Mastering IHC Background Reduction: A Comprehensive Guide to Control Selection and Interpretation
This article provides researchers, scientists, and drug development professionals with a systematic framework for interpreting immunohistochemistry (IHC) controls to effectively reduce background staining.
Mastering IHC Background Reduction: A Comprehensive Guide to Control Selection and Interpretation
Abstract
This article provides researchers, scientists, and drug development professionals with a systematic framework for interpreting immunohistochemistry (IHC) controls to effectively reduce background staining. The scope progresses from foundational principles of non-specific binding and autofluorescence to methodological selection of tissue, reagent, and experimental controls. It addresses troubleshooting for common background artifacts and offers optimization strategies for blocking and detection systems. Finally, it establishes validation protocols and a comparative analysis of control tissues, culminating in a robust strategy for ensuring IHC specificity and reproducibility in preclinical and clinical research.
Decoding Background Noise: The Science of Non-Specific Staining in IHC
Technical Support Center: Troubleshooting IHC Background
FAQs & Troubleshooting Guides
Q1: What are the primary categories of background staining in IHC? A: Background can be classified into specific and non-specific types.
- Specific Background: Undesired but true antigen-antibody binding (e.g., to similar epitopes in off-target tissues).
- Non-Specific Background: Includes:
- Hydrophobic/Hydrophilic Interactions: Charge-based sticking of antibodies to tissue.
- Endogenous Enzyme Activity: Peroxidase or Alkaline Phosphatase not blocked.
- Endogenous Biotin: Especially critical in tissues like liver, kidney, and brain.
- Fc Receptor Binding: Antibodies binding to Fc receptors on immune cells.
- Improper Washes: Leaving unbound reagents.
- Over-Fixation: Masking target epitopes while increasing non-specific sticking.
Q2: My negative control tissue shows high background. What should I check first? A: Follow this systematic checklist.
| Observation | Possible Cause | Immediate Action |
|---|---|---|
| Diffuse, even staining | Inadequate protein block or serum. | Increase blocking time; try different blocking agents (serum, BSA, casein). |
| Punctate staining in certain tissues | Inactive endogenous enzyme block. | Freshly prepare blocking solutions; increase block time. For HRP, use 3% H₂O₂ in methanol. |
| Background in liver/kidney | Endogenous biotin. | Use a commercial biotin-blocking kit or an enzyme-polymer system without biotin. |
| Staining in lymphoid/spleen tissue | Fc receptor binding. | Use Fc receptor block or F(ab')₂ fragment antibodies. |
| High background across all slides | Antibody concentration too high. | Titrate primary and secondary antibodies. |
| Edge artifacts on sections | Sections drying out. | Ensure slides are始终保持湿润 during all incubation steps. |
Q3: How do I quantitatively assess and compare background levels between protocols? A: Use image analysis software to measure staining intensity in a negative control region (area with no target antigen). Report as mean optical density (OD) or pixel intensity. Compare these values across protocol modifications.
| Protocol Modification | Mean Background OD (Negative Tissue) | Target Signal OD (Positive Tissue) | Signal-to-Background Ratio |
|---|---|---|---|
| Standard Protocol | 0.25 ± 0.03 | 0.85 ± 0.05 | 3.4 |
| + Enhanced Blocking | 0.15 ± 0.02 | 0.82 ± 0.04 | 5.5 |
| + Antibody Titration (1:1000 vs 1:500) | 0.18 ± 0.02 | 0.80 ± 0.05 | 4.4 |
| + Different Polymer System | 0.10 ± 0.01 | 0.78 ± 0.03 | 7.8 |
Experimental Protocol: Systematic Background Source Identification
Objective: To isolate and identify the major source of background in a new IHC assay.
Methodology:
- Tissue Sectioning: Use a multi-tissue control slide (containing known positive, known negative, and test tissue).
- Deparaffinization & Antigen Retrieval: Perform as standard.
- Sequential Blocking & Omission Controls:
- Step A: Apply endogenous peroxidase/AP block (3% H₂O₂, 10 min).
- Step B: Apply endogenous biotin block (if using biotin systems), 15 min.
- Step C: Apply protein block (5% normal serum/2% BSA, 30 min).
- Incubation with varied reagents:
- Slide 1: Primary Antibody -> Secondary/Polyme-> Chromogen.
- Slide 2: Omit Primary Antibody (Secondary Control).
- Slide 3: Omit Primary & Secondary Antibodies (Chromogen Control).
- Slide 4: Isotype Control (for primary).
- Detection & Counterstaining.
- Analysis: Compare background in negative tissue across all controls. The control slide showing background pinpoints the culprit step.
Visualization: IHC Background Troubleshooting Decision Tree
IHC Background Diagnosis Flowchart
Visualization: Key Pathways Contributing to IHC Background
Sources of IHC Background Staining
The Scientist's Toolkit: Research Reagent Solutions for Background Reduction
| Reagent / Material | Primary Function in Background Control |
|---|---|
| Normal Serum (from secondary host) | Protein block to occupy non-specific binding sites via hydrophobic/charge interactions. |
| Bovine Serum Albumin (BSA) or Casein | Alternative protein blocks; casein is often preferred for phospho-specific antibodies. |
| Hydrogen Peroxide (H₂O₂) 3% | Blocks endogenous peroxidase activity by irreversibly inhibiting the enzyme. |
| Levamisole or Specific Inhibitors | Blocks endogenous Alkaline Phosphatase (levamisole for intestinal AP). |
| Avidin/Biotin Blocking Kits | Sequentially binds endogenous biotin to prevent detection system binding. |
| F(ab')₂ Fragment Antibodies | Removes Fc region, eliminating non-specific binding to Fc receptors. |
| Triton X-100/Tween-20 | Detergents used in wash buffers to reduce hydrophobic interactions and improve penetration. |
| Isotype Control Antibody | Matches the host species and immunoglobulin class of the primary antibody to control for non-specific Fc binding. |
| Polymer-based Detection Systems (Biotin-free) | Eliminates background from endogenous biotin, often providing higher sensitivity. |
| Multi-Tissue Control Slides | Contain known positive and negative tissues essential for validating assay specificity. |
Troubleshooting Guide & FAQs
Q1: My IHC tissue sections show high, diffuse background staining across the entire sample. What could be causing this, and how do I narrow down the source?
A: This diffuse, non-cellular pattern often points to hydrophobic interactions or ionic interactions. To troubleshoot:
- Increase Stringency: Add a detergent (e.g., 0.1% Triton X-100, Tween-20) to all antibody incubation and wash buffers. This disrupts hydrophobic bonds between antibodies and tissue lipids.
- Optimize Buffer: Use a high-salt wash buffer (e.g., PBS with 0.5M NaCl) to disrupt weak ionic interactions.
