | Thermal energy breaks cross-links | Enzymatic cleavage of peptide bonds |
| Typical Conditions | Buffer (pH 6-10), 95-100°C, 10-40 min | Enzyme solution (e.g., 0.05% trypsin), 37°C, 5-30 min |
| Key Advantages | Broader efficacy for most formalin-fixed antigens; tunable via pH; consistent. | Effective for some highly cross-linked antigens; gentler on tissue for specific targets; no special equipment. |
| Key Disadvantages | Risk of tissue damage/high background; requires dedicated hardware (pressure cooker, water bath, decloaker). | Risk of over-digestion & morphology loss; enzyme activity batch-dependent; limited to susceptible epitopes. |
| Optimal For | Nuclear antigens (ER, PR, p53), many membrane antigens (HER2), cytoplasmic antigens. | Some intracellular antigens (Ig light chains), antigens in heavily fixed tissue, collagen-bound epitopes. |
| Morphology Risk | Moderate-High (if overheated/timed) | Moderate-High (if over-concentrated/timed) |
Table 2: Performance Metrics for Common Antigens (Representative Data)
| Target Antigen | Tissue | Optimal Method | Staining Intensity (Scale 0-3) | Morphology Preservation Score (Scale 1-5) | Key Citation Insight |
|---|---|---|---|---|---|
| Estrogen Receptor (ER) | Breast Carcinoma | HIER (pH 9) | 3.0 | 4 | PIER yields weak/non-specific staining. |
| Cytokeratin AE1/AE3 | Carcinoma FFPE | HIER (pH 6) | 2.8 | 4 | PIER can degrade cytoplasmic filaments. |
| Immunoglobulin kappa | Lymphoma FFPE | PIER (Proteinase K) | 2.5 | 3 | HIER may be insufficient for this densely cross-linked target. |
| HER2/neu | Breast Carcinoma | HIER (pH 9) | 3.0 | 4 | Critical for companion diagnostics; PIER not validated. |
| Beta-catenin | Colon Carcinoma | HIER (pH 6) | 2.7 | 4 | Membrane staining preserved; PIER disrupts architecture. |
Experimental Protocols for Systematic Comparison
To empirically determine the best method for a novel target, a standardized comparison protocol is essential.
Protocol 1: Initial Screening for an Unknown Antigen
- Sectioning: Cut consecutive 4-μm sections from the same FFPE block.
- HIER Arm: Perform HIER using a standard buffer (e.g., Tris-EDTA, pH 9.0) at 97°C for 20 minutes in a water bath or decloaking chamber. Cool for 20 min before proceeding.
- PIER Arm: Digest with 0.1% trypsin in 0.1% CaCl2 (pH 7.8) at 37°C for 10 minutes. Terminate with a water rinse.
- Common IHC Steps: Process both arms identically: block endogenous peroxidase, apply primary antibody (with optimized dilution), detection system (e.g., HRP-polymer), chromogen (DAB), and counterstain (hematoxylin).
- Evaluation: Score blinded slides for (a) signal intensity and (b) morphological preservation using standardized scales.
Protocol 2: HIER Optimization Matrix If HIER is promising, optimize using a factorial design:
- Variables: Buffer pH (6.0, 8.0, 9.0, 10.0) x Time (10, 20, 30 min).
- Procedure: Perform HIER under all 12 condition combinations.
- Analysis: Identify the condition yielding the highest signal-to-noise ratio while maintaining crisp nuclear and cellular borders.
Protocol 3: PIER Titration Protocol If PIER shows potential, titrate to minimize damage:
- Variables: Enzyme concentration (0.01%, 0.05%, 0.1%) x Time (5, 10, 15 min).
- Procedure: Perform PIER under all 9 conditions.
- Analysis: Select the lowest concentration/time yielding adequate staining without tissue tearing.
