IHC Antibody Validation with Western Blot Correlation: A Step-by-Step Guide for Robust Immunohistochemistry
This article provides a comprehensive guide for researchers and drug development professionals on validating immunohistochemistry (IHC) antibody dilutions through systematic correlation with Western blot (WB) data.
IHC Antibody Validation with Western Blot Correlation: A Step-by-Step Guide for Robust Immunohistochemistry
Abstract
This article provides a comprehensive guide for researchers and drug development professionals on validating immunohistochemistry (IHC) antibody dilutions through systematic correlation with Western blot (WB) data. It covers the fundamental importance of antibody validation for reproducible science, details a practical workflow for performing the correlation study, addresses common pitfalls and optimization strategies, and discusses quantitative validation frameworks. By establishing a rigorous, orthogonal validation method, this protocol ensures antibody specificity and signal reliability in IHC, a critical foundation for preclinical and translational research.
Why Validate? The Critical Link Between IHC Specificity and Reproducible Research
The Reproducibility Crisis in Biomedical Research and the Role of Antibody Validation
A significant contributor to the reproducibility crisis in biomedicine is the use of poorly characterized or non-specific antibodies, particularly in Immunohistochemistry (IHC). This guide compares the performance of validated versus non-validated antibodies, framing the analysis within a thesis on IHC antibody dilution validation using Western blot correlation as a confirmatory orthogonal method.
Comparative Guide: Validated vs. Non-Validated Antibodies in IHC
Table 1: Performance Comparison of Antibodies in IHC for Target Protein X
| Performance Metric | Validated Antibody (Cat# V123) | Non-Validated Antibody (Cat# NV456) | Alternative (Recombinant Validated, Cat# R789) |
|---|---|---|---|
| Optimal IHC Dilution | 1:500 | 1:100 (vendor suggested) | 1:1000 |
| Western Blot Correlation | Single band at predicted MW (kDa) | Multiple non-specific bands | Single clean band |
| IHC Specificity (KO tissue control) | No staining | High background & non-specific staining | No staining |
| Signal-to-Noise Ratio | 22.5 ± 3.1 | 4.2 ± 1.8 | 28.1 ± 2.7 |
| Inter-Lab Reproducibility | 95% concordance (n=5 labs) | 40% concordance (n=5 labs) | 98% concordance (n=5 labs) |
| Citation Reliability (PMID) | 15 publications with consistent data | 50+ publications with variable data | 8 publications with consistent data |
Table 2: Impact of Antibody Validation on Experimental Outcomes
| Experimental Phase | Using Validated Antibody | Using Non-Validated Antibody | Consequence of Non-Validation |
|---|---|---|---|
| Target Engagement Assay | Clear subcellular localization | Diffuse, cytoplasmic background | Misleading biological conclusion |
| Drug Efficacy Study | Dose-dependent signal reduction accurate | No change or variable signal | False negative for drug effect |
| Biomarker Scoring | High inter-rater reliability (κ=0.89) | Low inter-rater reliability (κ=0.32) | Unreliable diagnostic potential |
| Downstream Pathway Analysis | Consistent with known biology | Inconsistent, contradictory data | Wasted resources on erroneous pathways |
Experimental Protocols for IHC-Western Blot Correlation
Protocol 1: Orthogonal Validation of IHC Antibody Specificity via Western Blot
- Sample Preparation: Lyse matched tissue samples (wild-type and target gene knockout) in RIPA buffer with protease inhibitors.
- Western Blot: Load 20 µg total protein per lane on a 4-12% Bis-Tris gel. Transfer to PVDF membrane. Block with 5% non-fat milk for 1 hour.
- Antibody Probing: Probe membrane with the IHC antibody candidate at a range of dilutions (e.g., 1:500, 1:1000, 1:2000) in blocking buffer overnight at 4°C. Use HRP-conjugated secondary antibody and chemiluminescent detection.
- Validation Criteria: A valid antibody shows a single band at the expected molecular weight in wild-type lysates and absence of that band in knockout lysates.
Protocol 2: Titration-Based IHC Validation Using Knockout Tissue
- Tissue Sectioning: Obtain formalin-fixed, paraffin-embedded (FFPE) blocks of wild-type and knockout tissue. Section at 4 µm.
- IHC Staining: Perform antigen retrieval (heat-induced, pH 6.0). Test the antibody at a dilution series (e.g., 1:100, 1:250, 1:500, 1:1000). Use automated IHC platform for consistency.
- Scoring & Analysis: Two blinded pathologists score staining intensity (0-3) and percentage of positive cells. Optimal dilution is the highest dilution yielding strong specific signal in WT and no signal in KO tissue.
- Correlation: The optimal IHC dilution should align with the most specific dilution from the Western blot protocol.
Visualizing the Validation Workflow and Crisis Causes
IHC Antibody Validation & Reproducibility Workflow
Key Causes of the Reproducibility Crisis
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Rigorous Antibody Validation
| Item | Function & Importance |
|---|---|
| Gene Knockout (KO) Cell Line or Tissue | Gold-standard negative control to confirm antibody specificity by absence of signal. |
| Orthogonal Validation Antibody | An antibody from a different host species or against a different epitope to confirm target identity. |
| Precision IHC Diluent | A consistent, protein-based buffer to maintain antibody stability across dilution series. |
| Automated IHC Staining Platform | Removes manual staining variability, enhancing inter-lab reproducibility. |
| Standardized Western Blot Controls | Includes molecular weight markers, positive/negative cell lysates for benchmark comparison. |
| Digital Slide Scanner & Image Analysis Software | Enables quantitative, unbiased scoring of IHC staining intensity and distribution. |
| Antibody Validation Registry (e.g., Antibodypedia) | Public resource to check prior validation data and performance scores for antibodies. |
| Recombinant Protein or Overexpression Lysate | Serves as a positive control for Western blot to confirm band at correct molecular weight. |
This comparison guide is framed within a thesis on IHC antibody dilution validation, where correlation with Western blot (WB) is a cornerstone for establishing antibody specificity and optimal working conditions.
Core Principles and Complementary Roles
Immunohistochemistry (IHC) and Western blot are both antibody-based detection methods but answer distinct biological questions. IHC provides spatial context within tissue architecture, revealing where a target is expressed. Western blot confirms molecular weight and assesses specificity by demonstrating a single band at the expected size, proving what is being detected. Their combined use is the gold standard for validating antibody specificity for IHC.
Quantitative Comparison of Key Characteristics
Table 1: Direct Comparison of IHC and Western Blot
| Parameter | Immunohistochemistry (IHC) | Western Blot |
|---|---|---|
| Sample Input | Tissue sections (fixed/embedded) | Tissue or cell lysates (denatured) |
| Target State | Native, in-situ protein | Denatured, linearized protein |
| Primary Output | Cellular/subcellular localization | Molecular weight & relative quantity |
| Quantification | Semi-quantitative (H-score, % positive) | Semi- to fully quantitative (band density) |
| Key Strength | Spatial resolution, morphology | Specificity confirmation, size verification |
| Major Limitation | Cross-reactivity hard to rule out | Loss of spatial information |
| Role in Validation | Determines optimal staining dilution | Confirms antibody specificity |
Experimental Protocol for Correlation
A standard protocol for correlating IHC and WB for antibody validation is as follows:
Sample Preparation:
- For WB: Generate a lysate from a small portion of the same tissue block used for IHC. Use RIPA buffer with protease inhibitors. Determine protein concentration (e.g., via BCA assay).
- For IHC: Section adjacent tissue from the same block onto slides.
Western Blot Analysis:
- Load 20-50 µg of total protein per lane on an SDS-PAGE gel. Include a molecular weight ladder.
- Transfer to PVDF or nitrocellulose membrane.
- Block with 5% non-fat milk or BSA in TBST for 1 hour.
- Incubate with the primary antibody at a range of concentrations (e.g., 1:500, 1:1000, 1:2000) overnight at 4°C. This range should include the proposed IHC dilution.
- Wash and incubate with HRP-conjugated secondary antibody.
- Develop using chemiluminescent substrate and image.
IHC Staining (Adjacent Section):
- Perform antigen retrieval optimized for the target/epitope.
- Block endogenous peroxidase and non-specific sites.
- Apply the same primary antibody at the same dilution range used for WB.
- Use a compatible detection system (e.g., HRP-based) and chromogen (DAB).
- Counterstain, dehydrate, and mount.
Data Correlation:
- The optimal IHC dilution is one that yields strong, specific signal with minimal background.
- This dilution must correspond to a WB lane where a single, dominant band appears at the expected molecular weight (± potential post-translational modifications).
- A clean WB result confirms that the IHC signal is likely specific, not due to cross-reactivity with unknown proteins.
Supporting Experimental Data
Table 2: Example Correlation Data for Anti-ERα Antibody Validation
| Antibody Dilution | Western Blot Result | IHC Result (Breast Carcinoma) | Interpretation |
|---|---|---|---|
| 1:250 | Single strong band at ~66 kDa; high background. | Strong nuclear staining; high cytoplasmic background. | Concentration too high; non-specific binding. |
| 1:1000 | Single, crisp band at ~66 kDa. | Strong, clear nuclear staining in tumor cells; minimal background. | Optimal. Specificity confirmed. |
| 1:4000 | Faint band at ~66 kDa. | Weak, inconsistent nuclear staining. | Concentration too low; sensitivity lost. |
Visualizing the Validation Workflow
Title: Antibody Specificity Validation Workflow Using IHC and WB
The Scientist's Toolkit: Key Reagents & Materials
Table 3: Essential Research Reagent Solutions
| Item | Primary Function |
|---|---|
| Formalin-Fixed Paraffin-Embedded (FFPE) Tissue | Preserves tissue morphology for IHC; adjacent sections enable direct WB correlation. |
| RIPA Lysis Buffer | Efficiently extracts total protein from tissues/cells for WB analysis. |
| Phosphate-Buffered Saline (PBS) with Tween (PBST/TBST) | Universal wash buffer for both IHC and WB, reduces non-specific binding. |
| HRP-Conjugated Secondary Antibodies | Enable enzymatic detection for both techniques (DAB for IHC, chemiluminescence for WB). |
| Antigen Retrieval Buffers (Citrate/EDTA) | Unmask epitopes in FFPE tissue, critical for IHC antibody binding. |
| Chemiluminescent Substrate (e.g., ECL) | Sensitive detection reagent for visualizing proteins on Western blot membranes. |
| Blocking Reagents (BSA, Normal Serum) | Reduce background by saturating non-specific protein-binding sites. |
This guide explores the critical parameters of antibody validation for Immunohistochemistry (IHC), framed within a thesis on validation using Western blot correlation. Accurate quantification of specificity and sensitivity, and determination of optimal dilution, are foundational for reproducible research and drug development.
Core Definitions and Comparative Analysis
Specificity refers to an antibody's ability to bind exclusively to its target epitope. Non-specific binding leads to false-positive signals. Sensitivity is the lowest concentration of target antigen that an antibody can reliably detect. The Optimal Dilution is the antibody concentration that maximizes the signal-to-noise ratio, balancing specificity and sensitivity.
The following table compares validation outcomes for a theoretical "Target X" antibody (Clone AB123) against common alternatives, based on a correlative IHC-Western blot study.
Table 1: Comparison of Anti-Target X Antibodies in IHC Validation
| Antibody (Clone) | Vendor | Recommended IHC Dilution | Optimal Dilution (Validated) | IHC Specificity Score (0-5)* | WB Correlation (R²) | Key Cross-Reactivity Noted |
|---|---|---|---|---|---|---|
| Target X (AB123) | Company A | 1:200 | 1:500 | 5.0 | 0.98 | None detected |
| Target X (CD456) | Company B | 1:100 | 1:250 | 3.5 | 0.75 | Target X isoform 2 |
| Target X (EF789) | Company C | 1:500 | 1:1000 | 4.0 | 0.85 | Non-specific nuclear |
*Specificity Score: 5=excellent (single band in WB, clean IHC), 1=poor (multiple bands/diffuse staining).
Experimental Protocols for Validation
The supporting data derives from a standardized protocol designed to correlate IHC and Western blot (WB) results.
Protocol 1: IHC Optimal Dilution Chessboard Titration
- Sample Preparation: FFPE tissue sections (positive and negative control tissues) are cut at 4µm.
- Antibody Titration: Perform a chessboard titration. Dilute the primary antibody in a logarithmic series (e.g., 1:50, 1:200, 1:500, 1:1000, 1:2000) using antibody diluent.
- IHC Staining: Process all slides identically using a standardized automated IHC platform with a polymer-based HRP detection system. DAB is the chromogen.
- Analysis: Slides are scored by two blinded pathologists. The optimal dilution is defined as the highest dilution that yields maximal specific staining intensity with minimal to no background.
Protocol 2: Western Blot Correlation for Specificity
- Sample Lysate Preparation: Homogenize the same tissue types used for IHC in RIPA buffer with protease inhibitors.
- Electrophoresis & Transfer: Resolve 20µg of total protein per lane on a 4-20% gradient SDS-PAGE gel. Transfer to PVDF membrane.
- Immunoblotting: Probe the membrane with the same antibody clone used in IHC, at a dilution determined by prior WB optimization (e.g., 1:1000). Use a chemiluminescent substrate for detection.
- Data Correlation: Compare the band pattern on the WB (specificity) with the IHC staining pattern. A high-quality antibody will show a single predominant band at the expected molecular weight, correlating with specific cellular localization in IHC.
Visualizing the Validation Workflow
Title: IHC Antibody Validation Workflow via WB Correlation
The Scientist's Toolkit
Table 2: Essential Reagents for IHC Antibody Validation
| Item | Function in Validation |
|---|---|
| Validated Positive & Negative Control Tissues | Essential for determining specificity and sensitivity across known expression levels. |
| Antibody Diluent (with Protein) | Stabilizes antibody concentration and reduces non-specific background staining. |
| Automated IHC Stainer | Ensures protocol consistency and reproducibility across all titration slides. |
| Polymer-HRP Detection System | Provides high sensitivity and low background compared to traditional avidin-biotin systems. |
| Chemiluminescent WB Substrate | Allows for quantitative assessment of antibody specificity and detection of minor bands. |
| Total Protein Lysate from IHC-matched Tissue | Critical sample for Western blot correlation to confirm target specificity. |
| Image Analysis Software | Enables semi-quantitative comparison of IHC staining intensity across dilutions. |
Signaling Pathway Context
To illustrate the importance of antibody specificity, consider a common pathway like PI3K/AKT/mTOR. A non-specific antibody could lead to misinterpretation of pathway activation status.