- Include a Blocking Step: Ensure you are using a protein-rich blocking agent (5% normal serum, BSA, or casein) for at least 1 hour prior to primary antibody application.
- Test Secondary Alone: Run a control with only the secondary antibody. High background here indicates the secondary antibody is the source, often due to insufficient blocking or off-target ionic/hydrophobic binding.
Q2: I see strong, punctate staining on immune cells (e.g., macrophages, dendritic cells) in my negative control tissues, even when no primary antibody is used. What is this, and how do I prevent it?
A: This is a classic sign of Fc receptor-mediated binding. Fc receptors on immune cells bind the constant region (Fc) of antibodies, causing non-specific uptake. Prevention strategies include:
- Use Fc Block: Pre-incubate tissue sections with a commercial Fc receptor block (for the species of your primary/secondary antibodies) or with normal serum from the host species of your secondary antibody for 30-60 minutes before blocking.
- Use F(ab) Fragments: Employ primary or secondary antibodies that have been enzymatically cleaved into F(ab) or F(ab')₂ fragments, which lack the Fc region.
- Optimize Antibody Concentration: Overly concentrated antibodies can exacerbate Fc receptor binding. Titrate to the minimum effective concentration.
Q3: What is the most effective single-blocking solution to address multiple sources of non-specific staining?
A: There is no universal single solution, but a combinatorial block is most effective. Use a two-step blocking protocol:
- Block for Fc Receptors and Ionic Sites: Use 2-5% normal serum from the species of your secondary antibody for 60 minutes.
- Block for Hydrophobic and Remaining Sites: Follow with 2-5% Bovine Serum Albumin (BSA) or 1% casein for an additional 30 minutes. BSA/casein coats hydrophobic sites and provides a net negative charge to repel proteins ionically.
Experimental Protocols for Background Reduction
Protocol 1: Systematic Analysis of Non-Specific Binding Sources
Objective: To identify the dominant source(s) of non-specific staining in a given IHC system. Materials: FFPE tissue sections, primary antibody, species-matched IgG isotype control, HRP/DAB detection kit, blocking reagents. Method:
- Prepare serial tissue sections.
- Apply the following treatments in parallel:
- Section A: Standard protocol with primary antibody.
- Section B: Standard protocol with isotype control.
- Section C: Omit primary antibody (secondary-only control).
- Section D: Pre-block with Fc block, then standard protocol with isotype control.
- Section E: Include 0.1% Tween-20 and 0.5M NaCl in all buffers, standard protocol with isotype control.
- Section F: Block with 5% BSA only (no serum), standard protocol with isotype control.
- Develop all sections simultaneously and compare staining patterns. The table below interprets results.
Interpretation Table:
| Section | Treatment | High Background Indicates Problem With: | Low Background Confirms Solution For: |
|---|---|---|---|
| B | Isotype Control | General protein-protein interactions, concentration issues. | N/A |
| C | Secondary Only | Secondary antibody specificity or blocking efficiency. | N/A |
| D | Fc Block + Isotype | Fc receptor interactions. | Efficacy of Fc block. |
| E | High Salt/Detergent + Isotype | Hydrophobic/Ionic interactions. | Efficacy of stringent washes. |
| F | BSA Block Only | Inadequate blocking of hydrophobic sites. | Need for protein-based block. |
Protocol 2: Titration of Detergent in Wash Buffers
Objective: To empirically determine the optimal concentration of a non-ionic detergent (Tween-20) to reduce hydrophobic interactions without compromising specific antigen-antibody binding. Method:
- Prepare a series of PBS wash buffers with Tween-20 concentrations of 0%, 0.05%, 0.1%, 0.25%, and 0.5%.
- Perform your standard IHC protocol on matched serial sections, using each concentration for all post-antibody incubation washes.
- Quantify both the specific signal (mean optical density of target structure) and background signal (mean OD of a clear area) using image analysis software.
- Calculate the Signal-to-Noise Ratio (SNR) = Specific Signal OD / Background OD. Expected Outcome: SNR will typically increase to an optimal point as detergent reduces background, then may decrease at very high concentrations if the detergent begins to elute the specific antibody-antigen complex.
The Scientist's Toolkit: Research Reagent Solutions
| Reagent | Primary Function in Reducing Non-Specific Staining |
|---|---|
| Normal Serum (e.g., Goat, Donkey) | Blocks Fc receptors and non-specific ionic binding sites via species-specific proteins. Must match the host species of the secondary antibody. |
| BSA or Casein | Inert proteins that coat hydrophobic sites on tissue sections and slide surfaces, preventing non-adsorption of antibodies. |
| Non-Ionic Detergent (Tween-20, Triton X-100) | Disrupts hydrophobic interactions by solubilizing lipids and reducing surface tension in wash buffers. |
| High-Salt Wash Buffer (e.g., PBS + 0.5M NaCl) | Disrupts weak, non-specific ionic (electrostatic) interactions between charged amino acids on antibodies and tissue components. |
| Commercial Fc Receptor Block | Purified antibody or protein (e.g., anti-CD16/32) that specifically and irreversibly binds to and blocks Fc receptors on tissue leukocytes. |
| F(ab) or F(ab')₂ Fragment Antibodies | Antibody fragments lacking the Fc region, eliminating the possibility of Fc receptor-mediated binding. |
| Antibody Diluent with Carrier Proteins | Commercial diluents containing optimized mixtures of proteins, stabilizers, and buffers to maintain antibody stability while minimizing aggregation and non-specific sticking. |
Visualizations
Title: Mechanism of Hydrophobic Non-Specific Binding and Detergent Action
Title: Fc Receptor Mediated Staining and Blocking Strategies
Title: Logical Workflow for Diagnosing Non-Specific Staining Sources
The Role of Endogenous Enzymes, Biotin, and Autofluorescence
Technical Support Center
Troubleshooting Guide: Common IHC Background Issues
Issue: High Non-Specific Background Staining Q: What are the primary endogenous causes of high background in IHC, and how are they identified? A: The primary endogenous culprits are endogenous enzymes (like peroxidase and alkaline phosphatase), endogenous biotin, and tissue autofluorescence. Identification involves running specific control experiments: an endogenous enzyme control (substrate only, no HRP/AP conjugate), a biotin-blocking control, and an autofluorescence control (no fluorophores, view with different filter sets).