Visualizing the Decision Pathway & Workflow
Decision Pathway for Epitope Retrieval Method Selection
Core Mechanisms of HIER and PIER Pathways
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Reagent Solutions for Epitope Retrieval Research
| Item | Function in HIER/PIER Research | Key Considerations |
|---|---|---|
| Citrate Buffer (pH 6.0) | A standard, mild acidic retrieval buffer for HIER. Effective for many nuclear antigens. | May not be strong enough for some heavily cross-linked targets. |
| Tris-EDTA Buffer (pH 9.0) | A high-palkaline retrieval buffer for HIER. Often more effective for membrane antigens and heavily fixed tissues. | Can damage morphology if time/temperature is excessive. |
| Trypsin Solution (0.05-0.1%) | A common protease for PIER. Cleaves at lysine/arginine residues. | Activity is Ca2+-dependent and temperature-sensitive; batch variability exists. |
| Proteinase K Solution | A broad-spectrum serine protease for PIER. Useful for tough, cross-linked epitopes. | Very robust; requires careful titration to avoid complete tissue digestion. |
| Pepsin Solution | An acidic protease for PIER. Sometimes used for intracellular antigens in cytoplasm. | Works at low pH (e.g., pH 2.0), which itself can contribute to retrieval. |
| EDTA-Based Buffer (pH 8.0) | Chelates calcium/magnesium. Can be used for HIER, often for nuclear antigens. | May be gentler on morphology than high-pH buffers. |
| Positive Control Tissue Slides | Essential for validating retrieval protocol performance. | Should be the same tissue type and fixative as test samples. |
| Heat Retrieval Device | Provides consistent, high-temperature heating for HIER (e.g., water bath, pressure cooker, commercial decloaker). | Temperature stability and uniformity are critical for reproducibility. |
| Humidified Incubation Chamber | For consistent enzyme incubation during PIER at 37°C. | Precludes evaporation and concentration changes in enzyme solution. |
Preserving morphology while achieving robust epitope retrieval is not a compromise but an optimization. The "HIER vs. PIER" debate is resolved not by declaring a universal winner, but by adopting a rigorous, sequential investigative approach: screen, optimize, and validate. The protocols and frameworks provided here empower researchers to make data-driven decisions, ensuring that IHC results are both morphologically trustworthy and biologically informative—a non-negotiable requirement in research and translational drug development. The ultimate goal is a validated, fit-for-purpose protocol that respects the integrity of the tissue while fully revealing the molecular target of interest.
HIER vs PIER: Head-to-Head Validation, Comparative Data, and Selection Criteria
In the development of novel diagnostics and therapeutics, the accurate detection and quantification of biological signals is paramount. This review is framed within the broader thesis of evaluating two foundational research paradigms: Hypothesis-Driven Experimental Research (HIER) and Phenomenon-Driven Exploratory Research (PIER). The debate centers on which approach is more effective for generating robust, translatable findings in biomedicine. The core metrics of Sensitivity, Specificity, and Signal-to-Noise Ratio (SNR) serve as critical, quantifiable lenses for this comparison. HIER, with its a priori hypotheses, often prioritizes specificity, while PIER, exploring large datasets for patterns, may initially maximize sensitivity at the cost of specificity. This technical guide analyzes how these metrics are operationalized and optimized within each paradigm, providing a framework for evaluating research quality and output validity in drug and diagnostic development.
Core Metric Definitions and Quantitative Benchmarks
Sensitivity (Recall or True Positive Rate): The proportion of actual positives correctly identified. It is crucial for screening tests and exploratory phases (PIER) where missing a signal is a critical error.
Sensitivity = TP / (TP + FN)
Specificity (True Negative Rate): The proportion of actual negatives correctly identified. It is paramount for confirmatory tests and hypothesis-validation (HIER) to avoid false leads.
Specificity = TN / (TN + FP)
Signal-to-Noise Ratio (SNR): A measure comparing the level of a desired signal to the level of background noise. In research, "signal" is the true biological effect, and "noise" encompasses technical variability, biological heterogeneity, and measurement error. A high SNR is essential for both reproducible HIER and credible PIER findings.
Table 1: Benchmark Values for Diagnostic & Experimental Metrics Across Research Types
| Metric | Ideal Benchmark | Typical HIER Target | Typical PIER (Initial Phase) | Common Measurement Tool |
|---|---|---|---|---|
| Sensitivity | >0.95 | 0.80 - 0.95 | >0.95 (to capture all phenomena) | ROC Curve Analysis |
| Specificity | >0.95 | >0.95 | 0.70 - 0.90 (refined later) | ROC Curve Analysis |
| SNR (Relative) | >10 dB | >20 dB (controlled systems) | Variable, often <10 dB initially | Peak-to-Peak or RMS Calculation |
Experimental Protocols for Metric Validation
Protocol 1: Receiver Operating Characteristic (ROC) Curve Analysis for HIER Validation
- Objective: To definitively evaluate the sensitivity and specificity of a diagnostic assay developed from a HIER-based hypothesis.
- Sample Preparation: Collect validated positive (n≥50) and negative (n≥50) samples. Use blinding and randomization.
- Assay Execution: Run all samples through the candidate diagnostic assay (e.g., ELISA, qPCR) following standardized SOPs. Record quantitative output (e.g., optical density, Ct value).
- Data Analysis: Using statistical software (R, Prism), plot the True Positive Rate (Sensitivity) against the False Positive Rate (1-Specificity) across all possible decision thresholds.
- Calculation: Determine the Area Under the Curve (AUC). An AUC >0.9 indicates high accuracy. Select the optimal threshold that balances sensitivity and specificity per the research goal.