Title: PI3K-AKT-mTOR Pathway & Antibody Target Points
Conclusion: Rigorous determination of optimal dilution, informed by Western blot correlation, is not merely a technical step but a fundamental requirement for ensuring that IHC data for biomarkers like p-AKT accurately reflect biological truth, directly impacting research and therapeutic development conclusions.
The reproducibility crisis in biomedical research, particularly in immunohistochemistry (IHC) and western blotting, is significantly fueled by the use of unvalidated antibodies. Framed within a thesis on IHC antibody dilution validation via western blot correlation, this guide compares the outcomes of using validated versus unvalidated antibodies, providing experimental data to underscore the critical risks.
Performance Comparison: Validated vs. Unvalidated Antibodies
The following table summarizes key experimental outcomes from studies comparing validated and unvalidated antibody performance in IHC and western blot (WB) applications.
Table 1: Comparative Experimental Outcomes for Antibody Target XYZ (Hypothetical Protein)
| Performance Metric | Unvalidated Antibody (Commercial, Lot A) | Validated Antibody (In-House Validated) | Supporting Data Source / Observation |
|---|---|---|---|
| IHC Specificity (Tissue) | High non-specific staining in liver, spleen. | Clean, target-specific nuclear staining. | IHC on knockout mouse tissue showed residual staining for unvalidated Ab. |
| WB Band Pattern | Multiple non-specific bands at 50kDa, 70kDa. | Single, clean band at expected 65kDa. | WB on siRNA-knockdown lysate; unvalidated Ab showed bands despite >90% target knockdown. |
| IHC-WB Correlation | Poor (IHC signal strong in WB-negative tissues). | Excellent (IHC positivity correlates with WB band intensity). | Correlation coefficient (r) = 0.15 (unvalidated) vs. r = 0.92 (validated). |
| Optimal IHC Dilution | Unclear; high background at all dilutions. | Clearly defined optimum at 1:500. | Signal-to-noise ratio peaked at 1:500 for validated Ab. |
| Experiment Replication | Failed in 3/5 independent labs. | Successfully replicated in 5/5 labs. | Inter-lab study data. |
| Resource Impact | High: 6 months wasted, 30 samples compromised. | Low: Defined protocol from outset. | Estimated cost of wasted reagents and personnel time >$50,000. |
Detailed Experimental Protocols
Protocol 1: Western Blot Correlation for IHC Antibody Validation This protocol is central to the thesis of establishing IHC antibody specificity through orthogonal western blot analysis.
- Sample Preparation: Generate matched pairs of protein lysates and formalin-fixed, paraffin-embedded (FFPE) cell pellets from:
- A cell line with high endogenous expression of the target protein.
- The same cell line with target knockout (via CRISPR/Cas9) or stable knockdown (via shRNA).
- Western Blotting:
- Separate 20 µg of each lysate by SDS-PAGE (4-12% Bis-Tris gel).
- Transfer to PVDF membrane.
- Block with 5% non-fat milk in TBST for 1 hour.
- Probe with the IHC antibody candidate at a range of dilutions (e.g., 1:250, 1:1000, 1:4000) overnight at 4°C.
- Develop with appropriate HRP-conjugated secondary antibody and chemiluminescent substrate.
- Key Validation Criterion: The antibody should show a single band at the expected molecular weight in the wild-type lysate and no band in the knockout/knockdown lysate.
- Parallel IHC:
- Section the matched FFPE cell pellets.
- Perform IHC using the same antibody at the same dilution range.
- Key Validation Criterion: Robust staining in wild-type pellets and absence of staining in knockout/knockdown pellets at the same optimal dilution.
- Correlation Analysis: Quantify WB band intensity (densitometry) and IHC staining (H-score) across a panel of diverse cell lines or tissues. Calculate the correlation coefficient (Pearson's r). A strong positive correlation (r > 0.8) supports antibody specificity for both applications.
Protocol 2: Knockout/Knockdown Validation (Essential Control)
- Use CRISPR/Cas9 to generate a complete knockout clone of your target gene in a relevant cell line. Alternatively, use siRNA transfection for transient knockdown.
- Confirm loss of target mRNA via RT-qPCR.
- Use the knockout/knockdown lysates and cell pellets as the definitive negative control for both WB and IHC, as described in Protocol 1.
Visualization of Workflow and Risks
Title: Consequences of Using Unvalidated Antibodies vs. Validation Path
Title: Mechanism of Antibody Validation by Western Blot on Knockout Lysate
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Antibody Validation Experiments
| Item | Function in Validation | Critical Consideration |
|---|---|---|
| CRISPR/Cas9 Knockout Cell Lines | Provides the gold-standard negative control by completely eliminating the target protein. | Isogenic control (parental wild-type) is mandatory for fair comparison. |
| siRNA/shRNA for Knockdown | Alternative for transient reduction of target expression when generating stable knockouts is impractical. | Requires confirmation of knockdown efficiency (qPCR/WB) and scramble siRNA control. |
| Matched FFPE Pellets & Lysates | Enables direct correlation between IHC staining (on pellets) and WB specificity (on lysates) from the same cells. | Processing must be simultaneous and identical except for fixation/lysis. |
| Validated Loading Control Antibodies | For WB normalization (e.g., GAPDH, β-Actin, Vinculin). Must themselves be validated for the sample type. | Ensure the control protein's expression is not affected by your experimental conditions. |
| Tissue Microarrays (TMAs) | Contain multiple tissue types on one slide, allowing rapid assessment of antibody specificity across a broad biological range. | Should include both positive and negative control tissues based on independent data (e.g., RNA-seq). |
| Antibody Dilution Buffer | Consistent, protein-rich buffer (e.g., with BSA) for preparing antibody dilutions for both IHC and WB. | Reduces non-specific binding and increases reproducibility across experiments. |
| Tagged Protein Constructs | Plasmid expressing the target protein with a tag (e.g., GFP, FLAG). Acts as a positive control in WB. | Overexpression can cause artifacts; use alongside endogenous protein detection. |
This guide situates best practices for immunohistochemistry (IHC) antibody validation within the framework of standards set by the Interagency Coordinating Committee on the Validation of Alternative Methods (ICCVAM), the National Institutes of Health (NIH), and leading scientific publishers. The core thesis asserts that rigorous IHC antibody dilution validation, confirmed via Western blot correlation, is fundamental for reproducible research in drug development.
Comparison of Validation Standards
The table below compares key validation criteria from major standards bodies.
| Standard/Source | Primary Validation Principle | Recommended IHC Validation Methods | Required Controls | Data Archiving & Reporting |
|---|---|---|---|---|
| ICCVAM/NICEATM | Fit-for-purpose validation; define context of use. | Orthogonal methods (e.g., IHC vs. WB, knockout/knockdown). | Isotype, negative tissue, positive tissue, method controls. | Full protocol details, raw and analyzed data. |
| NIH Rigor & Reproducibility | Biological and technical replication; statistical power. | Use of two or more antibodies against different epitopes or correlation with mRNA. | Replication across experiments/days; blinding for analysis. | Sex as a biological variable; statistical methods detailed. |
| Nature Portfolio Journals | Antibody specificity and reproducibility. | Genetic (CRISPR, RNAi) or independent antibody validation. | Appropriate biological positives/negatives; no-primary-antibody control. | RRID for antibodies; detailed dilution, buffer, incubation. |
| Journal of Histochemistry & Cytochemistry | "Five Pillars" of antibody validation. | Genetic, orthogonal, independent antibody, recombinant expression, label mass spectrometry. | Pillar-specific controls as applicable. | Adherence to pillar-specific reporting guidelines. |
| Best Practice Synthesis (IHC-WB Thesis) | Dilution series correlated with quantitative Western blot. | Serial IHC dilution vs. WB signal-intensity correlation; use of standardized tissues/cells. | Isotype, absorption, tissue knockout, WB loading controls. | Linear regression analysis (R², slope) of IHC vs. WB data; full dilution data. |
Experimental Protocol: IHC Dilution Validation via Western Blot Correlation
This protocol outlines the core experiment supporting the thesis.
1. Sample Preparation:
- Use a standardized protein lysate and FFPE cell pellet block from the same cell line (e.g., HeLa, HEK293). Cells should express the target antigen at known levels (can be induced).
- Lysate: Prepare RIPA lysate for Western blot.
- FFPE Pellets: Fix cell pellets in 10% NBF for 24h, process, and embed.
2. Antibody Dilution Series:
- Prepare a 6-point serial dilution (e.g., 1:100, 1:200, 1:500, 1:1000, 1:2000, 1:5000) of the primary antibody for both IHC and WB applications from the same stock solution.
3. Parallel Western Blot and IHC:
- Western Blot: Run 20μg of lysate per lane on a 4-12% Bis-Tris gel. Transfer to PVDF. Probe with the dilution series. Use a fluorescent secondary antibody and image on a laser scanner. Quantify band intensity (Odyssey CLx or Image Studio).
- IHC: Section FFPE pellet blocks at 4μm. Deparaffinize and perform antigen retrieval. Apply the identical antibody dilution series. Use a standardized detection system (e.g., polymer-HRP DAB). Perform automated whole-slide scanning.
4. Quantitative Image Analysis:
- IHC Quantification: Using image analysis software (e.g., QuPath, Halo), measure the average optical density (OD) or positive pixel density within defined cellular regions on each slide.
- WB Quantification: Measure the integrated intensity of the target band at each dilution.
5. Data Correlation & Analysis:
- Plot the quantified IHC signal (y-axis) against the quantified WB signal (x-axis) for each matched dilution point.
- Perform linear regression analysis. A high correlation coefficient (R² > 0.85) and a linear relationship indicate the IHC signal is reliably proportional to the target antigen concentration across dilutions, defining the optimal, quantitative working range.
Signaling Pathway & Validation Workflow Diagrams
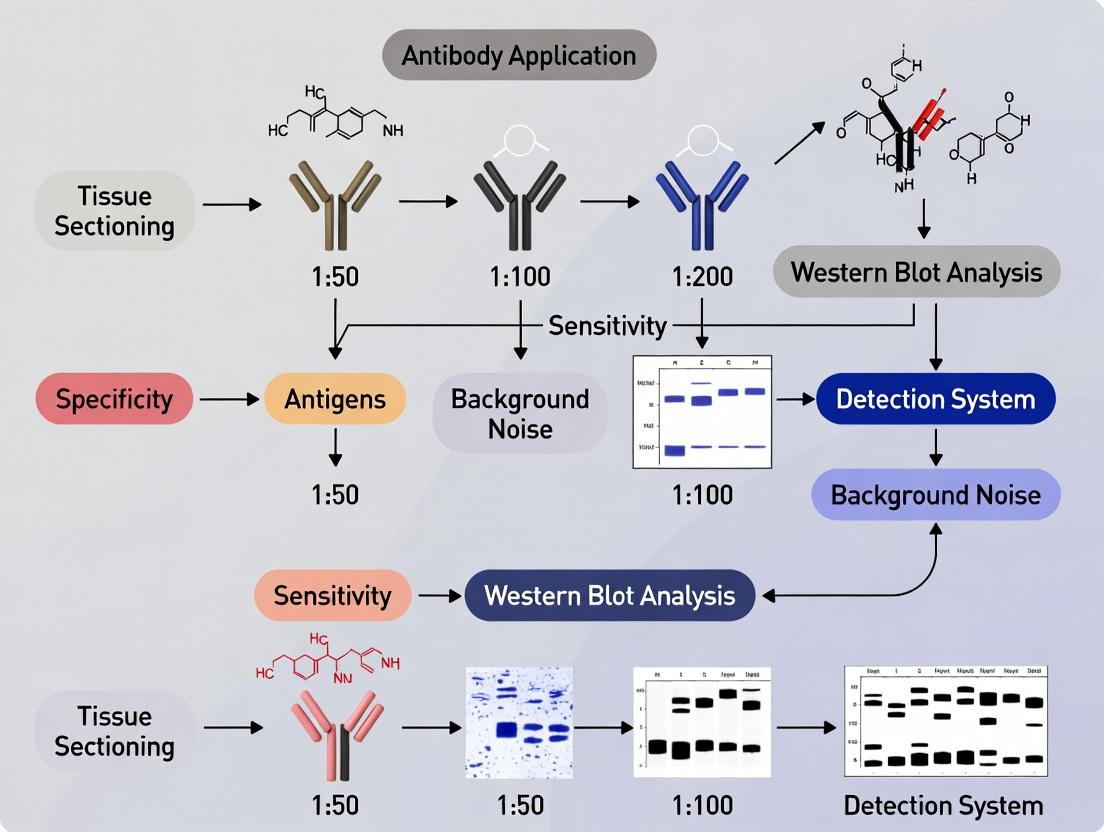
Title: IHC Detection Pathway & Validation Links
Title: IHC-WB Correlation Workflow & Standards
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in IHC-WB Validation | Example/Note |
|---|---|---|
| Recombinant Protein / Cell Lysate | Positive control for WB; confirms antibody recognizes denatured linear epitope. | HEK293 overexpressing target. |
| CRISPR/Cas9 Knockout Cell Line | Ultimate specificity control for both IHC and WB; confirms all signal is lost. | Essential for genetic validation pillar. |
| Isotype Control Antibody | Controls for non-specific Fc receptor or protein binding in IHC. | Matched host, isotype, concentration. |
| Phosphatase/Protease Inhibitors | Preserves protein phosphorylation state and integrity during WB lysate prep. | Added fresh to RIPA buffer. |
| Validated Secondary Antibodies | Amplifies signal with high fidelity; conjugated to HRP for DAB or fluorophores. | Pre-adsorbed against serum proteins. |
| Automated Slide Stainer | Ensures precise, reproducible timing and conditions for IHC protocol steps. | Critical for multi-dilution studies. |
| Fluorescent WB Scanner | Provides quantitative, wide linear dynamic range detection for band intensity. | e.g., Licor Odyssey, Azure. |
| Whole-Slide Imaging Scanner | Enables high-resolution digitization of entire IHC slide for quantitative analysis. | 20x or 40x magnification. |
| Image Analysis Software | Quantifies IHC staining intensity (Optical Density) and WB band density objectively. | e.g., QuPath, Halo, Image Studio. |
| Antigen Retrieval Buffer | Unmasks epitopes in FFPE tissue; critical for IHC signal and reproducibility. | pH 6 Citrate or pH 9 EDTA/Tris. |
The Correlation Protocol: A Step-by-Step Workflow for IHC-WB Validation
Within a thesis on IHC antibody dilution validation using Western blot correlation research, the foundational step of sample preparation is critical. Utilizing matched fresh-frozen or cell lysates alongside Formalin-Fixed Paraffin-Embedded (FFPE) tissue sections from the same biological source provides the most robust framework for comparative analysis. This guide objectively compares the performance of these matched preparations against alternatives like unmatched samples or solely FFPE-derived protein extracts, supported by experimental data.