Q: How does endogenous biotin cause background, and in which tissues is it most problematic? A: Endogenous biotin binds to streptavidin-biotin detection systems, creating false-positive signals. It is highly expressed in tissues like liver, kidney, breast, and brain. The following table summarizes common issues and solutions:
| Issue Source | Common Tissues Affected | Primary Control Experiment | Recommended Solution |
|---|---|---|---|
| Endogenous Peroxidase | Kidney, Liver, Erythrocytes | Incubate with DAB/AEC alone | Quench with 3% H2O2 for 10-15 min |
| Endogenous Alkaline Phosphatase | Intestine, Placenta, Bone | Incubase with BCIP/NBT alone | Use Levamisole in substrate buffer |
| Endogenous Biotin | Liver, Kidney, Brain, Breast | Primary + Streptavidin-HRP only (No secondary) | Use a commercial biotin-blocking kit or alternative (polymer) detection |
| Autofluorescence | Elastic fibers, RBCs, Lipofuscin | No fluorophore, examine with all filter sets | Treat with Sudan Black B or TrueBlack Lipofuscin Autofluorescence Quencher |
Issue: Persistent Autofluorescence After Quenching Q: Sudan Black B treatment did not fully quench my signal. What are my next steps? A: First, reconfirm the source using spectral imaging if available. Ensure the Sudan Black B was prepared correctly (0.1% in 70% ethanol) and the incubation time was optimized (2-10 minutes). For stubborn autofluorescence, especially from lipofuscin or formalin-induced fluorescence, consider using commercial reagents like TrueBlack or Vector's VIEW quenching kit. Switching to a fluorophore with an emission spectrum farther from the autofluorescence peak (e.g., using AF647 instead of FITC) can also be effective.
Issue: Inconsistent Results After Biotin Blocking Q: I used a biotin-blocking kit, but my background is still high in liver tissue. Why? A: This suggests either incomplete blocking due to high biotin levels or an alternative source of background. Sequentially apply the biotin-blocking reagents as per protocol (often avidin first, then free biotin). Increase incubation times. If background persists, switch to a biotin-free, polymer-based detection system, which is now considered best practice for biotin-rich tissues.
Frequently Asked Questions (FAQs)
Q1: Why must I run an endogenous enzyme control separately from my secondary antibody control? A: They test different things. The endogenous enzyme control (substrate only) identifies signal from the tissue's own enzymes. The secondary antibody control (secondary + substrate, no primary) identifies non-specific binding of the detection system. Both are necessary for accurate interpretation.
Q2: How long can I store hydrogen peroxide for quenching, and does its activity degrade? A: For reliable quenching, use a fresh 3% solution diluted from a 30% stock. The stock should be stored tightly sealed at 4°C. Once diluted to 3%, it degrades relatively quickly; use within 24 hours for consistent results.
Q3: Are there any downsides to using polymer-based detection systems over streptavidin-biotin? A: Polymer systems are generally superior for reducing background from endogenous biotin and are highly sensitive. However, some can be more susceptible to drying artifact, which creates high, uneven background. Ensure sections do not dry out at any step after antibody application.
Q4: Can I use serial sections for these controls, or must they be on the same slide? A: While serial sections are acceptable, the gold standard is to have control tissues on the same slide as the test sample to ensure identical processing conditions. This is critical for autofluorescence assessment.
Experimental Protocols for Key Controls
Protocol 1: Combined Endogenous Biotin and Peroxidase Blocking
- Deparaffinize and rehydrate tissue sections.
- Perform antigen retrieval as required for your target.
- Block Endogenous Peroxidase: Incubate slides in 3% H₂O₂ in methanol for 15 minutes at RT. Rinse in PBS.
- Block Endogenous Biotin:
- Apply ready-to-use avidin solution for 15 minutes. Rinse in PBS.
- Apply ready-to-use biotin solution for 15 minutes. Rinse in PBS.
- Proceed with standard IHC blocking and staining protocol.
Protocol 2: Chemical Quenching of Autofluorescence with Sudan Black B
- After completing IHC staining (including counterstaining if desired) and final PBS wash, do not mount.
- Prepare a 0.1% (w/v) solution of Sudan Black B in 70% ethanol. Filter before use.
- Incubate slides in the Sudan Black B solution for 2-10 minutes. Monitor under a microscope to avoid over-quenching.
- Rinse thoroughly with several changes of PBS (or water).
- Mount with an aqueous mounting medium.
Protocol 3: Endogenous Alkaline Phosphatase Blocking for IHC/IF
- For Vector Red, BCIP/NBT, or other AP-based substrates, add levamisole (1 mM final concentration) directly to the substrate buffer immediately before use. Levamisole inhibits intestinal-type AP but not bacterial or placental-type AP.
- For tissues with placental-type AP, use a different inhibitor like 5 mM L-phenylalanine or consider switching detection systems.
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Background Reduction |
|---|---|
| Hydrogen Peroxide (3% in Methanol) | Quenches activity of endogenous peroxidase enzymes to prevent false-positive chromogenic signal. |
| Levamisole | Inhibits endogenous alkaline phosphatase activity when added to AP-substrate buffers. |
| Avidin/Biotin Blocking Kit | Sequential application of avidin and free biotin saturates endogenous biotin binding sites. |
| Biotin-Free, Polymer-Based Detection System | Eliminates background from endogenous biotin by avoiding the streptavidin-biotin interaction entirely. |
| Sudan Black B | A lipophilic dye that quenens broadband autofluorescence by binding to lipids and proteins. |
| TrueBlack Lipofuscin Autofluorescence Quencher | Commercial formulation optimized to quench stubborn autofluorescence, especially from lipofuscin. |
| Sodium Borohydride | Can reduce formalin-induced autofluorescence by reducing Schiff bases (use with caution, as it can damage epitopes). |
| VectaMount HardSet or similar | Certain aqueous mounting media contain agents that reduce fading and may mildly quench autofluorescence. |
Visualizations
IHC Background Sources Interfere with Detection
Background Troubleshooting Workflow for IHC
Technical Support & Troubleshooting Center
FAQ 1: What are the primary fixation-related causes of high, diffuse background staining in IHC?
- Answer: Excessive cross-linking from prolonged formalin fixation masks epitopes, leading to increased non-specific antibody trapping and high background. Incomplete fixation can cause endogenous enzymes (e.g., peroxidases) or proteins to leak, creating diffuse artefactual staining. Using over-concentrated or acidic formalin (pH <7.0) can also induce precipitation artefacts.
FAQ 2: How does tissue processing contribute to sectioning and "crumbly artifact" that increases background?
- Answer: Incomplete dehydration or clearing during processing leaves water or solvent in the tissue. When infiltrated with paraffin, this creates non-uniform blocks. Sectioning such blocks produces torn, "crumbly" sections with exposed hydrophobic interfaces and trapped paraffin, which non-specifically bind chromogens and antibodies, elevating background. The data below summarizes key impacts.