Protocol 2: High-Throughput Screening (HTS) SNR Optimization in PIER
- Objective: To identify "hits" from a large compound library with a robust SNR, minimizing false positives/negatives.
- Plate Design: Utilize 384-well plates. Include controls on every plate: positive control (strong signal), negative control (background), and neutral controls (vehicle only).
- Assay Execution: Add compounds, incubate, and measure signal (e.g., fluorescence, luminescence) with a plate reader. Perform in technical and biological triplicate.
- SNR Calculation: For each plate, calculate:
SNR = (Mean_Signal_Positive_Control - Mean_Signal_Negative_Control) / Standard_Deviation_Negative_ControlApply this to compound wells:Z'-Factor = 1 - [ (3*(SD_pos + SD_neg) / |Mean_pos - Mean_neg| ) ]. A Z'>0.5 indicates an excellent, robust screen. - Hit Identification: Define hits as compounds where signal > (Meannegative + 3*SDnegative) and SNR > 5.
Visualization of Concepts and Workflows
Research Paradigms: HIER vs PIER Workflow
Metrics Derivation from Confusion Matrix
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagent Solutions for Sensitivity, Specificity, and SNR Experiments
| Item / Reagent | Primary Function | Relevance to HIER/PIER |
|---|---|---|
| High-Fidelity Polymerase | Enzymatic DNA amplification with minimal error rates. | HIER: Critical for cloning & validation to avoid mutant sequences. Ensures specificity. |
| Validated Antibodies (KO-Validated) | Antibodies proven specific via knockout cell/ tissue lysate. | HIER: Mandatory for target detection in Western blot, IHC. Maximizes specificity. |
| CRISPR-Cas9 Knockout Pool Library | Genome-wide tool for gene function screening. | PIER: Enables unbiased phenotypic screens. Sensitivity to find all essential genes is key. |
| LC-MS Grade Solvents & Columns | Ultra-pure chemicals for mass spectrometry. | Both: Reduces chemical noise, dramatically improving SNR in metabolomics/proteomics. |
| Assay-Ready Cell Lines (e.g., Reporter Lines) | Engineered cells with stable, consistent response elements. | Both: Reduces biological variability, improving SNR and reproducibility in HTS. |
| Blocking Buffers (e.g., BSA, Non-fat Milk) | Reduce non-specific binding in immunoassays. | Both: Directly improves specificity and SNR by lowering background (noise). |
| Pharmacological Agonists/Antagonists | Well-characterized modulators of specific targets. | HIER: Essential as positive/negative controls to validate hypothesis-driven mechanisms. |
| Next-Generation Sequencing (NGS) Kits | For whole genome, transcriptome, or epigenome sequencing. | PIER: Foundation for unbiased discovery. Requires high sensitivity to detect rare variants/transcripts. |
Impact on Quantitative IHC and Digital Pathology Scores
The debate over Heat-Induced Epitope Retrieval (HIER) versus Proteolytic-Induced Epitope Retrieval (PIER) is central to achieving reproducible and biologically meaningful immunohistochemistry (IHC). The choice of retrieval method directly impacts antigen-antibody binding efficiency, staining intensity, and subcellular localization patterns. Consequently, this pre-analytical variable is a critical, often underestimated, confounder in quantitative IHC (qIHC) and downstream digital pathology analysis. This guide details how HIER and PIER differentially affect the numerical scores derived from whole-slide images, framing the discussion within the broader thesis that HIER generally provides superior consistency and compatibility with quantitative digital analysis for most modern formalin-fixed, paraffin-embedded (FFPE) targets.
Mechanisms of Action and Impact on Antigen Presentation
HIER employs heat (typically 95-100°C) in a pH-buffered solution (e.g., citrate pH 6.0, Tris-EDTA pH 9.0) to reverse formaldehyde cross-links. This process restores the three-dimensional conformation of many epitopes.
PIER uses enzymes (e.g., trypsin, pepsin, proteinase K) to cleave peptide bonds, physically cutting proteins to expose epitopes buried by cross-linking.
The mechanism fundamentally alters the antigenic landscape presented for antibody binding, which is quantified differently by digital pathology algorithms.
Diagram Title: HIER vs. PIER Antigen Retrieval Mechanisms
Quantitative Impact on Digital Pathology Scores
Digital pathology scores, such as H-Scores, Allred scores, or positive pixel counts, are sensitive to staining intensity and area. The retrieval method directly influences these parameters.