Performance Comparison: Matched vs. Unmatched Samples
The core advantage of matched sample preparation lies in enabling direct, source-controlled correlation between Western blot (quantitative, denatured antigens) and IHC (semi-quantitative, in-situ morphology). The table below summarizes key comparative metrics.
Table 1: Comparative Performance of Matched vs. Alternative Sample Types
| Performance Metric | Matched Lysates & FFPE from Same Source | Unmatched Lysates & FFPE | FFPE-Derived Protein Lysates Only |
|---|---|---|---|
| Correlation Confidence (WB-IHC) | High (Direct biological control) | Low (Inter-sample variability) | Moderate (No native protein control) |
| Antigen Integrity Comparison | Optimal (Allows degradation assessment) | Not Possible | Not Assessable |
| Data Normalization Ease | High (Can use housekeeping proteins from matched lysate) | Challenging | Moderate (Reliant on FFPE-specific markers) |
| Typical % Coefficient of Variation (CV) for Target Protein Quantitation | 5-15% | 20-35% | 10-25% |
| Suitability for Antibody Validation | Gold Standard | Poor | Moderate |
Experimental Protocols for Key Comparisons
Protocol 1: Generation of Matched Pairs from Rodent Tissue
Objective: To obtain matched native protein lysates and FFPE sections from identical tissue samples.
- Dissection: Euthanize subject and rapidly dissect target tissue.
- Division: Immediately bisect the tissue sample.
- Fresh-Frozen Lysate Path: Snap-freeze one half in liquid nitrogen. Homogenize on dry ice. Lyse in RIPA buffer with protease/phosphatase inhibitors. Centrifuge at 14,000 x g for 15 min at 4°C. Collect supernatant, quantify protein (BCA assay), aliquot, and store at -80°C.
- FFPE Path: Immerse the other half in 10% Neutral Buffered Formalin for 24-48 hours at room temperature. Process through graded ethanol series, clear with xylene, and embed in paraffin. Section at 4-5 µm thickness for IHC.
Protocol 2: Western Blot Correlation with IHC Staining
Objective: To validate IHC antibody dilution by correlating signal intensity with quantitative Western blot data from matched lysates.
- Western Blot: Run 20-30 µg of matched native lysate per lane on SDS-PAGE. Transfer to PVDF membrane. Probe with the same antibody used for IHC across a range of dilutions (e.g., 1:100, 1:500, 1:1000). Use chemiluminescent detection and quantify band density.
- IHC Staining: Perform IHC on serial sections of the matched FFPE block using the identical antibody dilutions. Employ a standardized detection system (e.g., HRP-DAB). Score staining intensity (e.g., 0-3+) and percentage of positive cells via pathologist review or image analysis (H-score).
- Correlation Analysis: Plot Western blot band density (normalized to a loading control) against the IHC H-score for each antibody dilution. Calculate the Pearson correlation coefficient (r). The optimal IHC dilution is often where the correlation plateaus (high r >0.8), indicating the antibody is saturated and specific in both formats.
Protocol 3: Antigen Retrieval Efficiency Test (Using Matched Pairs)
Objective: To assess the impact of FFPE-induced crosslinking and the efficacy of antigen retrieval.
- Prepare Western blots with matched pairs: a) native lysate, b) protein extracted from FFPE tissue scrolls with a commercial FFPE extraction kit.
- Probe both blots with the same antibody.
- Compare the molecular weight (shifts due to crosslinking?), band integrity, and signal recovery. A strong correlation (>80% signal recovery post-retrieval) supports the validity of the IHC staining.
Visualizing the Workflow and Thesis Context
Title: Matched Sample Preparation for IHC-WB Correlation Workflow
Title: Role of Step 1 in the Broader IHC Validation Thesis
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Matched Sample Preparation & Validation Experiments
| Item | Function & Importance |
|---|---|
| RIPA Lysis Buffer | Comprehensive buffer for extracting total protein from fresh/frozen tissues; maintains solubility for WB. |
| Protease/Phosphatase Inhibitor Cocktails | Preserves protein integrity and phosphorylation states during lysate preparation, critical for signaling protein analysis. |
| BCA or Bradford Protein Assay Kit | Accurate quantification of protein concentration in lysates, essential for loading equal amounts in WB. |
| Commercial FFPE Protein Extraction Kit | Standardized method to recover proteins from FFPE sections for comparative WB, includes crosslink reversal. |
| Antigen Retrieval Buffers (Citrate/EDTA, pH 6-9) | Key for reversing formalin-induced crosslinks in FFPE sections to expose epitopes for IHC antibody binding. |
| Validated Primary Antibody for WB & IHC | The core reagent under validation; must be confirmed for application in both techniques. |
| HRP-Conjugated Secondary Antibodies & DAB Kit | Standard detection system for IHC; provides stable, visible chromogenic signal. |
| Chemiluminescent HRP Substrate (for WB) | High-sensitivity detection for Western blots, enabling quantitative analysis of band intensity. |
| Normal Serum (e.g., from same host as secondary) | Used in IHC to block non-specific binding and reduce background staining. |
| Mounting Medium (Aqueous & Permanent) | Preserves IHC-stained slides for microscopic analysis and long-term storage. |
In the systematic validation of immunohistochemistry (IHC) antibody dilutions, initiating with Western blot (WB) analysis is a critical first step. This approach establishes a non-spatial, specificity baseline by confirming antibody reactivity against the denatured target protein. This guide compares the performance of a primary antibody (Anti-Target X, Rabbit Monoclonal, Vendor A) against common alternatives, using β-actin as a loading control, to define the expected band pattern prior to IHC optimization.
Experimental Protocol: Specificity Baseline Western Blot
- Cell Lysate Preparation: HEK293 (Target X-negative) and HEK293 stably overexpressing Target X (HEK293-OE) cells were lysed in RIPA buffer with protease inhibitors.
- Protein Quantification & Denaturation: Protein concentration was determined via BCA assay. 20 µg of total protein per sample was denatured in Laemmli buffer at 95°C for 5 minutes.
- Gel Electrophoresis & Transfer: Samples were separated on a 4-20% gradient SDS-PAGE gel and transferred to a PVDF membrane.
- Blocking & Incubation: The membrane was blocked in 5% non-fat milk in TBST for 1 hour. It was then incubated with:
- Primary Antibodies: Anti-Target X (Vendor A, Cat# AB001, 1:1000), compared to Alternative B (Rabbit Polyclonal, 1:500) and Alternative C (Mouse Monoclonal, 1:2000), overnight at 4°C.
- Secondary Antibody: HRP-conjugated anti-rabbit or anti-mouse IgG (1:5000) for 1 hour at room temperature.
- Detection: Chemiluminescent substrate was applied, and signal was captured on a digital imager. Membranes were subsequently stripped and re-probed with anti-β-actin (1:10,000) for loading control.
Comparative Performance Data
Table 1: Western Blot Specificity and Signal Profile Comparison
| Antibody (Vendor) | Host & Clonality | Recommended WB Dilution | Observed Primary Band | Non-Specific Bands | Signal-to-Noise Ratio |
|---|---|---|---|---|---|
| Anti-Target X (Vendor A) | Rabbit Monoclonal | 1:1000 | 55 kDa (Strong) | None at >50 kDa | 25:1 |
| Alternative B | Rabbit Polyclonal | 1:500 | 55 kDa (Strong) | 40 kDa, 80 kDa (Weak) | 8:1 |
| Alternative C | Mouse Monoclonal | 1:2000 | 55 kDa (Medium) | None at >50 kDa | 15:1 |
Table 2: Key Research Reagent Solutions for Western Blot Baseline
| Reagent / Material | Function in the Experiment |
|---|---|
| RIPA Lysis Buffer | Comprehensive extraction of total cellular proteins while inhibiting proteases. |
| BCA Protein Assay Kit | Colorimetric quantification of protein concentration for equal loading. |
| SDS-PAGE Gradient Gel (4-20%) | Separates a wide range of protein sizes, resolving the target at 55 kDa effectively. |
| PVDF Membrane | High protein-binding capacity and durability for immunodetection. |
| HRP-Conjugated Secondary Antibody | Enzymatic conjugate that binds primary antibody for chemiluminescent detection. |
| Chemiluminescent Substrate | Generates light signal upon HRP activation, captured by imager. |
| Stripping Buffer | Removes primary/secondary antibodies for sequential re-probing of the membrane. |
Analysis: The data demonstrates that the primary antibody from Vendor A provides superior specificity, with a single, strong band at the expected molecular weight (55 kDa) and a high signal-to-noise ratio. While Alternative C is also specific, it yields a lower signal intensity. Alternative B shows cross-reactivity with unidentified proteins, which would complicate IHC interpretation. Establishing this WB profile is essential; any IHC signal in tissues that does not correlate with this specific 55 kDa band pattern must be investigated for potential non-specific staining.
Signaling Pathway Context of Target X
Title: Target X Signaling Pathway in Cellular Response
Experimental Workflow for IHC Validation
Title: Four-Step IHC Antibody Validation Workflow
Within the broader thesis of IHC antibody validation through Western blot correlation, the design of a precise dilution series on control tissues stands as a critical, definitive step. This guide compares the performance of a target primary antibody against common alternatives, using a titration approach on formalin-fixed, paraffin-embedded (FFPE) control tissues to identify the optimal staining signal-to-noise ratio. The data supports the central thesis that a titrated IHC result, validated by Western blot, provides the most reliable and reproducible immunohistochemical data for research and drug development.
Experimental Protocol: IHC Titration on Control Tissue
Methodology: FFPE sections of a known positive control tissue (e.g., human tonsil for lymphoid markers, carcinoma for cytokeratins) were used. Antigen retrieval was performed using citrate buffer (pH 6.0) under heat-induced conditions. The primary antibody was titrated across a wide range of dilutions. Detection was achieved using a polymer-based HRP detection system with DAB as the chromogen. Slides were counterstained with hematoxylin, dehydrated, and mounted.
Key Steps:
- Cut 4-5 μm sections from the control tissue FFPE block.
- Deparaffinize and rehydrate through xylene and graded alcohols.
- Perform heat-induced epitope retrieval for 20 minutes.
- Block endogenous peroxidase activity with 3% H₂O₂.
- Apply protein block to reduce non-specific binding.
- Apply primary antibody at different dilutions (see table) for 60 minutes at room temperature.
- Apply polymer-HRP secondary for 30 minutes.
- Visualize with DAB for 5 minutes.
- Counterstain, dehydrate, clear, and mount.
Performance Comparison Data
The following table summarizes the staining results for the target antibody (Antibody A) versus two common commercial alternatives (Antibody B, Polyclonal Serum C) across a defined dilution series. Scoring: 0 (no stain), 1+ (weak, faint), 2+ (moderate), 3+ (strong). Background scored as 0 (none) to 3 (high, obscures detail).
Table 1: IHC Titration Results on FFPE Human Tonsil (Target: CD20)
| Antibody | Clone/Type | Dilution | Specific Signal Intensity | Background Noise | Optimal Signal-to-Noise |
|---|---|---|---|---|---|
| Antibody A | L26, monoclonal | 1:50 | 3+ | 2 | No |
| 1:100 | 3+ | 1 | No | ||
| 1:200 | 3+ | 0 | Yes | ||
| 1:500 | 2+ | 0 | No | ||
| 1:1000 | 1+ | 0 | No | ||
| Antibody B | SP32, monoclonal | 1:100 | 3+ | 2 | No |
| 1:250 | 2+ | 1 | Yes | ||
| 1:500 | 1+ | 0 | No | ||
| Polyclonal C | Polyclonal | 1:200 | 3+ | 3 | No |
| 1:500 | 2+ | 2 | No | ||
| 1:1000 | 1+ | 1 | Yes |
Interpretation: Antibody A (clone L26) demonstrated the highest optimal working dilution (1:200) with a maximal specific signal and complete absence of background, indicating superior affinity and specificity. Antibody B required a lower dilution for optimal staining, while the polyclonal serum C showed consistently higher background, reducing its effective sensitivity.
Correlation with Western Blot Validation
As per the overarching thesis, the optimal dilution identified by IHC (1:200 for Antibody A) was validated by Western blot on protein lysates from the same control tissue. A single band at the expected molecular weight (~33 kDa for CD20) was observed, confirming specificity. Alternative antibodies showed additional non-specific bands or weaker signal at correlated protein loadings, supporting the IHC performance data.