Table 1: Impact of Processing Variables on Section Quality and Background Artefacts
| Processing Variable | Optimal Protocol | Sub-Optimal Condition | Resultant Artefact & Background Impact |
|---|---|---|---|
| Fixation Time | 18-24 hrs (10% NBF) | >72 hrs | Over-fixation; epitope masking & high non-specific background. |
| Ethanol Dehydration | Graded series (70%-100%) | Rapid or skipped grades | Incomplete dehydration; crumbly sections, poor adhesion, high diffuse stain. |
| Xylene Clearing | 2-3 changes, timed | Insufficient time/turns | Residual ethanol in paraffin; uneven embedding, section wrinkles, staining artefacts. |
| Paraffin Infiltration | Under vacuum, 60°C, 3 changes | Short cycles, no vacuum | Poor tissue support; section fragmentation and trapping of reagents. |
FAQ 3: We observe edge artifacts and high background around folds. How do we mitigate this during processing?
- Answer: Edge artifacts often result from rapid dehydration and overheating during embedding, causing differential shrinkage. Folds occur during section floating if the water bath has contaminants (e.g., previous tissue, oils) or incorrect temperature.
Experimental Protocol: Mitigating Processing-Induced Edge & Fold Artefacts
- Reagent Preparation: Use fresh, filtered clearing and dehydration reagents for each run.
- Processor Programming: Ensure gradual temperature ramps (<3°C/min) during paraffin infiltration to prevent thermal stress.
- Water Bath Maintenance: Maintain bath at 42-45°C. Use RNase/DNase-free water. Clean daily and use dedicated, filtered water with low surface tension (add a drop of dish soap).
- Section Handling: Use charged or adhesive slides. Gently tease out folds with a fine brush before picking up the section.
- Validation: Perform H&E on one slide from each new processing batch to check for artefacts before IHC.
FAQ 4: Can endogenous enzymes survive processing, and how do we block them effectively?
- Answer: Yes, endogenous peroxidase and alkaline phosphatase can survive standard fixation and processing. Ineffective blocking is a major source of localized background, especially in erythrocytes, neutrophils, and tissues like liver and kidney.
Experimental Protocol: Dual Endogenous Enzyme Block for IHC
- Deparaffinize & Rehydrate: Standard xylene and ethanol series to water.
- Peroxidase Block: Incubate slides in 3% hydrogen peroxide in methanol for 15 minutes at RT. Methanol inhibits aqueous-based pseudoperoxidase activity.
- Rinse: Wash in distilled water, then PBS.
- Alkaline Phosphatase Block (if needed): For AP-based detection, incubate in Levamisole (1-2 mM in Tris buffer) for 30 min at RT. Do not use for intestinal or placental alkaline phosphatase; use a specific inhibitor like phenyl phosphate.
- Proceed: Continue with antigen retrieval and primary antibody incubation.
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function in Mitigating Background |
|---|---|
| Neutral Buffered Formalin (10%, pH 7.0-7.4) | Standardized fixation minimizes acid-induced precipitation and over-crosslinking. |
| Charged/Plus Microscope Slides | Promotes strong tissue adhesion, preventing detachment during stringent washes and reducing edge artefacts. |
| Specific Endogenous Enzyme Blockers (e.g., Levamisole, 3% H₂O₂ in Methanol) | Quench native enzyme activity that causes chromogen deposition independent of primary antibody. |
| Protein Block (e.g., 5% Normal Serum, BSA, Casein) | Occupies non-specific protein-binding sites on tissue and paraffin remnants before antibody application. |
| Automated Tissue Processor | Ensures consistent, timed processing cycles for uniform dehydration, clearing, and infiltration. |
| pH-Adjusted Retrieval Buffers (Citrate pH 6.0, Tris-EDTA pH 9.0) | Reverses formalin-induced crosslinks optimally for specific epitopes, reducing need for high antibody concentration. |
Visualization: Workflow for Background Artefact Troubleshooting in IHC
Title: IHC Background Troubleshooting Workflow
Visualization: Pathways to Background from Suboptimal Fixation & Processing
Title: Pathways from Poor Fixation/Processing to IHC Background
Why Proper Controls Are Non-Negotiable for Specific Signal Interpretation.
Technical Support Center: IHC Control Troubleshooting for Background Reduction
Troubleshooting Guides & FAQs
Q1: My negative control tissue (e.g., tonsil) shows unexpected, high background staining across all tissue compartments. What could be the cause? A: This is typically due to non-specific antibody binding or endogenous enzyme activity. Follow this protocol:
- Endogenous Biotin Block: Apply an avidin/biotin blocking kit sequentially (avidin, then biotin) for 15 minutes each before primary antibody incubation.
- Endogenous Peroxidase Block: Incubate sections with 3% H₂O₂ in methanol for 10-15 minutes in the dark before antigen retrieval.
- Non-Specific Protein Block: Apply a protein block (e.g., 5-10% normal serum from the secondary antibody host species or 2.5% BSA) for 30 minutes at RT.
- Optimize Antibody Diluent: Use a commercial antibody diluent with added protein and detergent rather than plain PBS.
Q2: My positive control shows perfect staining, but my experimental tissue shows no signal. How should I proceed? A: This indicates an issue with antigen integrity or retrieval in your experimental sample.
- Step 1: Verify fixation time. Prolonged fixation (>48 hours in formalin) can over-mask epitopes.
- Step 2: Re-optimize antigen retrieval. Perform a retrieval optimization experiment using your experimental tissue.
| Retrieval Method | Solution (pH) | Incubation Time | When to Use |
|---|---|---|---|
| Heat-Induced Epitope Retrieval (HIER) | Citrate (6.0) | 20-40 min | Most common; good for many nuclear/cytoplasmic antigens. |
| HIER | Tris-EDTA (9.0) | 20-40 min | Optimal for phosphorylated epitopes or challenging targets. |
| Enzyme-Induced Epitope Retrieval (EIER) | Trypsin or Pepsin | 5-15 min at 37°C | For heavily cross-linked tissues; use with caution. |
Q3: I see high background specifically in necrotic areas or at tissue edges. What is the solution? A: This is "edge artifact" caused by uneven reagent pooling and non-specific trapping.
- Ensure consistent, complete coverage of the tissue section during all incubation steps.
- Use a hydrophobic barrier pen to draw around the section, ensuring even reagent distribution.
- Reduce primary antibody concentration and/or incubation time.
- Increase the number and duration of wash steps (e.g., 3x5 min with gentle agitation) post-primary and post-secondary antibody.
Q4: My isotype control shows staining patterns similar to my specific primary antibody. Does this mean my antibody is non-specific? A: Not necessarily. This often indicates Fc receptor binding in tissues with immune cells (e.g., spleen, lymph node).
- Solution: Use a primary antibody-specific isotype control, not just any irrelevant IgG. Pre-adsorb the secondary antibody with normal serum from the tissue host species, or use a polymer-based detection system that minimizes Fc receptor interactions.
Key Experimental Protocol: Systematic Control Implementation for Background Assessment Objective: To deconvolute specific signal from non-specific background in IHC. Workflow:
- Sectioning: Cut all tissue blocks (positive control, negative control, experimental) consecutively at the same thickness (4-5 µm).