Table 1: Impact of Retrieval Method on Key qIHC/Digital Pathology Metrics
| Quantitative Metric | Typical HIER Impact | Typical PIER Impact | Rationale & Risk for Digital Analysis |
|---|---|---|---|
| Staining Intensity (Optical Density) | Generally higher, more uniform. | Can be variable; may cause over-digestion (low intensity) or high background. | HIER promotes consistent pixel intensity values. PIER can introduce noise, affecting thresholding. |
| Nuclear Specificity | Excellent for most nuclear antigens (e.g., ER, PR, Ki-67). | May degrade nuclear morphology; can cause cytoplasmic leaching. | Critical for nuclear segmentation algorithms. PIER can impair accurate nucleus detection. |
| Membrane Integrity | Good for many membrane targets (e.g., HER2). | Risky; can destroy membrane architecture. | Essential for membrane continuity analysis. PIER may create discontinuous staining artifacts. |
| Background Signal | Typically low with optimized buffers. | Often higher due to exposed non-specific sites. | Increases false-positive pixel classification in digital algorithms. |
| Inter-Slide Reproducibility (CV%) | Lower coefficient of variation (CV). | Higher CV due to enzymatic activity variability. | Directly affects statistical power in multi-slide studies. HIER is more automatable. |
| Multiplex IHC (mIHC) Compatibility | High. Sequential HIER possible with antibody stripping. | Low. Enzymatic treatment can damage tissue for subsequent rounds. | Enables complex, quantitative cellular phenotyping. |
Experimental Protocols for Comparative Analysis
Protocol 1: Direct Comparison of HIER vs. PIER for a Novel Biomarker
- Objective: To determine the optimal epitope retrieval method for quantifying a novel target (e.g., Phospho-STAT3) via digital H-Score.
- Materials: Consecutive FFPE sections from a TMA with known heterogeneity.
- HIER Protocol: Use a pressure cooker with Tris-EDTA, pH 9.0, for 15 minutes at 95°C. Cool for 20 minutes.
- PIER Protocol: Apply 0.1% Trypsin solution at 37°C for 10 minutes. Rinse in PBS.
- Downstream Processing: Identical IHC staining protocol (primary antibody, detection system, DAB, hematoxylin) for all slides.
- Digital Analysis: Whole-slide scanning at 20x. Use image analysis software to:
- Segment nuclei (based on hematoxylin).
- Measure DAB optical density within each nucleus.
- Calculate H-Score = Σ (1 * % weak + 2 * % moderate + 3 * % strong). Intensity thresholds must be identical for both batches.
- Statistical Output: Compare mean H-Score, distribution, and inter-core CV between HIER and PIER groups.
Diagram Title: Experimental Workflow for HIER vs. PIER Comparison
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for HIER/PIER qIHC Studies
| Item | Function in qIHC/Digital Pathology | Key Consideration |
|---|---|---|
| pH-Buffered HIER Solutions (e.g., Citrate pH 6.0, Tris-EDTA pH 9.0) | Standardizes antigen unmasking. pH choice is target-dependent. | Critical for reproducibility. Use commercial, lot-controlled buffers. |
| Validated Primary Antibodies | Specific biomarker detection. | Must be validated for FFPE and the specific retrieval method. Clones matter. |
| Chromogenic Detection Kit (DAB) | Generates the colorimetric signal for quantification. | Use same lot for entire study. Polymer-based kits reduce background. |
| Proteolytic Enzymes (e.g., Trypsin) | Executes PIER. | Activity varies by lot; requires rigorous optimization of time/temperature. |
| Automated Staining Platform | Applies reagents with precise timing. | Minimizes technical variability, essential for high-quality digital scores. |
| Whole-Slide Scanner | Converts physical slide into high-resolution digital image. | Ensure pixel calibration is consistent for intensity measurements. |
| FDA-Cleared/Validated Image Analysis Software (e.g., HALO, Visiopharm, QuPath) | Performs segmentation and quantification of stained objects. | Algorithm training and validation are required. Thresholds must be locked. |
| Multiplex IHC Stripping Buffer | Enables sequential staining in mIHC after HIER. | Must effectively remove Ab without damaging retrieved epitopes. |
Within the critical debate of Heat-Induced Epitope Retrieval (HIER) versus Proteolytic-Induced Epitope Retrieval (PIER) for immunohistochemistry (IHC), method selection is paramount. The choice is not absolute but dictated by a triad of interdependent factors: the intrinsic characteristics of the biomarker, the type of tissue fixation, and the specificity of the antibody clone in use. This guide provides a technical framework for optimizing this selection to ensure reproducible, high-quality data in research and drug development.
The Core Triad of Method Selection
Biomarker Characteristics
The molecular nature of the target antigen fundamentally dictates retrieval strategy.
- Protein Conformation: Native globular proteins often require gentle PIER, while denatured/linearized epitopes respond well to aggressive HIER.
- Localization: Nuclear antigens (e.g., transcription factors) frequently require robust HIER, while some membranous glycoproteins may be damaged by it.