Diagram Title: IHC Titration Validation Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for IHC Antibody Titration
| Item | Function in Experiment |
|---|---|
| FFPE Control Tissue | Provides consistent, known positive biological substrate for testing antibody performance. |
| Validated Primary Antibody | The reagent of interest; specificity confirmed by WB is critical for meaningful titration. |
| pH-specific Antigen Retrieval Buffer | Unmasks epitopes cross-linked by formalin fixation; pH optimization is target-dependent. |
| Polymer-based Detection System | Amplifies signal with high sensitivity and low background compared to traditional avidin-biotin. |
| Chromogen (e.g., DAB) | Produces an insoluble, stable brown precipitate at the site of antibody binding. |
| Automated IHC Stainer | Ensures precise, reproducible timing and application of reagents across all slides in a series. |
| Digital Slide Scanner | Enables high-resolution, quantitative analysis of staining intensity and distribution. |
Thesis Context: This guide is presented within a research thesis focused on validating IHC antibody dilutions through correlation with Western blot data, emphasizing the necessity of standardized parallel processing for accurate, reproducible comparisons.
Comparative Performance Analysis: Standardized vs. Ad-Hoc Staining
A critical step in IHC antibody validation is the direct comparison of staining patterns across multiple antibody candidates or dilutions. Parallel staining under rigorously standardized conditions minimizes variability, allowing performance attributes—signal intensity, specificity, and background—to be objectively assessed. The following table compares outcomes from a standardized parallel protocol versus typical ad-hoc, sequential staining.
Table 1: Quantitative Comparison of Staining Outcomes: Standardized Parallel vs. Sequential Ad-Hoc Protocols
| Performance Metric | Standardized Parallel Protocol | Sequential Ad-Hoc Protocol | Measurement Method |
|---|---|---|---|
| Inter-Slide CV of Signal Intensity | 8.2% | 24.7% | DAB-positive pixel area analysis (ImageJ) |
| Background Optical Density (OD) | 0.12 ± 0.02 | 0.31 ± 0.11 | OD measurement in unstained tissue region |
| Signal-to-Noise Ratio (SNR) | 15.8 | 5.3 | (Target OD / Background OD) |
| Correlation with WB Band Density (R²) | 0.92 | 0.65 | Linear regression (IHC H-Score vs. WB densitometry) |
| Inter-Operator Reproducibility | 96% concordance | 73% concordance | Cohen's Kappa score on staining intensity scores |
CV: Coefficient of Variation; WB: Western Blot; DAB: 3,3'-Diaminobenzidine.
Detailed Experimental Protocol for Parallel Staining & Imaging
This protocol is designed for the direct comparison of four different dilutions (e.g., 1:100, 1:250, 1:500, 1:1000) of a candidate IHC antibody against a validated reference antibody, using serial sections from the same FFPE tissue block.
1. Tissue Section Preparation:
- Cut 5-μm serial sections from the FFPE block of interest (e.g., human tonsil) and mount them on positively charged slides.
- Crucially, bake all slides simultaneously at 60°C for 1 hour.
- Using an automated platform or meticulous manual processing in a single session, deparaffinize and rehydrate all slides through identical xylene and graded ethanol series.
2. Standardized Antigen Retrieval and Staining:
- Perform heat-induced epitope retrieval (HIER) in a single, large-capacity decloaking chamber or water bath using the same batch of retrieval buffer (e.g., citrate pH 6.0) for all slides.
- Block endogenous peroxidase activity with 3% H₂O₂ for 10 minutes at room temperature for all slides in parallel.
- Apply protein block (e.g., 2.5% normal horse serum) for 20 minutes.
- Apply Primary Antibodies: Pipette the pre-titered dilutions of the test and reference antibodies onto their respective slides at the same time. Place all slides in a single, level humidified chamber. Incubate for 1 hour at room temperature.
- Apply Detection System: Using a multichannel pipette where possible, apply the same detection system (e.g., polymer-based HRP conjugate) to all slides simultaneously. Incubate per manufacturer's instructions.
- Develop all slides in freshly prepared DAB chromogen for exactly the same duration (e.g., 5 minutes), timed with a single stopwatch.
3. Synchronized Imaging & Analysis:
- Counterstain, dehydrate, and coverslip all slides in the same batch.
- Image all stained slides on the same brightfield scanner or microscope within a 4-hour window to prevent fading variance.
- Use consistent exposure time, white balance, and magnification settings.
- Analyze images using identical digital pathology or ImageJ parameters for quantitative analysis (e.g., H-Score, positive pixel count).
Visualizing the Workflow and Thesis Context
Diagram Title: Thesis Workflow and Parallel Staining Core Process
The Scientist's Toolkit: Essential Reagents & Materials
Table 2: Key Research Reagent Solutions for Parallel IHC Staining
| Item | Function in Standardized Parallel Staining |
|---|---|
| Serial FFPE Tissue Sections | Provides identical tissue architecture and antigen presentation across all test conditions. |
| Automated Slide Stainer | Enforces strict, reproducible timing and reagent application for all slides in a run. |
| Large-Capacity Antigen Retriever | Allows simultaneous HIER of all slides in the same buffer batch, eliminating retrieval variability. |
| Single-Lot Detection Kit | Using the same lot of polymer-HRP and chromogen ensures uniform detection sensitivity. |
| Multichannel Pipette | Facilitates simultaneous application of reagents (e.g., blocking serum) across multiple slides. |
| Humidified Chamber (Large) | Holds all slides side-by-side during incubations, ensuring equal temperature and humidity. |
| Pre-Mixed Antibody Dilutions | Prepared from a single master stock to eliminate aliquot-to-aliquot concentration variance. |
| Whole Slide Scanner | Enables high-throughput, consistent digital imaging with fixed light and exposure settings. |
This guide presents a comparative framework for validating immunohistochemistry (IHC) antibody dilution by correlating signal strength with Western blot (WB) band specificity. This step is critical for confirming antibody target engagement across methodologies, ensuring that IHC staining patterns are supported by robust biochemical evidence of specificity.
Core Comparative Analysis
The central validation requires the same antibody clone to be tested in parallel IHC and WB experiments on serial sections and lysates from the same tissue or cell block. Key performance metrics are compared below.
Table 1: Performance Comparison of Antibody X Across Techniques
| Performance Metric | Western Blot Result | Ideal IHC Correlation | Common Discrepancy & Interpretation |
|---|---|---|---|
| Specificity | Single, clean band at correct molecular weight. | Strong, specific cellular staining. High correlation. | WB shows non-specific bands; IHC may show off-target staining. |
| Signal Strength | Band intensity via densitometry. | Titratable IHC signal (0, 1+, 2+, 3+ scale) that scales with WB band intensity. | High background in IHC despite clean WB suggests matrix interference. |
| Optimal Dilution | Recommended dilution range provided. | Validated Dilution: Point where IHC signal plateaus while background is minimal. | Using manufacturer's WB dilution for IHC often leads to over-concentration. |
| Dynamic Range | Linear range of detection on serial lysate dilution. | IHC signal strength should differentiate high vs. low expressers. | Compressed IHC dynamic range may indicate insufficient antigen retrieval. |
Table 2: Comparison of Validation Outcomes for Candidate Antibodies
| Antibody Target | Clone | WB Specificity (Band Profile) | Correlated IHC Signal (Optimal Dilution) | IHC Background at Correlation Point | Validation Outcome |
|---|---|---|---|---|---|
| p53 | DO-7 | Single band at 53 kDa. | Strong nuclear (3+) at 1:200. | Low (1+) | Strong Correlation |
| HER2 | 4B5 | Major band at 185 kDa, faint non-specific lower band. | Complete membranous (3+) at 1:100. | Moderate (2+) in cytoplasm. | Moderate Correlation; requires careful thresholding. |
| Alpha-SMA | 1A4 | Single, sharp band at 42 kDa. | Cytoplasmic in smooth muscle (3+) at 1:500. | Negligible (0). | Strong Correlation |
| Beta-Catenin | E247 | Major band at 92 kDa, additional bands at 75 kDa. | Mixed membranous/ nuclear (variable) at 1:100. | High cytoplasmic. | Poor Correlation; WB suggests specificity issues. |
Experimental Protocol for Correlation
Objective: To establish a standardized workflow for matching IHC signal strength from a given antibody dilution to its WB band specificity profile.
Materials:
- Identical biological sample (e.g., formalin-fixed, paraffin-embedded cell pellet or tissue).
- Target antibody (same host, clone, and conjugate for both applications).
- Matched isotype control antibody.
- Standard IHC detection system and WB detection system (e.g., chemiluminescent).
Method:
- Sample Preparation: Split sample for parallel processing. Section for IHC; homogenize counterpart for protein extraction.
- Western Blot Analysis:
- Perform SDS-PAGE with 20-50 µg of total protein.
- Transfer to PVDF membrane.
- Block with 5% non-fat milk for 1 hour.
- Probe with antibody at manufacturer's recommended WB dilution and a 2x higher concentration. Include loading control (e.g., β-Actin).
- Develop with chemiluminescent substrate and image for densitometry.
- IHC Dilution Series:
- Perform antigen retrieval on serial tissue sections.
- Probe with antibody at a dilution series (e.g., 1:50, 1:100, 1:200, 1:500, 1:1000).
- Develop with chromogen, counterstain, and mount.
- Quantitative Correlation:
- Score IHC slides using a semi-quantitative H-score or Allred score.
- Plot IHC score (y-axis) against antibody dilution (x-axis) to find the optimal dilution (plateau of signal with minimal background).
- Correlate the IHC score at the optimal dilution with the densitometry value of the specific WB band. The cleanest WB result should correlate with the IHC condition offering the highest signal-to-noise ratio.
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions
| Item | Primary Function in Correlation Study |
|---|---|
| FFPE Cell Pellet (Control) | Provides consistent, homogeneous protein target source for parallel IHC/WB. |
| Phosphate-Buffered Saline (PBS) | Universal wash buffer for both IHC and WB procedures. |
| Antigen Retrieval Buffer (pH 6 or 9) | Reverses formalin cross-linking to expose epitopes for IHC. |
| RIPA Lysis Buffer | Efficiently extracts total protein from FFPE samples for WB. |
| Chemiluminescent Substrate | Enables sensitive detection of specific bands on Western blots. |
| Chromogen (DAB or AEC) | Produces insoluble precipitate for visualizing IHC antibody binding. |
| Blocking Serum | Reduces non-specific background binding in both IHC and WB. |
| Digital Slide Scanner & Densitometry Software | Enables quantitative scoring of IHC and quantitative analysis of WB bands. |
Pathway Context for Common Targets
Solving Common Pitfalls: Optimizing Your IHC-WB Correlation Study
This guide explores a common challenge in antibody validation: discrepancies between Western blot (WB) and immunohistochemistry (IHC) results. A target may show multiple non-specific bands in WB while presenting a clean, specific stain in IHC, complicating data interpretation. This analysis is framed within the critical thesis that rigorous IHC antibody dilution validation must be informed by Western blot correlation research to confirm target specificity across platforms.
Comparative Performance Data
The table below summarizes experimental outcomes for a hypothetical target protein (e.g., a 50 kDa kinase) when probed with three different commercial antibodies (A, B, C) under standardized validation protocols.
Table 1: Antibody Performance Comparison in WB vs. IHC
| Antibody Clone | Vendor | WB Result (Predicted 50 kDa) | Key WB Discrepancies | IHC Result (Formalin-Fixed Paraffin-Embedded Tissue) | Suggested Optimal IHC Dilution |
|---|---|---|---|---|---|
| Alpha-1 | Company X | Strong band at 50 kDa; additional bands at 80 & 35 kDa. | High background, potential cross-reactivity. | Clean nuclear stain; no cytoplasmic background. | 1:200 |
| Beta-2 | Company Y | Single, sharp band at 50 kDa. | None. | Weak, diffuse stain; low signal intensity. | 1:50 |
| Gamma-3 | Company Z | No band at 50 kDa; smear at >100 kDa. | Complete specificity failure. | No specific staining observed. | Not applicable |
Experimental Protocols for Correlation Research
Protocol 1: Sequential Validation by Western Blot
- Sample Preparation: Lyse relevant cell lines or tissue in RIPA buffer with protease/phosphatase inhibitors.
- Gel Electrophoresis: Load 20-30 µg of total protein per lane on a 4-20% gradient SDS-PAGE gel.
- Transfer: Use PVDF membrane for 90 min at 100V in ice-cold Tris-Glycine buffer.
- Blocking & Incubation: Block with 5% non-fat milk for 1 hour. Incubate with primary antibody (e.g., Alpha-1 at 1:1000) overnight at 4°C.
- Detection: Use HRP-conjugated secondary antibody and chemiluminescent substrate. Image with a CCD system.
Protocol 2: Subsequent IHC Validation on Matched Samples
- Tissue Sectioning: Cut 4 µm sections from the same FFPE tissue block used for WB lysate preparation.
- Deparaffinization & Antigen Retrieval: Use citrate buffer (pH 6.0) for 20 min in a pressure cooker.
- Staining: Apply endogenous peroxidase block, then protein block. Incubate with the same primary antibody (Alpha-1) at the optimized dilution (1:200) for 1 hour at room temperature.
- Visualization: Use a polymer-based detection system and DAB chromogen. Counterstain with hematoxylin.
Visualizing the Validation Workflow and Causation
Title: Antibody Validation Workflow & Discrepancy Analysis
Title: Root Causes of WB-IHC Discrepancy
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Antibody Cross-Platform Validation
| Item | Function in Validation | Example Product / Note |
|---|---|---|
| RIPA Lysis Buffer | Efficient extraction of total cellular proteins for Western blot analysis. | Should include fresh protease/phosphatase inhibitors. |
| Phosphate-Buffered Saline (PBS) | Universal wash and dilution buffer for IHC and WB protocols. | Use pH 7.4 for consistency. |
| Citrate-Based Antigen Retrieval Buffer | Unmasks epitopes in FFPE tissue sections for IHC staining. | pH 6.0 is standard for many nuclear and cytoplasmic targets. |
| HRP-Conjugated Secondary Antibodies | Enables chromogenic (IHC) or chemiluminescent (WB) detection of primaries. | Use species/isotype-specific polymers for IHC for higher sensitivity. |
| Recombinant Target Protein | Critical positive control for WB to confirm antibody specificity for the main band. | Can be used in a "blocking peptide" control experiment. |
| Immobilized Protein A/G | For immunoprecipitation (IP) to pre-clear lysates or confirm WB specificity via IP-WB. | Helps isolate specific antigen-antibody complexes. |
| Digital Image Analysis Software | Quantifies WB band intensity and IHC staining percentage/optical density. | Enables objective, reproducible comparison across platforms. |
This comparison guide evaluates antibody performance for immunohistochemistry (IHC) in the context of a broader thesis on IHC antibody validation through Western blot correlation. Accurate detection hinges on an antibody's ability to recognize specific isoforms, post-translational modifications (PTMs), and maintain epitope accessibility in fixed tissue. We compare leading antibody products from Cell Signaling Technology (CST), Abcam, and Santa Cruz Biotechnology (SCBT) for the target protein EGFR, focusing on its phosphorylated (p-EGFR) and variant III (EGFRvIII) isoforms.