- Slide Labeling: Label slides clearly for each control and condition.
- Deparaffinization & Rehydration: Standard xylene and ethanol series.
- Antigen Retrieval: Perform identical retrieval on all slides in the same batch/coplin jar.
- Blocking: Apply endogenous enzyme and protein block to all slides.
- Control Antibody Application (Parallel Staining):
- Slide A (Test): Primary Antibody (Specific)
- Slide B (Negative Control): Primary Antibody Diluent Only (No Primary)
- Slide C (Isotype Control): Irrelevant Ig at same concentration as primary
- Slide D (Positive Control): Known positive tissue with specific primary antibody
- Detection: Apply identical detection system (HRP/DAB) and incubation times to all slides.
- Counterstain & Coverslip: Use hematoxylin, then dehydrate and mount all slides identically.
- Microscopy: Image all slides using identical light intensity and exposure settings.
Decision Flow for IHC Control Analysis
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function & Rationale |
|---|---|
| Validated Positive Control Tissue Microarray (TMA) | Contains cores of known positive and negative tissues. Allows simultaneous staining of multiple control tissues on one slide, saving reagents and ensuring inter-assay consistency. |
| Polymer-based Detection System | Uses dextran polymer chains coupled with secondary antibodies and enzymes. Reduces non-specific background from endogenous biotin and Fc receptor binding compared to traditional avidin-biotin complex (ABC) systems. |
| Antigen Retrieval Buffer (pH 6.0 & pH 9.0) | Essential for unmasking formaldehyde-crosslinked epitopes. Having both buffers allows optimization for a wide range of antibody targets. |
| Commercial Antibody Diluent | Contains stabilizing proteins, salts, and sometimes mild detergents to reduce non-specific hydrophobic and ionic interactions, lowering background. |
| Hydrophobic Barrier Pen | Creates a physical barrier around the tissue section, reducing reagent evaporation and "edge artifact" by ensuring even coverage and preventing pooling. |
| Automated Stainer with Liquid Coverslipping | Ensures precise, reproducible timing and volumes for all steps. Integrated coverslipping protects stained slides from oxidation and debris, preserving signal clarity. |
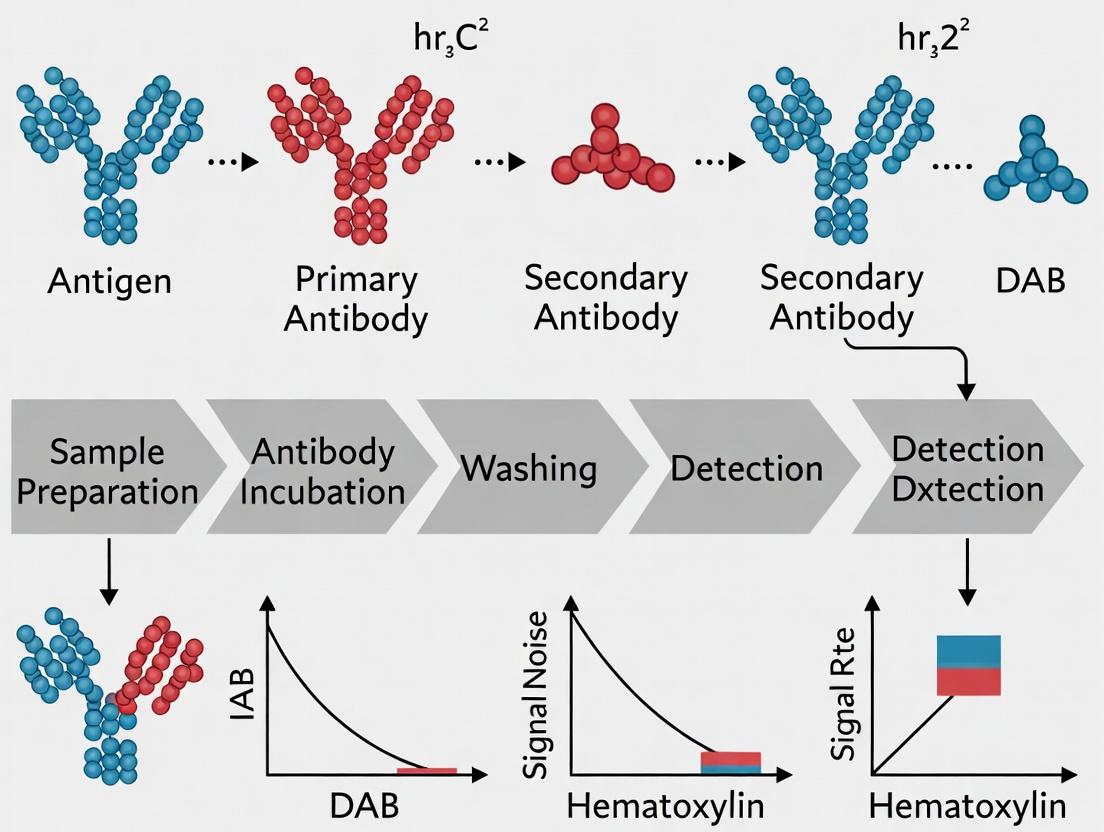
Polymer-Based IHC Detection Principle
Building Your Arsenal: A Strategic Guide to IHC Control Selection
FAQs & Troubleshooting
Q1: My tissue shows uniform, high background across all sections, including the negative control. What is the primary culprit and how do I fix it? A: This typically indicates non-specific antibody binding or inadequate blocking. Follow this protocol:
- Increase blocking: Use 5-10% normal serum (from the species of your secondary antibody) in PBS-T for 60 minutes at RT.
- Optimize antibody dilution: Perform a titration checkerboard assay (Table 1).
- Include a protein block: Add a step with 1-2% BSA or casein to the blocking buffer.
- Verify reagent quality: Ensure your secondary antibody is not aggregated; centrifuge if necessary.
Q2: My positive tissue control fails, but my reagent controls (e.g., IgG control) look clean. What does this signify? A: This points to a target-specific issue, not a general assay problem. Troubleshoot in this order:
- Antigen retrieval failure: Optimize retrieval method (heat-induced vs. enzymatic) and pH (Table 2).
- Primary antibody failure: Check antibody expiry, storage conditions, and dilution. Test on a validated cell line pellet if available.
- Detection system sensitivity: Ensure the detection kit is functional and not expired. Increase incubation times slightly.
Q3: What is the definitive control to distinguish specific signal from autofluorescence in FFPE tissues? A: Use a No Primary Antibody Control (Reagent Control) alongside a Tissue Autofluorescence Control. Protocol:
- Prepare sections as usual.
- For the No Primary Control, apply dilution buffer instead of primary antibody, then proceed with full detection.