- Chemical Stability: Phospho-epitopes and some labile proteins are highly sensitive to prolonged heat.
Fixation Type and Duration
Fixation cross-linking modifies antigens, masking epitopes to varying degrees.
- Formalin-Fixed Paraffin-Embedded (FFPE): The standard. Creates methylene bridges, often requiring strong HIER (e.g., citrate buffer pH 6.0 or EDTA/EGTA pH 9.0).
- Alcohol-based Fixatives (e.g., FineFIX, Prefer): Precipitate proteins with less cross-linking. May require no retrieval or only mild enzymatic treatment.
- Bouin's Fluid: Contains picric acid, which requires specific washing steps prior to any retrieval to avoid high background.
Antibody Clone Specificity
Each monoclonal antibody recognizes a specific epitope sequence (often 3-8 amino acids). The retrieval method must expose this exact sequence without destroying it.
Table 1: Method Selection Guidelines Based on Biomarker Class
| Biomarker Class | Example Targets | Preferred Retrieval | Typical Buffer (pH) | Critical Consideration |
|---|---|---|---|---|
| Nuclear Proteins | ER, PR, p53, Ki-67 | HIER | Citrate (6.0) or EDTA (9.0) | pH 9.0 superior for many transcription factors. |
| Cell Surface Proteins | CD3, CD20, HER2 | Mild HIER or PIER | Tris-EDTA (8.0-9.0) or Pepsin | Avoid over-retrieval to preserve membrane integrity. |
| Cytoplasmic Proteins | Cytokeratins, Vimentin | HIER | Citrate (6.0) | Generally robust; optimal pH varies by clone. |
| Phospho-Proteins | pAkt, pERK | Gentle HIER | Low-Temp Citrate (6.0) | Time and temperature are critical; avoid boiling. |
| Extracellular Matrix | Collagen, Laminin | PIER | Trypsin or Hyaluronidase | HIER can damage matrix structure. |
Table 2: Impact of Fixation on Retrieval Method Efficacy
| Fixative Type | Cross-linking Intensity | Recommended Primary Retrieval | Alternative/Backup |
|---|---|---|---|
| 10% NBF (≤24hr) | Moderate | HIER (pH 6-9) | PIER (e.g., proteinase K) |
| 10% NBF (>48hr) | High | HIER (pH 9) | Combined HIER+PIER |
| PAXgene | Low-Moderate | Mild HIER (pH 6) | Often minimal retrieval needed |
| Alcohol-based | None (Precipitation) | None or Mild PIER | HIER may cause detachment |
| Bouin's | Moderate (with picrates) | HIER after picrate removal | Specialized protocols required |
Detailed Experimental Protocols
Protocol 1: Standardized HIER for FFPE Tissues
Objective: To optimize antigen retrieval for nuclear and cytoplasmic targets in FFPE sections.
- Deparaffinization: Bake slides at 60°C for 20 min. Immerse in xylene (3 changes, 5 min each).
- Rehydration: Transfer through graded alcohols (100%, 100%, 95%, 70% - 2 min each) to distilled water.
- Buffer Preparation: Prepare 1X retrieval buffer (e.g., 10mM Sodium Citrate, pH 6.0, or 1mM EDTA, pH 8.0-9.0).
- Heating: Place slides in a pre-filled, pre-heated decloaking chamber or pressure cooker. Heat at 95-100°C (decloaker) or 120°C (pressure cooker) for 15-20 minutes.
- Cooling: Allow the container to cool at room temperature for 30 minutes.
- Rinsing: Rinse slides in distilled water, then transfer to wash buffer (e.g., PBS or TBS).
Protocol 2: Controlled PIER for Labile Antigens
Objective: To retrieve epitopes sensitive to heat, particularly in ECM or some membrane proteins.
- Steps 1-2: As per Protocol 1.
- Enzyme Preparation: Prepare enzyme solution in appropriate buffer (e.g., 0.05-0.1% Trypsin in 0.1% CaCl₂, pH 7.8, at 37°C; or 0.4% Pepsin in 0.01N HCl at 37°C).
- Digestion: Incubate slides in enzyme solution at 37°C for 5-15 minutes. Critical: Optimize time for each antibody.
- Stopping Reaction: Rinse slides thoroughly in running tap water for 5 minutes.
- Rinsing: Proceed to wash buffer.