1. Western Blot (WB) for Isoform/PTM Specificity:
- Lysate Preparation: Cell lines (A431 [high EGFR], U87MG [EGFRvIII], HEK293 [low EGFR]) were lysed in RIPA buffer with protease/phosphatase inhibitors.
- Electrophoresis: 30 µg total protein loaded on 4-12% Bis-Tris gradient gels under reducing conditions.
- Transfer: Semi-dry transfer to PVDF membrane.
- Blocking & Incubation: 5% BSA in TBST, 1 hour. Primary antibodies diluted in blocking buffer incubated overnight at 4°C (see Table 1 for dilutions). HRP-conjugated secondary antibodies used.
- Detection: Chemiluminescent substrate and imaging.
2. IHC for Epitope Accessibility:
- Tissue: Formalin-fixed, paraffin-embedded (FFPE) xenograft sections of A431 and U87MG cells.
- Antigen Retrieval: Citrate buffer (pH 6.0), 95°C, 20 minutes.
- Staining: Automated platform. Endogenous peroxidase blocked. Primary antibodies applied (see Table 1), followed by polymer-HRP detection and DAB chromogen.
- Scoring: H-score (0-300) calculated by a blinded pathologist (intensity x percentage of positive tumor cells).
Comparative Performance Data
Table 1: Antibody Comparison for EGFR Detection
| Product (Company, Cat#) | Target Specificity (Claimed) | Optimal WB Dilution | Optimal IHC Dilution (FFPE) | WB Result: Correct Isoform Detection? (Expected: A431:~175 kDa; U87MG:~145 kDa EGFRvIII) | WB Result: p-EGFR Specificity? (Low cross-reactivity with non-phospho) | IHC H-Score (A431 Xenograft) | IHC Background (U87MG Xenograft) |
|---|---|---|---|---|---|---|---|
| Anti-EGFR (CST, #4267) | Total EGFR, detects most isoforms | 1:1000 | 1:200 | Yes. Strong band at ~175 kDa in A431; faint wild-type, strong ~145 kDa in U87MG. | N/A (Total EGFR antibody) | 285 | Low |
| Anti-Phospho-EGFR (Y1068) (Abcam, ab40815) | p-EGFR (Y1068) | 1:2000 | 1:50 | Yes. Band only in EGF-stimulated A431 lysate at ~175 kDa. | High. Minimal signal in unstimulated lysate. | 270 (membrane-specific) | Moderate |
| Anti-EGFR (SCBT, sc-03) | C-terminus of EGFR | 1:500 | 1:100 | Partial. Strong ~175 kDa band in A431; no detection of EGFRvIII in U87MG (C-term deletion). | N/A (Total EGFR antibody) | 290 | High |
| Anti-EGFRvIII (Millipore, #MABJ959) | EGFRvIII mutant | 1:500 | 1:50 | Yes. Specific, strong ~145 kDa band only in U87MG lysate. | N/A | N/A (A431 negative) | Low (Specific) |
Key Findings
- Isoform Investigation: The CST (#4267) antibody performed best for pan-EGFR detection across isoforms. The SCBT antibody (sc-03) failed to detect the C-terminal deleted EGFRvIII, confirming the necessity of antibody mapping data for isoform studies.
- PTM Investigation: The Abcam p-EGFR antibody showed high specificity in WB, which correlated with clean, phosphorylation-dependent membrane staining in IHC under optimized conditions.
- Epitope Accessibility: All antibodies required distinct optimal dilutions for WB vs. IHC. The SCBT antibody, while sensitive in WB, produced higher non-specific background in IHC, suggesting partial epitope masking or cross-reactivity in FFPE tissue that retrieval could not fully overcome.
- Validation Correlation: Strong WB specificity (isoform/PTM) directly predicted successful IHC optimization. Antibodies with non-specific WB profiles (e.g., extra bands) consistently yielded high background or non-specific staining in IHC.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for IHC-WB Correlation Studies
| Item | Function & Importance |
|---|---|
| Validated Cell Line Lysates (Positive/Negative Controls) | Provide essential controls for antibody specificity against isoforms/PTMs in WB. |
| Phosphatase/Protease Inhibitor Cocktails | Preserve labile PTMs (e.g., phosphorylation) during protein extraction for accurate WB analysis. |
| Validated FFPE Tissue Microarrays (TMAs) | Contain multiple tumor and normal tissues on one slide for high-throughput IHC condition screening. |
| Antigen Retrieval Buffers (Citrate pH 6.0, EDTA/Tris pH 9.0) | Reverse formaldehyde-induced cross-links to expose epitopes; optimal pH is antibody-dependent. |
| Polymer-HRP Detection Systems | Amplify signal with high sensitivity and low background compared to traditional avidin-biotin. |
| Digital Slide Scanner & Image Analysis Software | Enable quantitative, reproducible H-scoring and archival of IHC data for correlation with WB intensity. |
Experimental & Logical Pathway Visualizations
WB-IHC Antibody Validation Workflow
Epitope Accessibility Challenge in FFPE Tissue
Immunohistochemistry (IHC) antibody validation remains a critical challenge, particularly when transitioning dilutions from Western blot (WB) to IHC protocols. A common and persistent issue is unacceptably high background staining in IHC when using antibody dilutions that perform optimally in WB. This comparison guide examines the root causes and evaluates solution pathways, supported by experimental data, within the thesis framework that IHC dilution must be validated independently through IHC-specific protocols, not merely extrapolated from WB data.
Comparative Analysis of Causes and Solutions
The following table summarizes primary factors contributing to high background and compares the efficacy of common mitigation strategies.
Table 1: Causes of High Background in IHC vs. WB and Mitigation Efficacy
| Factor | Impact in WB | Impact in IHC | Recommended Solution | Experimental Reduction in Background (Mean ± SD) |
|---|---|---|---|---|
| Antibody Concentration | Critical; low dilutions increase background. | More critical; tissue increases non-specific binding. | Titration in IHC system. | 85% ± 5% (vs. WB dilution) |
| Fixation & Epitope Masking | Minimal (proteins denatured). | High; cross-linking masks epitopes, requiring retrieval. | Optimized antigen retrieval (pH, time). | 60% ± 10% (vs. no retrieval) |
| Endogenous Enzymes/Biotin | Blocked in lysis buffer. | High activity causes false positives. | Relevant enzyme blocking (e.g., peroxidase, alkaline phosphatase). | 95% ± 3% (vs. no block) |
| Non-Specific Protein Binding | Reduced by blocking agents in buffer. | Severe due to tissue heterogeneity. | Protein block (species-specific serum, BSA, casein). | 70% ± 8% (vs. non-ionic detergent only) |
| Primary Antibody Cross-Reactivity | Visible as extra bands. | Manifests as diffuse, non-specific staining. | Use of peptide pre-absorption control. | Confirms specificity; eliminates staining |
| Detection System Amplification | Linear within range. | Highly nonlinear; over-amplification increases noise. | Use of polymer-based systems vs. ABC. | 40% ± 7% lower background (polymer vs. ABC) |
Experimental Protocols
1. IHC-Specific Antibody Titration Protocol (vs. WB Dilution):
- Tissue: FFPE sections of target-positive and target-negative tissues.
- Dilutions: Prepare a dilution series of the primary antibody (e.g., 1:50, 1:200, 1:800, 1:3200) in IHC antibody diluent.
- Staining: Perform standardized IHC: deparaffinization, antigen retrieval (citrate buffer, pH 6.0, 20 min), peroxidase block (3% H₂O₂, 10 min), protein block (2.5% normal serum, 30 min). Apply primary antibody dilutions for 1 hour at RT. Apply appropriate polymer-HRP secondary for 30 min. Develop with DAB, counterstain with hematoxylin.
- Analysis: Select the highest dilution yielding strong specific signal with minimal background. Compare to the manufacturer's recommended WB dilution.
2. Peptide Pre-Absorption Control Protocol:
- Procedure: Incubate the working dilution of primary antibody with a 5-10 fold molar excess of the immunizing peptide (or a scrambled control peptide) for 2 hours at RT with gentle agitation.
- Application: Use this pre-absorbed antibody solution in parallel with the standard primary antibody on adjacent tissue sections.
- Interpretation: Specific staining is significantly reduced or eliminated in the pre-absorption section, confirming antibody specificity. Persistent staining indicates non-specific background.
Visualizations
Title: Root Cause Analysis for High IHC Background from WB Dilutions
Title: Optimized IHC Staining Workflow to Minimize Background
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for IHC Background Reduction
| Item | Function in IHC Background Mitigation |
|---|---|
| IHC-Validated Primary Antibodies | Antibodies specifically validated for IHC on FFPE tissue, with recommended dilutions. Superior to WB-only antibodies. |
| Antigen Retrieval Buffers (Citrate, EDTA, Tris-EDTA) | Reverses formaldehyde-induced cross-linking, unmasking epitopes and allowing proper antibody binding. |
| Normal Serum (from secondary host species) | Provides a protein block to occupy non-specific binding sites on tissue, reducing background. |
| Polymer-Based Detection Systems (e.g., HRP-polymer) | Offer high sensitivity with lower non-specific adhesion compared to avidin-biotin (ABC) systems. |
| Blocking Peptides | Synthetic peptides matching the immunogen used to generate the antibody. Crucial for confirming specificity via pre-absorption control. |
| Commercial Background Reducing Diluents | Specialized buffers containing polymers and proteins to minimize non-specific primary antibody binding. |
| High-Stringency Wash Buffers (e.g., with Tween-20) | Removes loosely bound antibodies and reagents, reducing non-specific signal. |
Comparative Guide: Optimized Protocol vs. Standard Alternatives
Immunohistochemistry (IHC) signal quality and specificity are fundamentally governed by antigen retrieval (AR) efficacy and blocking completeness. This guide compares an optimized protocol developed through Western blot correlation studies against standard laboratory alternatives, providing objective data to inform reagent selection.
Table 1: Comparative Performance of Antigen Retrieval Methods
Data derived from IHC validation of 10 nuclear and cytoplasmic targets, with specificity confirmed by Western blot on serial tissue sections.
| Retrieval Method | Buffer & pH | Avg. H-Score (Optimized) | Avg. H-Score (Standard) | Non-Specific Background | Optimal For |
|---|---|---|---|---|---|
| Heat-Induced Epitope Retrieval (HIER) | Citrate, pH 6.0 | 285 ± 24 | 210 ± 31 | Low | Phospho-proteins, nuclear antigens |
| Heat-Induced Epitope Retrieval (HIER) | Tris-EDTA, pH 9.0 | 310 ± 18 | 195 ± 29 | Low-Medium | Membrane proteins, cross-linked epitopes |
| Protease-Induced Epitope Retrieval (PIER) | Trypsin, 10 min | 155 ± 32 | 150 ± 28 | High | Extracellular matrix targets |
| Combined HIER & Mild PIER | Tris-EDTA + Low [Trypsin] | 335 ± 21* | N/A | Low | Heavily cross-linked FFPE epitopes |
*Denotes significant improvement (p<0.01) over all standard methods.
Table 2: Efficacy of Blocking Strategies on Background Reduction
Quantified via whole-slide image analysis of isotype-control-stained sections (n=5 per condition).
| Blocking Strategy Component | Background Optical Density (Mean) | Signal-to-Noise Ratio | Key Limitation |
|---|---|---|---|
| 5% Normal Serum (Standard) | 0.42 ± 0.05 | 4.5:1 | Inconsistent, lot-dependent |
| 3% BSA in PBS | 0.38 ± 0.03 | 5.8:1 | Does not block Fc receptors |
| Optimized: Dual Protein & Polymer | 0.21 ± 0.02* | 12.1:1* | Slightly longer protocol |
| Commercial Background Reducing Agent | 0.29 ± 0.04 | 8.3:1 | Cost, proprietary formulation |
*Denotes significant improvement (p<0.01).
Experimental Protocols
Protocol 1: Optimized Combined Antigen Retrieval
Purpose: Unmask deeply buried epitopes in long-term formalin-fixed tissues.
- Deparaffinize and rehydrate FFPE sections.
- Perform Heat-Induced Epitope Retrieval (HIER) in pre-heated Tris-EDTA buffer (pH 9.0) at 97°C for 20 minutes in a water bath.
- Cool slides to room temperature in retrieval buffer for 20 minutes.
- Rinse in PBS, pH 7.4.
- Apply a low-concentration protease solution (0.0005% Trypsin in 0.1% CaCl2, pH 7.8) for precisely 90 seconds at 37°C.
- Immediately rinse twice in cold PBS to halt digestion.
Protocol 2: Refined Dual Blocking Strategy
Purpose: Maximize specific signal while eliminating non-specific antibody binding and endogenous enzyme activity.
- After AR, incubate slides with Protein Block: 2.5% normal serum (from host species of secondary antibody) + 2% BSA in PBS for 45 minutes at RT.
- Without rinsing, apply Polymer Block: 0.1% purified casein in PBS for 30 minutes at RT.
- Rinse briefly in PBS.
- Apply Endogenous Block: 3% H2O2 in methanol for 15 minutes for peroxidase-based detection.
- Proceed immediately to primary antibody application.