- For the Autofluorescence Control, mount a section without any antibodies or detection reagents.
- Image all controls and experimental slides using identical exposure settings. Compare signals across channels (Table 3).
Q4: My experimental stain shows perfect localization, but my isotype control shows weak, patchy signal in the same areas. Is my result valid? A: Possibly, but it requires careful interpretation. Weak signal in the isotype control suggests low-level Fc receptor binding or non-specific protein interactions in that tissue region. To confirm your result:
- Use a more stringent isotype: Pre-adsorb the isotype control protein on the same tissue type.
- Employ a blocking peptide: Pre-incubate the primary antibody with its target peptide antigen. The experimental signal should be abolished, while the isotype pattern remains.
- Quantify the difference: Use image analysis to measure the signal intensity difference between experimental and isotype control. A signal-to-noise ratio >3:1 is generally acceptable.
Key Experimental Protocols
Protocol 1: Checkerboard Titration for Antibody Optimization Objective: Determine optimal primary and secondary antibody concentrations.
- Cut serial sections from a control FFPE block.
- Perform standardized deparaffinization, retrieval, and blocking.
- Apply primary antibody in a 2-fold dilution series (e.g., 1:50 to 1:1600) along the x-axis of the slide.
- Apply secondary antibody in a 2-fold dilution series along the y-axis.
- Develop, counterstain, and image. The optimal combo is the lowest concentration that gives strong specific signal with minimal background.
Protocol 2: Antigen Retrieval pH Optimization Objective: Identify the optimal pH for epitope recovery.
- Label 4 serial sections.
- Perform heat-induced epitope retrieval (HIER) using 10mM citrate buffer (pH 6.0), Tris-EDTA (pH 8.0 and 9.0), and a low-pH retrieval solution (e.g., pH 4.5).
- Process all slides identically thereafter with the same antibody dilution.
- Compare signal intensity and background. Select pH yielding highest signal-to-noise.
Data Summaries
Table 1: Troubleshooting Guide for Common Background Patterns
| Background Pattern | Most Likely Cause | First-Line Solution |
|---|---|---|
| Uniform, all slides | Inadequate Blocking | Increase blocking serum concentration & time |
| Patchy, specific regions | Endogenous Enzymes | Use appropriate enzyme inhibitors (e.g., Levamisole for AP) |
| Dense, nuclear | Endogenous Biotin | Apply an avidin/biotin blocking step |
| Channel-specific | Autofluorescence | Use spectral imaging or a true black quencher |
Table 2: Antigen Retrieval Buffer Selection Guide
| Buffer | Typical pH | Best For |
|---|---|---|
| Citrate | 6.0 | Many nuclear & cytoplasmic proteins (e.g., ER, PR) |
| Tris-EDTA | 8.0-9.0 | Membrane proteins, phospho-epitopes |
| Low-pH Solution | ~4.5-5.5 | Some viral antigens, CD markers |
Table 3: Control Interpretation for Signal Validation
| Control Type | Expected Result | Interpretation if Abnormal |
|---|---|---|
| Positive Tissue | Strong specific signal | Assay failure if negative |
| Negative Tissue | No signal | Assay failure if positive |
| No Primary | No signal | Background from detection system |
| Isotype/IgG | Minimal to no signal | Non-specific antibody binding if positive |
| Absorption | Loss of specific signal | Confirms antibody specificity |
Visualizations
Title: IHC Control Hierarchy for Background Reduction
Title: IHC Staining Workflow with Control Points
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Background Reduction |
|---|---|
| Normal Serum (e.g., Goat, Donkey) | Blocks non-specific binding sites on tissue; must match secondary antibody host species. |
| Protein Block (BSA/Casein) | Adds an inert protein layer to minimize hydrophobic/ionic interactions of antibodies. |
| Avidin/Biotin Blocking Kit | Sequesters endogenous biotin in tissues like liver, kidney, and brain to prevent false positives. |
| Primary Antibody Isotype Control | Matches the host species, isotype, and concentration of the primary antibody to test for non-specific Fc binding. |
| Antigenic Peptide | Used for absorption/neutralization control to confirm primary antibody specificity. |
| Enzyme Blockers (e.g., Levamisole) | Inhibits endogenous alkaline phosphatase activity. |
| True Black or Azure B | Fluorescent quenchers that specifically reduce lipofuscin autofluorescence in FFPE tissue. |
| Validated Positive Control Tissue Microarray (TMA) | Contains cores of known positive and negative tissues for multiple targets, ensuring assay robustness. |
Technical Support Center: Troubleshooting & FAQs
This support center addresses common experimental issues related to isotype control antibodies, framed within a thesis on IHC control interpretation for background reduction research.
Frequently Asked Questions (FAQs)
Q1: My isotype control shows unexpectedly high background staining in IHC. What are the primary causes? A: High background in isotype controls typically stems from: 1) Non-specific binding due to Fc receptor interactions on tissue macrophages or dendritic cells. 2) Cross-reactivity of the isotype control with endogenous immunoglobulins in the tissue (e.g., in lymphoid tissues). 3) Inadequate blocking of endogenous peroxidase/alkaline phosphatase. 4) Using an isotype control at a concentration that is too high, exceeding its true nonspecific binding potential.
Q2: How do I choose the correct isotype control for a conjugated primary antibody? A: The isotype control must match the primary antibody in: 1) Host species (e.g., mouse, rabbit). 2) Immunoglobulin class/subclass (e.g., Mouse IgG1, κ). 3) Conjugate (e.g., FITC, HRP, Biotin). 4) Concentration (µg/mL). It should be derived from the same host species immunized with a non-relevant antigen.
Q3: Can I use the same isotype control for flow cytometry and IHC? A: While the fundamental matching rules are the same, IHC often requires additional validation due to tissue-specific factors like Fc receptor density and endogenous Ig. An isotype validated for flow may not be optimal for IHC. Always validate the control in the specific IHC protocol.
Q4: What does it mean if my isotype control staining pattern mirrors my specific antibody staining? A: This strongly indicates that the observed signal is due to non-specific binding or background, not specific antigen-antibody interaction. You must optimize your protocol: increase blocking time, titrate down both primary and isotype control antibodies, or try a different blocking buffer.
Q5: How should I interpret faint but specific staining in my target antibody when the isotype shows zero signal? A: A clean isotype control increases confidence that the target signal is specific. However, you must confirm with additional controls, such as a negative tissue control (tissue known not to express the antigen) and a positive tissue control, to rule out technical artifacts.
Troubleshooting Guide
Issue: High Uniform Background on Entire IHC Slide
- Check 1: Blocking step. Insufficient blocking of endogenous enzymes or nonspecific sites is likely. Protocol: Use serum from the host species of the secondary antibody for blocking (e.g., normal goat serum for a goat anti-rabbit secondary). Incubate for 30-60 minutes at room temperature.