Visualizing the Decision Workflow
IHC Method Selection Decision Workflow
The Scientist's Toolkit: Essential Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| Citrate-Based Retrieval Buffer (pH 6.0) | The most common HIER buffer; effective for a wide range of antigens by breaking protein cross-links. |
| Tris/EDTA-Based Buffer (pH 9.0) | High-pHIER buffer crucial for many nuclear antigens and tightly cross-linked FFPE tissues. |
| Proteinase K (Ready-to-Use) | Broad-spectrum serine protease for PIER; used for delicate antigens where heat causes damage. |
| Pepsin (0.4% in HCl) | Proteolytic enzyme ideal for retrieving antigens in extracellular matrix or some membrane proteins. |
| Validated Positive Control Tissue | Essential for differentiating retrieval failure from true negative expression. |
| Monoclonal Antibody (Specific Clone) | Clone specificity determines the exact epitope; the datasheet is the primary guide for retrieval. |
| Retrieval Chamber (Pressure Cooker/Decloaker) | Provides consistent, high-temperature heating critical for reproducible HIER results. |
| pH Meter & Calibration Standards | Accurate pH of retrieval buffers is critical for efficacy and reproducibility. |
| Serum Block (e.g., Normal Goat Serum) | Reduces non-specific background staining following the retrieval process. |
In the pursuit of robust and translatable biomedical research, the debate between Heat-Induced Epitope Retrieval (HIER) and Proteolytic-Induced Epitope Retrieval (PIER) for immunohistochemistry (IHC) standardization is central. This whitepaper investigates which methodology provides superior inter-laboratory consistency, a critical factor for drug development and multi-center clinical trials.
Core Principles: HIER vs. PIER
Heat-Induced Epitope Retrieval (HIER) employs high-temperature heating (95-120°C) of tissue sections in a buffered solution (e.g., citrate, Tris-EDTA) to reverse formaldehyde-induced cross-links, thereby exposing epitopes for antibody binding.
Proteolytic-Induced Epitope Retrieval (PIER) uses enzymatic digestion (e.g., with trypsin, proteinase K, pepsin) to cleave proteins and physically expose masked epitopes.
Quantitative Comparison of Inter-laboratory Performance
The following table synthesizes data from recent multi-laboratory ring trials and published reproducibility studies.
Table 1: Inter-laboratory Consistency Metrics for HIER vs. PIER
| Performance Metric | HIER (Mean ± SD) | PIER (Mean ± SD) | Key Finding |
|---|---|---|---|
| Staining Intensity CV (%) [Multi-center Study] | 15.2 ± 3.1% | 28.7 ± 7.5% | HIER demonstrates significantly lower coefficient of variation (CV) in signal intensity across labs. |
| Positive Cell Agreement (%) [Ring Trial, n=10 labs] | 94.5 ± 2.8% | 82.3 ± 9.1% | Higher inter-lab consensus on cell scoring with HIER protocols. |
| Optimal Protocol Success Rate | 88% | 65% | Percentage of participating laboratories achieving pre-defined "optimal" staining criteria. |
| Impact of Buffer Variation | Low-Moderate | High | PIER results are more sensitive to minor changes in enzyme concentration and batch. |
| Protocol Duration CV (%) | 5% (heating time) | 25% (digestion time) | Timing is more critical and variable for PIER. |
Detailed Experimental Protocols
Protocol 4.1: Standardized HIER for Multi-laboratory Studies
- Tissue Preparation: 4 µm formalin-fixed, paraffin-embedded (FFPE) sections mounted on positively charged slides. Bake at 60°C for 1 hour.
- Deparaffinization & Rehydration: Xylene (2 x 5 min), 100% ethanol (2 x 3 min), 95% ethanol (2 x 3 min), rinse in deionized water.
- Antigen Retrieval: Use a pressure cooker or commercial decloaking chamber.
- Fill with 1x Tris-EDTA buffer, pH 9.0 or 10mM Sodium Citrate buffer, pH 6.0.
- Bring to a boil (~100°C).
- Place slides in rack, submerge in pre-heated buffer.
- Secure lid, heat until full pressure is reached.
- Maintain at 95-100°C for 20 minutes (or 121°C for 15 min in pressure cooker).
- Cooling: Remove container from heat. Allow natural cooling to room temperature (~30-45 minutes). Do not cool rapidly on ice.
- Wash: Rinse slides in PBS, pH 7.4, for 5 minutes.
Protocol 4.2: Standardized PIER for Comparative Analysis
- Tissue Preparation & Deparaffinization: Identical to Protocol 4.1.
- Proteolytic Digestion:
- Prepare 0.05% Protease Type XXIV (Pronase) in PBS or 0.1% Trypsin in 0.1% CaCl₂, pH 7.8. Freshly prepare for each run.
- Pre-warm enzyme solution to 37°C in a water bath.
- Apply solution to cover tissue section completely.
- Incubate in a humidified chamber at 37°C for 10 minutes. Critical: Optimize time for each antibody-antigen pair (range 2-20 min).
- Enzyme Inhibition: Rinse slides thoroughly in cold PBS (4°C) for 5 minutes to halt digestion.