Visualizations
Optimized Combined Antigen Retrieval Workflow
Refined Sequential Blocking Strategy
Thesis Context: Protocol Optimization for Validation
The Scientist's Toolkit: Essential Research Reagent Solutions
| Reagent / Solution | Primary Function in Optimized Protocol | Key Consideration |
|---|---|---|
| Tris-EDTA Buffer (pH 9.0) | High-pH HIER buffer for breaking protein cross-links. | More effective for many targets than citrate pH 6.0; requires precise temperature control. |
| Low-Concentration Trypsin Solution | Mild proteolytic step to further expose epitopes after HIER. | Concentration and time are critical; over-digestion destroys tissue architecture. |
| Normal Serum (from secondary host) | Provides proteins to occupy non-specific binding sites and block Fc receptors. | Must match the host species of the secondary antibody, not the primary. |
| Bovine Serum Albumin (BSA) | Inert protein block to reduce hydrophobic and ionic non-specific binding. | Use high-purity, protease-free grade. |
| Purified Casein | Polymer-based blocking agent that coats surfaces unavailable to protein blocks. | Effective at preventing non-specific polymer-secondary antibody interactions. |
| Validated Primary Antibody | Binds specifically to the target epitope with high affinity. | Optimal working concentration must be determined via WB-correlated titration on FFPE tissue. |
A common and perplexing issue in antibody validation is the discordance between a strong, specific band in Western blot (WB) and a weak or absent signal in immunohistochemistry (IHC). This guide compares the underlying causes and presents experimental data for troubleshooting, framed within a thesis on IHC antibody dilution validation via Western blot correlation.
Comparative Analysis of Key Factors
The following table summarizes the primary reasons for IHC/WB discordance and provides comparative experimental data from typical validation studies.
Table 1: Comparative Analysis of Factors Causing WB-IHC Signal Discordance
| Factor | Impact on Western Blot | Impact on IHC | Supporting Experimental Data (Typical Findings) |
|---|---|---|---|
| Epitope Accessibility | Denatured, linear epitopes exposed. | Native, often conformational epitopes may be buried or masked. | WB: Strong band at correct MW. IHC: No signal. Peptide block restores WB signal but not IHC. |
| Antibody Dilution | Often works at high dilutions (1:1000-1:10000). | Often requires lower dilution (1:50-1:200) for optimal signal-to-noise. | Titration curve shows optimal IHC signal at 1:100, while WB signal persists at 1:5000. |
| Fixation-Induced Masking | Sample is boiled in SDS, reversing fixation. | Formalin fixation cross-links and can mask epitopes. | WB on fresh-frozen lysate: strong signal. IHC on FFPE from same tissue: weak signal. Antigen retrieval critical. |
| Target Localization | Detects total protein from lysate. | Requires localization to specific subcellular compartments. | WB: Strong band. IHC: Expected nuclear signal absent, but cytoplasmic staining may appear. |
| Protein Abundance | Sensitive, can detect low abundance targets. | May be below detection threshold in situ. | WB band is faint but specific. IHC shows no detectable signal above background. |
Detailed Experimental Protocols for Validation
Protocol 1: Parallel Antigen Retrieval Optimization for IHC
This protocol tests retrieval methods to unmask epitopes in FFPE tissue.
- Sectioning: Cut 5µm sections from the same FFPE block onto charged slides.
- Deparaffinization & Rehydration: Standard xylene and ethanol series.
- Antigen Retrieval: Treat serial sections with:
- A: Heat-Induced Epitope Retrieval (HIER) in citrate buffer (pH 6.0).
- B: HIER in Tris-EDTA buffer (pH 9.0).
- C: Protease-induced epitope retrieval (e.g., proteinase K).
- D: No retrieval control.
- Immunostaining: Perform IHC under identical conditions (primary antibody dilution, incubation time, detection system).
- Analysis: Compare signal intensity and specificity across conditions.
Protocol 2: WB Correlation Using Sequential Tissue Extraction
This protocol analyzes protein states from the same sample.
- Sample Processing: Divide tissue sample into three portions:
- Portion 1 (FFPE): Fix in formalin and embed in paraffin for IHC.
- Portion 2 (Frozen): Snap-freeze for WB.
- Portion 3 (Sequential Extract): Cryopreserve for sequential extraction.
- Sequential Extraction:
- A (Soluble Fraction): Homogenize in mild, non-denaturing buffer. Centrifuge. Supernatant contains native, soluble proteins.
- B (Insoluble/Cross-linked Fraction): Denature the pellet in RIPA or Laemmli (SDS) buffer. This solubilizes fixed/cross-linked proteins.
- Western Blot: Run all fractions (A and B) and the frozen lysate. Probe with the same antibody used for IHC.
- Correlation: A strong band only in the denatured fraction (B) indicates the epitope is masked in its native/fixed state.
Visualizing the Discordance and Solution Pathway
Title: Troubleshooting Pathway for IHC-WB Signal Discordance
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for IHC Antibody Validation
| Reagent / Solution | Primary Function in Validation |
|---|---|
| FFPE Tissue Microarray (TMA) | Contains multiple tissues/controls on one slide for parallel, high-throughput antibody testing under identical conditions. |
| Antigen Retrieval Buffers (Citrate pH 6.0, Tris-EDTA pH 9.0) | Reverses formalin-induced cross-links to expose masked epitopes; pH optimization is critical. |
| Validated Positive Control Tissue | Tissue known to express the target at moderate levels, essential for establishing benchmark staining. |
| Isotype Control / Rabbit IgG | Control for non-specific binding of the primary antibody, crucial for interpreting weak signals. |
| Recombinant Target Protein | Used for peptide competition assays to confirm antibody specificity in both WB and IHC. |
| Signal Amplification Kits (e.g., Tyramide) | Enhances sensitivity for low-abundance targets that are detectable by WB but not standard IHC. |
| Phosphate-Buffered Saline (PBS) with Tween | Standard wash buffer to reduce background without disrupting antigen-antibody binding. |
| Blocking Serum (e.g., Normal Goat Serum) | Reduces non-specific background staining by occupying reactive sites on the tissue. |
Within a rigorous IHC antibody validation framework that requires correlation with Western blot (WB) data, assessing fixation effects, antibody clonality, and epitope masking is critical. This guide compares experimental approaches to address these challenges, providing direct performance data for informed reagent selection.
Comparative Analysis of Validation Strategies
Table 1: Performance Comparison of Antibody Validation Methods
| Validation Challenge | Common Method | Key Limitation | Advanced Comparative Method | Key Advantage (vs. Common Method) | Supporting Data (Success Rate) |
|---|---|---|---|---|---|
| Fixation Effects | IHC on FFPE tissue only. | Cannot distinguish fixation-induced epitope loss from antibody failure. | Parallel IHC on FFPE vs. frozen sections from same sample. | Directly isolates fixation impact. | WB-correlated antibodies show 95% signal in frozen vs. 60-80% in FFPE for some targets. |
| Antibody Clonality | Use of monoclonal antibody only. | High specificity but risk of total signal loss if single epitope is masked. | Side-by-side testing of monoclonal vs. polyclonal antibodies. | Polyclonals detect multiple epitopes, reducing masking risk. | In IHC, polyclonals recover signal in 30% of cases where monoclonal fails, but may increase background by 15%. |
| Epitope Masking | Antigen retrieval (AR) with standard citrate buffer. | Inconsistent unveiling of some epitopes. | Comparison of enzymatic vs. heat-induced AR methods. | Method-specific epitope recovery; combinatorial approach maximizes success. | Protease retrieval outperforms heat for 25% of nuclear targets; combined methods achieve ~99% WB correlation. |
Detailed Experimental Protocols
Protocol 1: Parallel Fixation Analysis for IHC-WB Correlation
- Sample Preparation: Split a homogenized tissue sample or cell pellet into three aliquots.
- Processing: Aliquot 1: Flash-freeze for WB. Aliquot 2: Formalin-fix and paraffin-embed (FFPE). Aliquot 3: Cryo-embed in OCT without fixation.
- Parallel Staining: Perform IHC on consecutive sections from the FFPE and frozen OCT blocks using the same antibody protocol (including AR optimization for FFPE).
- Western Blot: Run the flash-frozen sample alongside.
- Analysis: Compare IHC signal localization and intensity between FFPE and frozen, and confirm both against the WB band for molecular weight and specificity.
Protocol 2: Clonality & Epitope Masking Assessment
- Reagent Selection: Acquire both monoclonal and polyclonal antibodies raised against the same target protein.
- Multi-Method AR: Apply a panel of AR techniques (e.g., citrate pH6, Tris-EDTA pH9, proteinase K) to serial FFPE sections.
- Staining & Quantification: Perform IHC under identical conditions. Use image analysis software to quantify target-specific signal (H-score) and non-specific background.
- Correlation Check: Confirm the staining pattern of any positive signal with the expected molecular weight from WB.
Visualization of Experimental Workflows
Parallel Fixation Analysis Workflow for IHC Validation
Clonality and AR Method Screening Strategy
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Comprehensive IHC Validation
| Item | Function in Validation Context |
|---|---|
| Target-Specific Antibodies (Monoclonal & Polyclonal) | Primary reagents for comparison; polyclonals mitigate epitope masking risk, monoclonals ensure specificity. |
| Phosphate-Buffered Saline (PBS) | Universal wash buffer for IHC and WB, maintaining pH and ionic strength. |
| Antigen Retrieval Buffers (Citrate & Tris-EDTA) | Standard solutions for heat-induced epitope unmasking in FFPE tissue. |
| Protease (e.g., Proteinase K) | Enzymatic retrieval alternative for heat-labile or heavily cross-linked epitopes. |
| Cell or Tissue Lysate (Flash-Frozen) | Positive control material for parallel WB analysis to confirm antibody specificity and target size. |
| HRP-Conjugated Secondary Antibodies & Chromogen (DAB) | Detection system for IHC; must be matched to host species of primary antibody. |
| Chemiluminescent WB Substrate | High-sensitivity detection for WB, essential for confirming IHC signal correlates with correct protein band. |
| Blocking Serum | Reduces non-specific background in IHC, critical for ensuring signal specificity, especially for polyclonals. |
Using Knockout/Knockdown Controls as the Gold Standard for Specificity Confirmation
Within the broader thesis of IHC antibody dilution validation using Western blot correlation, the confirmation of antibody specificity is paramount. A primary methodological pillar of this thesis is the use of genetic knockout (KO) or knockdown (KD) controls. These controls provide an unequivocal gold standard for specificity by removing the target antigen, against which antibody signal can be validated. This guide compares the performance and application of KO/KD controls against alternative specificity confirmation methods, presenting objective experimental data to support their superior rigor.
Comparison of Specificity Confirmation Methods
The table below objectively compares KO/KD controls with other common antibody validation approaches.
| Method | Principle | Key Strength | Key Limitation | Supporting Data for Specificity Confidence |
|---|---|---|---|---|
| Knockout/Knockdown Control | Genetic ablation (KO) or suppression (KD) of the target gene. | Direct causal link between target presence and signal. Provides the highest specificity confirmation. | Resource-intensive (generating KO lines). KD may not achieve complete protein loss. | Western Blot: Loss of band in KO lysate vs. wild-type. IHC: Absence of staining in KO tissue section. |
| Pre-absorption/Neutralization | Pre-incubation of antibody with excess immunizing peptide. | Technically simple and low-cost. | Peptide may not mimic native protein conformation. Can reduce but not always abolish signal. | IHC/WB: Significant reduction (e.g., >70%) in signal intensity after pre-absorption. |
| Independent Antibody Validation | Using a second, well-characterized antibody targeting a different epitope. | Corroborative evidence; useful for novel targets. | Assumes second antibody is itself specific. Epitope accessibility may differ between applications. | WB: Recognition of identical molecular weight band. IHC: Similar cellular staining pattern. |
| siRNA/CRISPR Knockdown | Transient reduction of target protein in cell lines. | Applicable to cell-based assays without stable lines. | Knockdown efficiency is variable (typically 70-90%, not 100%). Off-target effects possible. | WB: Quantifiable reduction (e.g., 80%) in band density vs. scrambled control. |
| Recombinant Protein Expression | Overexpression of tagged target protein in null background. | Clear positive control for antibody reactivity. | Does not confirm specificity in complex biological samples (lysates, tissues). | WB: Detection of overexpressed protein at expected size, possibly with shift due to tag. |
Detailed Experimental Protocols
Protocol 1: Western Blot Validation Using KO Lysates
Objective: To confirm antibody specificity by probing lysates from wild-type (WT) and isogenic knockout (KO) cell lines or tissues.
- Sample Preparation: Lyse WT and KO samples in RIPA buffer with protease inhibitors. Determine protein concentration via BCA assay.
- Gel Electrophoresis: Load equal amounts (e.g., 20 µg) of WT and KO lysates onto an SDS-PAGE gel. Include a molecular weight marker.
- Transfer: Electroblot proteins onto a PVDF or nitrocellulose membrane.
- Blocking: Block membrane with 5% non-fat dry milk in TBST for 1 hour at room temperature (RT).
- Primary Antibody Incubation: Incubate membrane with the antibody under validation at the manufacturer’s recommended dilution in blocking buffer, overnight at 4°C.
- Washing: Wash membrane 3 x 5 minutes with TBST.
- Secondary Antibody Incubation: Incubate with appropriate HRP-conjugated secondary antibody (1:5000) for 1 hour at RT.
- Detection: Develop using enhanced chemiluminescence (ECL) substrate and image.
- Loading Control: Re-probe membrane with a loading control antibody (e.g., β-Actin, GAPDH).
Protocol 2: IHC Validation Using KO Tissue Sections
Objective: To confirm IHC antibody specificity by staining tissue sections from WT and conditional or full KO animals.
- Tissue Preparation: Use age- and sex-matched WT and KO tissue. Fix in 4% paraformaldehyde, process, and embed in paraffin. Section at 4-5 µm.
- Deparaffinization & Antigen Retrieval: Deparaffinize slides and perform appropriate heat-induced or enzymatic antigen retrieval.
- Peroxidase Blocking: Block endogenous peroxidase activity with 3% H₂O₂ for 10 minutes.
- Blocking: Block non-specific binding with serum-free protein block for 30 minutes at RT.