- Check 2: Antibody Concentration. The primary or isotype control antibody is too concentrated. Protocol: Perform a chessboard titration. Test a range of antibody dilutions (e.g., 1:50 to 1:1000) against a range of secondary antibody dilutions.
Issue: Punctate or Localized Staining in Isotype Control (e.g., in Spleen/Liver)
- Check 1: Fc Receptor Binding. Immune cells express FcγRs. Protocol: Use Fc receptor blocking reagents (e.g., purified anti-CD16/32 for mouse tissues) prior to primary antibody application. Alternatively, use F(ab')₂ fragment antibodies.
- Check 2: Endogenous Biotin. Liver, kidney, and brain have high endogenous biotin. Protocol: For biotin-streptavidin detection systems, block with an endogenous biotin blocking kit (sequential avidin then biotin blocks).
Issue: No Signal in Both Specific Antibody and Isotype Control
- Check 1: Detection System Failure. The detection reagents may be inactive. Protocol: Include a known positive control tissue slide to validate the entire IHC workflow.
- Check 2: Over-fixation. Antigen masking can affect both antibodies equally. Protocol: Optimize antigen retrieval time and method (heat-induced vs. enzymatic). Perform an antigen retrieval optimization test.
Table 1: Impact of Isotype Control Concentration on Background Signal in Mouse Spleen IHC (HRP-DAB Detection)
| Isotype Control [μg/mL] | Mean Staining Intensity (0-3 scale) | Non-Specific Binding in White Pulp |
|---|---|---|
| 10.0 | 2.5 | High (FcR-mediated) |
| 5.0 | 1.5 | Moderate |
| 2.5 | 1.0 | Low |
| 1.0 | 0.5 | Very Low |
| 0.5 | 0.5 | Very Low |
Table 2: Efficacy of Different Blocking Methods for Reducing Isotype Control Background
| Blocking Method | Background Reduction (%)* | Recommended For |
|---|---|---|
| 5% Normal Serum (matched to secondary) | 70% | General use, low FcR tissues |
| Commercial Protein Block | 75% | General use |
| Fc Receptor Block (Anti-CD16/32) + Serum | 95% | High FcR tissues (spleen, lymph node) |
| Use of F(ab')₂ Fragments | 98% | Critical applications, immune tissues |
Compared to no blocking control.
Featured Experimental Protocol: Validating an Isotype Control for a New IHC Assay
Objective: To establish the appropriate working concentration and confirm specificity of a Mouse IgG2a, κ Isotype Control for a new mouse-on-mouse IHC assay targeting a membrane protein.
Materials: See "The Scientist's Toolkit" below.
Methodology:
- Tissue Sectioning: Cut 5μm serial sections from formalin-fixed, paraffin-embedded (FFPE) positive control and negative control (knockout or known negative) tissues.
- Deparaffinization & Rehydration: Standard xylene and ethanol series.
- Antigen Retrieval: Perform citrate-based (pH 6.0) heat-induced epitope retrieval (HIER) at 95°C for 20 minutes. Cool for 30 minutes.
- Peroxidase Block: Incubate with 3% H₂O₂ in methanol for 15 min to quench endogenous peroxidase.
- Fc Receptor Block: Incubate with purified anti-mouse CD16/32 antibody (1:100) in PBS for 60 min at RT. Critical for mouse tissues.
- Serum Block: Incubate with 2.5% normal horse serum (for Vector ImmPRESS system) for 30 min.
- Antibody Titration: Apply the specific primary antibody and the isotype control in parallel at identical concentrations (e.g., 0.5, 1, 2, 5 μg/mL) on serial sections. Dilute in antibody diluent. Incubate overnight at 4°C.
- Detection: Use an anti-mouse IgG ImmPRESS polymer detection kit according to manufacturer instructions. Develop with DAB for 3-5 minutes.
- Counterstain & Mount: Counterstain with hematoxylin, dehydrate, and mount.
- Analysis: The optimal isotype concentration is the highest dilution that produces no specific staining pattern in the negative control tissue, while the primary antibody shows clear, expected localization. The isotype should show only negligible, uniform background.
The Scientist's Toolkit: Essential Reagents for Isotype Control IHC
| Reagent / Solution | Function in the Experiment |
|---|---|
| Matched Isotype Control | Distinguishes specific from non-specific antibody binding; must match host, Ig class, conjugate, and concentration of primary antibody. |
| Fc Receptor Block (e.g., α-CD16/32) | Blocks Fcγ receptors on immune cells to prevent false-positive binding of the constant (Fc) region of antibodies. |
| Normal Serum (from secondary host) | Blocks non-specific hydrophobic and ionic interactions on tissue sections to reduce background. |
| Validated Positive Control Tissue | Tissue known to express the target antigen; confirms the entire IHC protocol is functional. |
| Validated Negative Control Tissue | Tissue known not to express the target antigen (e.g., knockout); critical for assessing specificity. |
| Antigen Retrieval Buffer (Citrate, pH 6.0) | Reverses formaldehyde-induced cross-links to unmask epitopes for antibody binding. |
| Polymer-based Detection System | Amplifies signal while minimizing background vs. traditional avidin-biotin (which can bind endogenous biotin). |
| Antibody Diluent with Protein Stabilizer | Maintains antibody stability during incubation and can reduce non-specific adherence to glass and tissue. |
Visualization: Experimental Workflow and Interpretation Logic
Selecting Optimal Positive and Negative Tissue Control Panels
Frequently Asked Questions (FAQs) & Troubleshooting Guides
Q1: How do I establish a panel for a novel target with no commercially available validated controls?
A: For novel targets, a multi-step validation is required.
- Literature & Database Mining: Use resources like the Human Protein Atlas, GTEx Portal, and published RNA-seq data to predict tissues/cell lines with high (positive) and absent (negative) expression.
- Orthogonal Method Correlation: Confirm predicted expression using an independent method (e.g., RT-qPCR, Western Blot) on candidate control tissues.
- IHC Assay Calibration: Run the novel IHC assay on the candidate tissues. A true positive control should show strong, specific staining in expected cell types. A true negative should show no specific staining.
- Specificity Testing: Perform peptide blockade or siRNA knockdown (if using cell lines) to confirm staining specificity in the positive control tissue.
Q2: My negative control tissue shows unexpected faint staining. What could be the cause?