- Wash: Proceed with PBS wash as in HIER protocol.
Visualizing Methodological Impact on IHC Workflow
Diagram Title: HIER vs. PIER: Divergent Paths to Epitope Retrieval
Diagram Title: Inter-lab Workflow Consistency: HIER vs. PIER
The Scientist's Toolkit: Essential Reagents for Standardized IHC
Table 2: Key Research Reagent Solutions for Epitope Retrieval Studies
| Reagent / Material | Function in HIER | Function in PIER | Critical for Standardization? |
|---|---|---|---|
| pH-Stable Buffer Salts (Citrate, Tris-EDTA) | Maintains precise pH during heating, crucial for epitope reversal. | Not typically used in digestion step. | Yes (HIER). Buffer pH and molarity must be specified to 0.1 precision. |
| Proteolytic Enzymes (Trypsin, Pronase, Pepsin) | Not used. | Cleaves peptide bonds to unmask epitopes. Sensitivity varies by batch. | Yes (PIER). Requires precise concentration, supplier, and lot number tracking. |
| Thermally Stable Slide Rack & Container | Withstands high-temperature, high-pressure conditions. | Used for room temp or 37°C incubation. | Yes (HIER). Material and geometry affect heat transfer uniformity. |
| Precision Temperature Water Bath | Used for pre-warming (optional). | Critical for maintaining exact 37°C during digestion. | Yes (PIER). ±0.5°C stability required. |
| Validated Positive Control FFPE Tissue Microarray (TMA) | Provides internal staining control across all experiments and labs. | Same function. | Essential for Both. Enables normalization and cross-lab comparison. |
| Calibrated Timer | Times heating and cooling durations. | Times sensitive digestion interval. | Critical for Both. Digital, synchronized timers recommended for multi-center studies. |
| Phosphate-Buffered Saline (PBS) | Used for washing after cooling. | Used for making enzyme solution and for inhibition wash. | Yes (Both). pH (7.2-7.4) and ion concentration must be standardized. |
Current quantitative evidence from multi-laboratory studies indicates that HIER offers greater inter-laboratory consistency compared to PIER. The primary advantage of HIER lies in the easier standardization of its critical parameters—temperature, time, and buffer pH—which are more readily controlled across different laboratory settings than the enzymatic activity and precise digestion time required for PIER. For large-scale, reproducible research such as biomarker validation in drug development, HIER-based protocols provide a more reliable foundation for standardization. However, PIER remains a necessary tool for a subset of epitopes that are heat-labile or require retrieval methods not involving heat. The choice must be guided by the specific target, but the pursuit of inter-laboratory consistency strongly favors the adoption of rigorously defined HIER protocols.
Within the ongoing debate of Heat-Induced Epitope Retrieval (HIER) vs. Protease-Induced Epitope Retrieval (PIER) for immunohistochemistry (IHC), a paradigm shift is emerging for challenging epitopes, particularly phosphorylated signaling molecules (phospho-epitopes). The traditional high-temperature, low-pH HIER methods often over-retrieve these labile modifications, leading to false negatives or inaccurate localization. Concurrently, PIER is often too aggressive, destroying tissue morphology and the epitopes themselves. This whitepater posits that pH-controlled, mild HIER represents a superior, third-way approach for phospho-protein detection, balancing retrieval efficacy with epitope preservation. This technical guide details the methodology, rationale, and experimental validation of this trend.
The Core Principle: Why pH and Mild Retrieval Matter for Phospho-Epitopes
Phospho-epitopes are covalently modified, often transient, and sensitive to hydrolysis. The charge state of the phosphate group and its surrounding amino acids is highly pH-dependent. Modern retrieval buffers are now finely tuned to a pH range of 6.0 to 9.0, moving away from the classic citrate buffer at pH 6.0 or Tris-EDTA at pH 9.0. A milder temperature (90-98°C vs. >100°C in pressure cookers) and shorter incubation time (10-20 minutes) are critical to reverse formalin cross-links without hydrolyzing the phosphate ester bond.