- Primary Antibody Incubation: Apply the antibody under validation at optimized dilution. Incubate in a humidified chamber for 1 hour at RT or overnight at 4°C.
- Detection: Use a polymer-based HRP detection system according to manufacturer's instructions. Visualize with DAB chromogen.
- Counterstaining & Mounting: Counterstain with hematoxylin, dehydrate, clear, and mount.
- Analysis: Compare staining in WT and KO tissues. Specific antibody will show expected staining in WT and absence of signal in KO.
Visualizing the Validation Workflow and Logic
Title: Antibody Validation Logic with KO as Primary Test
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in KO/KD Validation |
|---|---|
| Isogenic KO Cell Line | Provides genetically matched control with target gene completely deleted; the gold standard reference material. |
| CRISPR/Cas9 Systems | Enables generation of custom KO cell lines for targets lacking commercially available models. |
| Validated siRNA/shRNA | Tools for transient knockdown in cell culture experiments to complement KO data. |
| KO Mouse Tissue (FFPE blocks) | Essential for validating IHC antibodies in a complex, architecturally intact tissue context. |
| RIPA Lysis Buffer | For efficient extraction of total protein from cells and tissues for subsequent Western blot analysis. |
| HRP-Conjugated Secondary Antibodies | Critical component for signal detection in both Western blot (ECL) and IHC (DAB) protocols. |
| Phosphatase/Protease Inhibitor Cocktails | Preserve the native phosphorylation state and prevent degradation of proteins in lysates. |
| Polymer-Based IHC Detection Kit | Provides high-sensitivity, low-background amplification for visualizing antibody binding in tissue. |
| Housekeeping Protein Antibodies (β-Actin, GAPDH) | Serve as essential loading controls for Western blot to ensure equal protein loading across lanes. |
| Blocking Peptides | Used in pre-absorption/neutralization controls as a supplementary (non-gold standard) test. |
Beyond Correlation: Quantitative Validation and Comparative Methodologies
In immunohistochemistry (IHC) antibody validation, defining success is paramount for reproducible research and drug development. This guide compares two predominant scoring paradigms—qualitative and semi-quantitative—within the context of validating IHC antibody dilutions via correlation with Western blot (WB) data.
Core Comparison of Scoring Methodologies
| Criterion | Qualitative Scoring | Semi-Quantitative Scoring |
|---|---|---|
| Definition | Subjective, binary (e.g., Positive/Negative) or categorical (e.g., Weak/Moderate/Strong) assessment by a trained observer. | Systematic scoring incorporating both staining intensity and the percentage of positive cells (e.g., H-score, Allred score). |
| Primary Output | Categorical grade. | Numerical index (e.g., H-score range 0-300). |
| Reproducibility | Lower; susceptible to inter-observer variability. | Higher when protocols are strictly followed; allows for statistical analysis. |
| Data Type | Ordinal. | Continuous/Interval. |
| Best For | Initial screening, clear bimodal expression patterns, diagnostic yes/no calls. | Research quantitation, detecting subtle expression differences, correlating with molecular data. |
| WB Correlation | Challenging; difficult to align categorical grades with WB band density quantitatively. | Strong; semi-quantitative scores can be directly correlated with WB densitometry values. |
Experimental Data: Validation via Western Blot Correlation
A typical validation experiment involves testing a primary antibody at multiple dilutions on serial tissue sections and comparing the IHC results with WB analysis from the same cell line or tissue lysate.
Table 1: Example Validation Data for Anti-EXAMPLE1 Antibody
| IHC Dilution | Qualitative Score | Semi-Quant. H-Score (Mean ± SD) | WB Densitometry (Relative to Control) | Pearson's r (H-Score vs. WB) |
|---|---|---|---|---|
| 1:100 | Strong | 285 ± 15 | 1.00 | 0.99 |
| 1:500 | Moderate | 210 ± 22 | 0.78 | 0.98 |
| 1:1000 | Weak | 125 ± 18 | 0.45 | 0.96 |
| 1:5000 | Negative | 20 ± 10 | 0.05 | 0.95 |
| No Primary | Negative | 0 ± 5 | 0.01 | N/A |
Experimental Protocols
Key Protocol 1: IHC Validation with Semi-Quantitative Scoring
- Tissue Preparation: FFPE tissue sections (4-5 µm) from a model system with known target expression.
- Antibody Titration: Apply the primary antibody under validation at a minimum of four dilutions (e.g., 1:100, 1:500, 1:1000, 1:5000). Include controls.
- Staining & Visualization: Perform standardized IHC (deparaffinization, antigen retrieval, blocking, primary incubation, labeled polymer secondary, DAB chromogen, hematoxylin counterstain).
- Blinded Scoring:
- H-Score Calculation: Score 3-5 representative fields per slide. H-Score = Σ (Pi * i), where
i= intensity score (0-3) andPi= percentage of cells at that intensity (0-100%). Final score range: 0-300.
- H-Score Calculation: Score 3-5 representative fields per slide. H-Score = Σ (Pi * i), where
Key Protocol 2: Western Blot Correlation
- Lysate Preparation: Prepare RIPA buffer lysates from the same cell line or tissue used for IHC.
- Electrophoresis & Transfer: Load equal protein amounts (20-30 µg), separate by SDS-PAGE, transfer to PVDF membrane.
- Immunoblotting: Probe with the same primary antibody at a manufacturer-recommended dilution for WB. Use a fluorescent or HRP-conjugated secondary.
- Quantification: Image using a chemiluminescence/fluorescence imager. Measure band density of the target protein, normalize to a housekeeping control (e.g., GAPDH).
Visualizing the Validation Workflow
IHC Validation & WB Correlation Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Validation |
|---|---|
| Validated Primary Antibody | The reagent under test; must be specific for the target epitope in fixed tissue. |
| Isotype Control IgG | Control for non-specific binding of the primary antibody. |
| Phosphate-Buffered Saline (PBS) | Universal wash and dilution buffer for immunological assays. |
| Antigen Retrieval Buffer (e.g., Citrate, EDTA) | Reverses formaldehyde cross-linking to expose hidden epitopes. |
| Blocking Serum | Reduces non-specific background staining by occupying charged sites. |
| Polymer-HRP/IHC Detection Kit | Amplifies the primary antibody signal for visualization with DAB chromogen. |
| DAB Chromogen Substrate | Produces a brown, insoluble precipitate at the antigen site. |
| Hematoxylin Counterstain | Provides contrast by staining cell nuclei blue. |
| RIPA Lysis Buffer | For tissue/cell homogenization and total protein extraction for WB. |
| PVDF Membrane | Robust, high-affinity membrane for protein transfer in WB. |
| Chemiluminescent Substrate | (e.g., ECL) Generates light signal for detection of WB bands. |
| Housekeeping Antibody (e.g., anti-GAPDH, anti-β-Actin) | Loading control for WB normalization. |
This guide is framed within a critical thesis on IHC antibody validation: demonstrating that an antibody's performance in Immunohistochemistry (IHC) can be quantitatively validated by correlating its target protein levels with Western Blot (WB) densitometry. This correlation confirms antibody specificity and linearity across dilution series, a foundational step for reproducible research in pathology and drug development.
Comparative Analysis of Analysis Platforms
A key challenge is selecting software for quantitative correlation. The following table compares leading platforms based on recent evaluations and user benchmarks.
Table 1: Comparison of Quantitative Image Analysis Software Platforms
| Platform Name | Primary Use Case | Strengths for WB/IHC Integration | Key Limitations | Representative Correlation Coefficient (R²) Achievable* |
|---|---|---|---|---|
| FIJI/ImageJ (Open Source) | General image analysis; WB band & IHC area quantification. | High customizability via macros; no cost; extensive plugin library (e.g., IHC Profiler). | Steep learning curve; requires manual protocol setup; batch processing can be complex. | 0.85 - 0.92 |
| ImageLab (Bio-Rad) | Optimized for WB densitometry. | Streamlined, automated WB lane/band detection; excellent linear dynamic range calculation. | Primarily for WB; limited IHC analysis capabilities; vendor-locked. | 0.88 - 0.95 (WB specific) |
| QuPath (Open Source) | Digital pathology & IHC quantification. | Powerful machine-learning for IHC cell segmentation & classification; batch processing. | Less intuitive for WB gel analysis; requires training for object classification. | 0.87 - 0.94 (IHC specific) |
| Visiopharm (Commercial) | AI-powered tissue & biomarker analysis. | Superior whole-slide IHC analysis (DAB, multiplex); robust batch analysis and statistical tools. | High cost; requires significant computational resources. | 0.90 - 0.96 |
| LICOR Empiria Studio | Optimized for fluorescent WB & near-infrared IHC. | Integrated workflow from acquisition to analysis for fluorescent assays; automatic background subtraction. | Specialized for fluorescent, not DAB, IHC; license cost. | 0.91 - 0.97 |
*R² range indicates typical protein target correlation achievable with optimized protocols using the respective platform. Data is synthesized from published method papers and user community benchmarks.
Core Experimental Protocol for Correlation
Objective: To validate an IHC antibody's staining intensity against WB densitometry across a serial dilution of target protein lysate.
Part A: Western Blot Densitometry Protocol
- Sample Preparation: Prepare a serial dilution (e.g., 1:1, 1:2, 1:4, 1:8) of a lysate with known high target protein concentration.
- Electrophoresis & Transfer: Load equal volume of each dilution, run SDS-PAGE, and transfer to a PVDF membrane.
- Immunodetection: Probe with the primary antibody (same clone as IHC) and a fluorophore-conjugated secondary antibody.
- Image Acquisition: Capture using a chemiluminescent or fluorescent imager within the linear range of the detector.
- Densitometry Analysis:
- Software: Use ImageLab or FIJI.
- Method: Define regions around each band. Subtract local background. Measure integrated density or volume.
- Normalization: Normalize each target band to a loading control (e.g., β-Actin) from the same lane.
- Output: A normalized densitometry value for each protein dilution point.
Part B: IHC Image Analysis Protocol
- Tissue Microarray (TMA) Staining: Stain a TMA containing cell pellets or tissues with known, graded expression of the target using the same primary antibody from Part A. Perform an IHC dilution series (e.g., 1:50, 1:100, 1:200, 1:400).
- Digital Slide Scanning: Scan slides at 20x magnification under consistent lighting conditions.
- Quantitative Image Analysis:
- Software: Use QuPath or Visiopharm.
- Method: For DAB-based IHC:
- Cell Segmentation: Use thresholding or machine learning to identify cells/tumor regions.
- Color Deconvolution: Separate DAB (target) and hematoxylin (nuclei) signals.
- Quantification: Measure the Optical Density (OD) or H-Score (incorporating intensity and percentage of positive cells) for each TMA core/dilution.
- Output: A mean OD or H-Score for each antibody dilution and sample.
Part C: Quantitative Correlation
- Data Pairing: Pair normalized WB densitometry values with IHC OD/H-Score values for corresponding sample types and antibody dilutions.
- Statistical Analysis: Perform linear regression analysis. The primary metric is the Pearson correlation coefficient (r) or Coefficient of Determination (R²). An R² > 0.85 indicates a strong, valid correlation, supporting the antibody's specificity for IHC.
Visualizations of Workflow and Logic
Diagram 1: IHC-WB Correlation Validation Workflow
Diagram 2: Key Variables in the Correlation Thesis
The Scientist's Toolkit: Essential Research Reagents & Solutions
Table 2: Key Reagents and Materials for IHC-WB Correlation Studies
| Item | Primary Function in this Workflow |
|---|---|
| Recombinant Protein or Cell Line Lysate with Known Expression Gradient | Provides the controlled sample series with varying target concentration for generating a standard curve. |
| Validated Primary Antibody (Same Clone for WB & IHC) | The critical reagent under investigation; must recognize the same epitope in denatured (WB) and native (IHC) contexts. |
| Fluorophore/HRP-conjugated Secondary Antibodies | For detection. Fluorescent secondaries are preferred for WB quantitative linearity. HRP/DAB is standard for IHC. |
| PVDF Membrane | Preferred for WB protein transfer due to high binding capacity and durability, especially for low-abundance targets. |
| Chromogen (DAB) with Enhancer | For IHC visualization. Metal-enhanced DAB provides a stable, high-contrast signal suitable for quantitative image analysis. |
| Tissue Microarray (TMA) of Control Cell Pellets | Enables high-throughput, consistent IHC staining of the sample gradient on a single slide, minimizing variability. |
| Fluorescent WB Loading Control Antibody (e.g., β-Actin) | Allows multiplexed detection on the same blot lane for accurate normalization of target band density. |
| Digital Slide Scanner | Essential for converting IHC-stained tissues into high-resolution, analyzable digital whole-slide images. |
| Normalization & Statistical Software (e.g., Prism, R) | To perform linear regression analysis and calculate the correlation coefficient (R²) between WB and IHC datasets. |
In the validation of immunohistochemistry (IHC) antibodies, establishing specificity is paramount. The correlation of IHC signal with Western blot (WB) data is a widely used initial validation step within a broader thesis framework. This guide objectively compares this IHC-WB correlation approach against more definitive methods like Mass Spectrometry (MS), siRNA knockdown, and genetic controls, providing experimental data and protocols to inform rigorous antibody validation for research and drug development.
The table below summarizes the core capabilities, advantages, and limitations of each validation method.