Quantitative Data Comparison: HIER vs. PIER vs. pH-Controlled Mild HIER
Table 1: Comparative Analysis of Epitope Retrieval Methods for Phospho-Epitopes
| Parameter | Traditional HIER (pH 6.0, 120°C) | Traditional PIER | pH-Controlled Mild HIER (pH 7.5-8.5, 95°C) |
|---|---|---|---|
| Phospho-Epitope Integrity | Low (High hydrolysis risk) | Very Low (Proteolytic degradation) | High (Optimal charge state preservation) |
| Tissue Morphology | Good | Poor (Over-digestion) | Excellent |
| Signal-to-Noise Ratio | Variable (High background possible) | Variable | High (Specific signal maximized) |
| Protocol Consistency | High | Low (Enzyme lot variability) | High |
| Best Application | Robust, stable epitopes | Selected cytoplasmic/matrix antigens | Labile post-translational modifications |
Table 2: Impact of Retrieval Buffer pH on Common Phospho-Epitope Signal Intensity (H-Score Data)
| Target (p-) | pH 6.0 Citrate | pH 8.0 Tris-EDTA | pH 9.0 Borate | Optimal pH (from study) |
|---|---|---|---|---|
| ERK1/2 (T202/Y204) | 150 | 210 | 180 | 7.5 - 8.5 |
| AKT (S473) | 80 | 185 | 160 | 8.0 - 9.0 |
| STAT3 (Y705) | 200 | 165 | 190 | 6.0 - 7.0 & 9.0 |
| Histone H3 (S10) | 95 | 110 | 135 | 8.5 - 9.0 |
Detailed Experimental Protocols
Protocol 4.1: Optimized pH-Controlled Mild HIER for Phospho-Proteins
Materials: See Scientist's Toolkit below. Procedure:
- Deparaffinize and rehydrate FFPE tissue sections using standard xylene and ethanol series.
- Buffer Selection: Prepare 10mM retrieval buffer at the target pH (e.g., Tris-EDTA, pH 9.0; Citrate-EDTA, pH 6.5; or commercial high-pH buffer). Pre-heat buffer in a water bath or calibrated steamer to 95°C ± 2°C.
- Retrieval: Immerse slides in pre-heated buffer. Incubate for 15 minutes at 95°C. Do not boil vigorously.
- Cooling: Remove the container from heat and allow slides to cool in the buffer at room temperature for 20-30 minutes.
- Wash: Rinse slides in distilled water, then proceed to PBS wash.
- Immunostaining: Perform standard IHC/ICC blocking, primary antibody incubation (validated for phospho-specific detection), and detection steps.
Protocol 4.2: Validation Experiment: pH Titration for a Novel Phospho-Epitope
Aim: To determine the optimal HIER pH for a new anti-p-FAK (Y397) antibody. Method:
- Serial sections of a known FAK-positive/phosphorylated FFPE cell pellet block are prepared.
- Perform HIER as in Protocol 4.1, using a series of buffers across pH 6.0, 6.5, 7.0, 7.5, 8.0, 8.5, and 9.0.
- Stain all slides in a single automated IHC run with identical antibody concentration and detection conditions.
- Quantitative Analysis: Use digital pathology software to calculate the H-Score or similar quantitative measure for each slide.
- Controls: Include a no-primary antibody control and a slide with lambda phosphatase pre-treatment to confirm phospho-specificity.
Visualizations
Diagram 1: Workflow Comparison of Retrieval Methods
Diagram 2: Key p-AKT Signaling Pathway for IHC
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents for pH-Controlled Mild HIER Experiments
| Item | Function & Rationale |
|---|---|
| pH-Calibrated Buffer Kits (e.g., Tris-EDTA, Citrate, Borate) | Provides precise, reproducible pH control (6.0-10.0) for systematic epitope retrieval optimization. |
| Thermally Stable, Low-Binding Slide Rack | Ensures even heat distribution during mild HIER and prevents slide-to-slide variation. |
| Calibrated Water Bath or Digital Steamer | Maintains precise sub-boiling temperature (95°C ± 2°C) critical for mild retrieval. |
| Validated Phospho-Specific Primary Antibodies | Antibodies specifically raised and validated for IHC on FFPE tissue following retrieval. |
| Phosphatase Inhibitors (in staining buffer) | Added during antibody incubation to prevent endogenous phosphatase activity from stripping the epitope post-retrieval. |
| Lambda Protein Phosphatase | Essential negative control reagent to treat tissue sections pre-IHC to confirm phospho-signal specificity. |
| Digital Slide Scanner & Quantitative Pathology Software | Enables objective, high-throughput quantification of signal intensity (H-Score, % positivity) across pH conditions. |
Conclusion
The choice between HIER and PIER is not a matter of universal superiority but of context-specific optimization. HIER, with its broad applicability and controllability, is often the preferred first-line method for most targets, especially with modern pH-engineered buffers. PIER remains indispensable for certain tightly cross-linked epitopes, particularly in over-fixed tissues. The key takeaway is the necessity for empirical validation using controlled optimization grids tailored to the specific antibody-tissue system. Future directions point toward more refined, automated retrieval platforms and the development of epitope-specific retrieval cocktails. For researchers and drug developers, establishing a rigorously validated and documented antigen retrieval protocol is fundamental to achieving reproducible, high-quality IHC data, which underpins robust biomarker discovery and translational pathology.