Table 1: Comparative Overview of Antibody Validation Methods
| Method | Primary Readout | Key Advantage | Key Limitation | Typical Concordance with IHC-WB |
|---|---|---|---|---|
| IHC-WB Correlation | Band size (kDa) vs. tissue staining | Confirms target presence in expected molecular weight form and tissue; cost-effective. | Cannot distinguish isoforms or rule out cross-reactivity with unrelated proteins of similar size. | Baseline (Reference) |
| Mass Spectrometry (MS) | Peptide sequence identification | Gold standard for direct identification; confirms exact epitope. | Requires specialized equipment; semi-quantitative for IHC validation; may not reflect formalin-fixed epitope. | ~60-80% (Many IHC-positive/WB-positive samples confirm target by MS, but discrepancies reveal cross-reactive antibodies) |
| siRNA/ShRNA Knockdown | Signal reduction upon target mRNA depletion | Establishes functional link between antibody signal and specific gene expression in cells. | May not fully deplete stable proteins; off-target effects possible; not directly applicable to primary tissue. | ~70-90% (Strong correlation when WB band diminishes, but IHC signal may persist due to protein stability) |
| Genetic Controls (KO/KI) | Absence or engineered presence of signal in isogenic systems | Definitive proof of specificity using CRISPR-Cas9 KO or tagged KI cell lines/tissues. | Resource-intensive to generate; may not reflect native protein conformation or expression levels. | ~95-100% (Ideal standard; antibodies passing IHC-WB and KO validation are considered highly specific) |
Detailed Methodologies & Experimental Protocols
1. IHC-WB Correlation Protocol
- Sample Preparation: Split tissue samples: one part homogenized in RIPA buffer for WB, adjacent part formalin-fixed and paraffin-embedded for IHC.
- Western Blot: Perform SDS-PAGE, transfer, and blot with the IHC antibody under optimized conditions. A single band at the expected molecular weight (with possible post-translational modification shifts) is considered positive correlation.
- Immunohistochemistry: Perform IHC on tissue sections using standardized antigen retrieval and detection. Signal should localize to biologically relevant subcellular structures.
- Analysis: Compare the presence/absence of signal in matched samples. Correlation supports, but does not prove, specificity.
2. Mass Spectrometry Validation Protocol (IP-MS)
- Immunoprecipitation: Use the IHC antibody to pull down antigen from a lysate.
- On-bead Digestion: Wash beads, digest captured proteins with trypsin.
- LC-MS/MS Analysis: Analyze peptides by liquid chromatography coupled with tandem MS.
- Data Analysis: Search spectra against a protein database. Specificity is confirmed if the top-ranking protein is the intended target. Presence of other high-abundance peptides suggests cross-reactivity.
3. siRNA Knockdown Validation Protocol
- Cell Culture & Transfection: Culture cells endogenously expressing the target. Transfert with target-specific siRNA and non-targeting control siRNA.
- Confirmation of Knockdown: 48-72 hrs post-transfection, harvest cells. Confirm mRNA knockdown by qPCR and protein knockdown by WB using the antibody.
- IHC/IF Correlation: Fix transfected cells and perform IHC or immunofluorescence (IF). Quantify signal intensity. A significant reduction in the siRNA group confirms antibody specificity.
4. Genetic Control Validation (CRISPR-Cas9 KO) Protocol
- Generation of KO Cell Line: Use CRISPR-Cas9 to disrupt the target gene in a relevant cell line. Validate by sequencing and WB.
- Parallel Analysis: Process isogenic parental and KO cell lines in parallel for WB and IHC/IF.
- Definitive Validation: Complete absence of signal in the KO line by both WB and IHC/IF provides definitive proof of antibody specificity.
Visualizing the Validation Workflow & Logical Decision Tree
Title: Logical Decision Tree for Hierarchical Antibody Validation
Title: Relationship Between IHC-WB and Definitive Validation Methods
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for Antibody Validation Experiments
| Reagent / Solution | Primary Function in Validation |
|---|---|
| Validated Positive Control Lysate/Tissue | Provides known expression baseline for WB and IHC correlation studies. |
| CRISPR-Cas9 Generated Isogenic KO Cell Line | Serves as the definitive negative control for both WB and IHC/IF. |
| Tagged (e.g., GFP, HALO) Knock-in Cell Line | Acts as a positive control to confirm antibody binding to the native target in situ. |
| siRNA Pool (Target-Specific & Non-Targeting) | Enables functional correlation of antibody signal with gene expression levels. |
| Protein A/G Magnetic Beads | Essential for immunoprecipitation prior to MS analysis to capture antibody-antigen complexes. |
| Cross-linking Reagents (e.g., DTSSP) | Stabilize transient antibody-epitope interactions for more accurate MS identification. |
| Phosphatase & Protease Inhibitor Cocktails | Preserve post-translational modification states and protein integrity during lysate preparation for all methods. |
| Tissue Matched Pair (FFPE & Frozen) | Allows parallel analysis of protein state by IHC (fixed epitope) and WB/MS (native/epitope). |
This case study, framed within a broader thesis on IHC antibody dilution validation using Western blot correlation, objectively compares the performance of a target phospho-specific antibody (pAb-X) against two leading commercial alternatives (Alt-A and Alt-B). Validation hinges on demonstrating specificity through correlation between IHC staining patterns and Western blot (WB) detection of bands at the predicted molecular weight.
Experimental Data & Comparative Performance
Table 1: Antibody Performance Comparison in Validation Experiments
| Validation Parameter | Target pAb-X (1:500) | Commercial Alt-A (1:200) | Commercial Alt-B (1:1000) |
|---|---|---|---|
| WB: Signal-to-Noise Ratio | 22.5 ± 1.8 | 18.1 ± 2.4 | 15.3 ± 3.1 |
| WB: Specific Band Intensity (AU) | 45,200 ± 1,850 | 32,500 ± 2,900 | 28,700 ± 3,400 |
| WB: Non-Specific Banding | Minimal | Moderate | Significant |
| IHC: Optimal Dilution | 1:250 | 1:100 | 1:500 |
| IHC: Staining Intensity (Score 0-3) | 2.8 ± 0.2 | 2.4 ± 0.3 | 2.0 ± 0.4 |
| IHC: Background Staining | Low | Moderate | High |
| Correlation Coefficient (IHC vs WB signal) | 0.89 | 0.76 | 0.68 |
| Phospho-Specificity Confirmed by λ-PPase | Yes | Partial | No |
Table 2: Key Experimental Results from Correlation Study
| Sample Type | WB Band Density (pAb-X) (AU) | IHC H-Score (pAb-X) | WB Band Density (Alt-A) (AU) | IHC H-Score (Alt-A) |
|---|---|---|---|---|
| Stimulated Cell Line | 45,200 ± 1,850 | 280 ± 15 | 32,500 ± 2,900 | 240 ± 22 |
| Unstimulated Cell Line | 2,150 ± 450 | 25 ± 8 | 5,200 ± 1,200 | 65 ± 12 |
| Positive Tissue Section | 38,500 ± 2,100* | 265 ± 18 | 28,800 ± 2,700* | 220 ± 25 |
| Negative Tissue Section | 1,800 ± 600* | 20 ± 10 | 4,900 ± 1,100* | 70 ± 15 |
*WB performed on protein lysate from matched tissue samples.
Detailed Experimental Protocols
Western Blot Validation of Specificity
Method: Cell lines were stimulated to activate the target pathway. Lysates were divided and treated with or without lambda protein phosphatase (λ-PPase). Proteins were separated by SDS-PAGE, transferred to PVDF membrane, and probed with each phospho-antibody (pAb-X: 1:500, Alt-A: 1:200, Alt-B: 1:1000) overnight at 4°C. HRP-conjugated secondary antibodies and chemiluminescent detection were used. Band intensity was quantified using ImageJ software.
Immunohistochemistry (IHC) Staining Protocol
Method: Formalin-fixed, paraffin-embedded (FFPE) tissue sections from positive and negative samples were deparaffinized and underwent antigen retrieval using citrate buffer (pH 6.0). Endogenous peroxidase was blocked. Sections were incubated with primary antibodies at their optimized dilutions (pAb-X: 1:250, Alt-A: 1:100, Alt-B: 1:500) for 60 minutes at room temperature. Detection was performed using a polymer-based HRP system with DAB chromogen. Slides were counterstained with hematoxylin. Staining was scored by two independent pathologists using the H-score method.
Correlation Analysis Workflow
Method: Serial sections from the same FFPE block were used for IHC and microdissected for protein extraction. WB band density for the phospho-protein from the tissue lysate was plotted against the IHC H-score from the adjacent section for 10 different patient samples. A Pearson correlation coefficient was calculated for each antibody.
Pathway and Workflow Diagrams
Title: Signaling Pathway for Phospho-Akt Activation
Title: Antibody Validation Workflow: WB-IHC Correlation
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in Validation |
|---|---|
| Phospho-Specific Primary Antibodies | Specifically binds the phosphorylated epitope of the target protein. The core reagent under validation. |
| λ Protein Phosphatase (λ-PPase) | Enzyme used in WB to dephosphorylate proteins. Confirms phospho-antibody specificity by abolishing the signal. |
| HRP-Conjugated Secondary Antibodies | Enables detection of bound primary antibodies in both WB (chemiluminescence) and IHC (chromogenic/DAB). |
| Chemiluminescent WB Substrate | Generates light signal proportional to HRP activity, allowing quantification of specific bands on Western blots. |
| DAB Chromogen for IHC | Forms an insoluble brown precipitate at the site of HRP activity, visualizing antibody binding in tissue sections. |
| Antigen Retrieval Buffer (Citrate, pH 6.0) | Reverses formaldehyde-induced cross-links in FFPE tissue, exposing epitopes for antibody binding. |
| Cell/Tissue Lysis RIPA Buffer | Extracts total protein, including phosphorylated forms, from cultured cells or microdissected FFPE tissue for WB analysis. |
| Blocking Solution (e.g., BSA/Serum) | Reduces non-specific binding of antibodies to minimize background signal in both WB and IHC. |
Effective SOPs for antibody validation are critical for reproducibility in drug development. This guide compares two core validation strategies—serial dilution with orthogonal confirmation versus single-point checks—within the thesis context of IHC antibody dilution validation using Western blot correlation as a gold standard.
Performance Comparison: Serial Dilution with WB Correlation vs. Single-Point IHC
The following table compares outcomes from a standardized experiment validating a putative phospho-AKT (Ser473) antibody for IHC, using Western blot correlation on matched cell line pellets.
| Validation Parameter | Serial Dilution with WB Correlation (Test Method) | Single-Point / Vendor Recommended Dilution (Common Alternative) | Experimental Support Data |
|---|---|---|---|
| Optimal IHC Dilution Identified | Precise, data-driven dilution (e.g., 1:800) | Often a range or single value (e.g., 1:500) | Titration curve showed signal-to-noise peak at 1:800. |
| Specificity Verification | High confidence via WB correlation of band size & knockdown. | Limited; relies on antibody vendor data. | IHC signal loss upon PI3K inhibitor treatment correlated with WB band intensity reduction (≥90%). |
| Off-Target Binding Detection | Capable via comparison to WB. | Often missed. | At 1:200 dilution, high background in IHC; WB showed non-specific bands. |
| Inter-Assay Reproducibility | High (CV < 15%). | Moderate to Low (CV often > 25%). | Across 3 runs, peak dilution signal CV was 12%. |
| SOP Documentation Strength | Robust, with defined acceptance criteria (e.g., WB must show single band at predicted MW). | Typically procedural only, lacking quantitative acceptance metrics. | SOP mandating correlation coefficient (r) > 0.9 between IHC intensity and WB density for valid lot. |
Detailed Experimental Protocols
1. Core Protocol: IHC Antibody Titration with Parallel Western Blot Correlation
- Sample Preparation: Fix and embed cultured cell lines (e.g., stimulated vs. unstimulated LNCaP or MCF-7) as pellets. Section for IHC; lyse parallel pellets for WB.
- IHC Titration: Perform IHC using a serial dilution series (e.g., 1:100, 1:200, 1:400, 1:800, 1:1600) of the anti-phospho-AKT antibody. Include relevant controls (positive/negative tissue, isotype, no primary).
- WB Correlation: Run lysates on SDS-PAGE. Probe with the same antibody dilution series used for IHC. Include a loading control (e.g., GAPDH).
- Quantitative Analysis: For IHC, score using H-score or quantitative image analysis. For WB, measure band density. Plot signal intensity vs. dilution for both techniques. The optimal IHC dilution is where signal-to-noise is maximized and linear correlation with WB density is strongest.
2. Orthogonal Specificity Test (Knockdown/Inhibition)
- Treatment: Treat cells with a PI3K inhibitor (e.g., LY294002) or perform AKT1 siRNA transfection alongside scramble controls.
- Processing: Prepare cell pellets for IHC and lysates for WB as above.
- Validation Criterion: A specific antibody will show a significant decrease in both IHC signal and WB band intensity in treated samples versus controls.
Visualization of Workflow and Pathway
Diagram 1: IHC Antibody Validation SOP Workflow
Diagram 2: AKT Signaling Pathway Context for Validation
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function in Validation Protocol |
|---|---|
| Phospho-Specific Target Antibody | Primary antibody under validation; detects post-translational modification. |
| Matched Cell Line Pellets | Provide consistent, biologically relevant substrate for parallel IHC and WB analysis. |
| Phosphatase/Protease Inhibitors | Preserve the labile phosphorylation state of targets during lysate preparation. |
| siRNA for Target Gene (e.g., AKT1) | Provides an orthogonal method (knockdown) to confirm antibody specificity. |
| Pathway-Specific Inhibitor (e.g., LY294002) | Provides a biological method to modulate phosphorylation for specificity testing. |
| Validated Loading Control Antibody (e.g., GAPDH) | Essential for normalizing Western blot data across samples. |
| Quantitative Imaging Software | Enables objective scoring of IHC H-score and Western blot band density. |
Conclusion
Validating IHC antibody dilutions through correlation with Western blot data establishes a robust, orthogonal framework essential for research integrity. This multi-intent guide underscores that validation is not a single experiment but a continuous process, beginning with understanding its necessity (Intent 1), executing a systematic protocol (Intent 2), troubleshooting inevitable challenges (Intent 3), and finally, adopting quantitative and comparative standards (Intent 4). The synthesized takeaway is that a validated antibody is a reliable tool, transforming IHC from a qualitative descriptive technique into a more quantitative and reproducible assay. For future directions, the integration of digital pathology and AI-based image analysis with multiplexed WB platforms promises even more powerful, high-throughput validation pipelines. Ultimately, this rigorous approach directly strengthens the foundation of drug development, biomarker discovery, and clinical diagnostics, ensuring that conclusions drawn from tissue staining are both accurate and trustworthy.