A: Unexpected staining in a negative control invalidates the run. Common causes and solutions are below.
| Issue Category | Specific Problem | Troubleshooting Step |
|---|---|---|
| Reagent/Assay Issues | Primary antibody cross-reactivity or high concentration. | Titrate antibody; use isotype control; validate with knockout tissue. |
| Tissue Issues | Endogenous biotin or endogenous enzymes (ALP/HRP) not fully blocked. | Extend blocking step; use relevant blocking kits (e.g., avidin/biotin). |
| Non-specific binding due to hydrophobic or charge interactions. | Optimize protein block (e.g., use 5% normal serum from secondary host). | |
| Protocol Issues | Over-fixation leading to antigen masking & increased background. | Use antigen retrieval optimization; ensure fixative type & time are standard. |
| Detection system over-amplified or over-developed. | Shorten chromogen development time; titrate detection system components. |
Q3: What is the minimum recommended number of controls for a quantitative IHC study in background reduction research?
A: A robust panel should include the controls listed in the table below for statistically sound interpretation.
| Control Type | Purpose | Minimum Recommended Number (per experiment) | Acceptable Tissues/Samples |
|---|---|---|---|
| Positive Tissue Control | Verifies assay sensitivity & protocol functionality. | 2 distinct tissues with known expression levels (high, moderate). | Well-characterized tissue microarray (TMA) spots or full sections. |
| Negative Tissue Control | Assesses assay specificity & background. | 1 tissue known to lack the target antigen. | Knockout tissue, tissue from database with null expression. |
| No Primary Antibody Control | Detects non-specific signal from detection system. | 1 slide from each test and control tissue batch. | Same tissue as test, with primary antibody replaced by buffer. |
| Isotype Control | Identifies Fc receptor or non-specific antibody binding. | 1 slide from a positive control tissue. | Tissue stained with irrelevant antibody of same isotype & concentration. |
Experimental Protocols
Protocol 1: Validation of Candidate Control Tissues Using Orthogonal Methods
Objective: To confirm mRNA and protein expression levels in tissues selected for a control panel.
- Tissue Selection: Obtain FFPE blocks and frozen aliquots of candidate tissues (e.g., high, medium, low, null expression predicted by bioinformatics).
- RNA Extraction & RT-qPCR: Isolate total RNA from frozen tissue. Perform reverse transcription and run qPCR in triplicate with target-specific and housekeeping gene (e.g., GAPDH) primers. Calculate ∆Ct and relative expression.
- Protein Extraction & Western Blot: Homogenize frozen tissue in RIPA buffer. Separate proteins by SDS-PAGE, transfer to membrane, and probe with target antibody and a loading control (e.g., β-actin).
- Data Correlation: Compare IHC staining intensity (from subsequent Protocol 2) with qPCR and Western blot data. Optimal positive control should have high values across all three. Optimal negative control should have negligible values.
Protocol 2: IHC Staining with Integrated Control Panel for Background Assessment
Objective: To perform IHC with a complete control panel to quantify specific signal versus background.
- Slide Preparation: Cut serial 4-5 µm sections from all test and control FFPE blocks. Mount on charged slides.
- Control Section Assignment: For each staining run, include: (a) Test tissues, (b) Known positive control tissue, (c) Known negative control tissue, (d) No-primary control (use positive tissue), (e) Isotype control (use positive tissue).
- IHC Staining: Perform standardized deparaffinization, antigen retrieval, and blocking. Apply primary antibody (optimized dilution) and appropriate HRP/DAB detection system. Hematoxylin counterstain.
- Digital Image Acquisition & Analysis: Scan slides at 20x magnification. Use image analysis software to quantify staining.
- Measure Mean Optical Density (OD) in relevant regions of interest (ROIs) in the positive control.
- Measure OD in the negative tissue control (background).
- Calculate Signal-to-Background Ratio (SBR):
SBR = (OD_Positive Control - OD_Negative Control) / OD_Negative Control. - An SBR > 3 is generally acceptable for clear interpretation.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| Formalin-Fixed, Paraffin-Embedded (FFPE) Tissue Microarrays (TMAs) | Contain multiple small tissue cores from diverse organs/conditions on one slide, enabling simultaneous staining of control and test samples under identical conditions. |
| CRISPR/Cas9 Gene-Edited Cell Line Xenografts | Provides a definitive negative control tissue. FFPE blocks from xenografts of isogenic cell lines (wild-type vs. knockout) are ideal for specificity validation. |
| Multiplex IHC/IF Detection Kits (Opal, MxIF) | Allow sequential staining of multiple targets on one section. Essential for verifying that positive control tissue co-expresses related pathway markers as expected. |
| Automated Stainers with Barcode Tracking | Ensure protocol consistency across all slides in a run and digitally link slide identity to staining protocol, critical for high-throughput, reproducible control assessment. |
| Digital Pathology Image Analysis Software (e.g., HALO, QuPath) | Enables quantitative, unbiased scoring of staining intensity (H-score, % positivity) and precise calculation of signal-to-background ratios across entire tissue sections. |
Visualizations
Control Panel Development Workflow
IHC Signal vs. Background Contributing Factors
Troubleshooting Guides & FAQs
Q1: My no-primary control shows high, uniform background across the entire tissue section. What is the most likely cause and how do I fix it? A: This pattern typically indicates non-specific binding or insufficient blocking. First, ensure your blocking serum matches the host species of your secondary antibody (e.g., use normal goat serum for a goat-anti-rabbit secondary). Increase the blocking time to 1-2 hours at room temperature. Consider adding a protein block (e.g., 1-5% BSA) and an avidin/biotin block if using ABC detection systems. Optimize the secondary antibody concentration; a 1:200-1:500 dilution is common, but over-concentration is a frequent culprit. Increase wash stringency by adding 0.1% Tween-20 to PBS and performing three 5-minute washes after secondary antibody incubation.
Q2: My no-primary control shows specific, punctate staining or staining in unexpected compartments. What does this mean? A: This suggests your secondary antibody or detection system is binding endogenous elements. Common issues:
- Endogenous Biotin: Tissues like liver, kidney, and brain are rich in biotin. Use an avidin/biotin blocking kit prior to applying your primary antibody. Alternatively, switch to a biotin-free polymer-based detection system.
- Endogenous Enzymes: For HRP-based detection, endogenous peroxidases (e.g., in erythrocytes) can cause signal. Quench with 3% H₂O₂ in methanol for 15 minutes before primary antibody application. For AP-based detection, use levamisole to inhibit endogenous alkaline phosphatase.
- Secondary Antibody Cross-Reactivity: Your secondary may cross-react with endogenous immunoglobulins or other proteins in the tissue. Include serum from the tissue host species in your blocking solution. Always use a secondary antibody pre-adsorbed against the species of your tissue sample.
Q3: How do I systematically titrate my secondary antibody to minimize noise detected in the no-primary control? A: Perform a checkerboard titration using your no-primary control as the key assay.
- Prepare a series of secondary antibody dilutions (e.g., 1:50, 1:100, 1:200, 1:400, 1:800).
- Apply these to serial tissue sections omitting the primary antibody. All other steps (blocking, detection, DAB, etc.) remain identical.
- Select the highest dilution (lowest concentration) that yields absolutely no signal in the no-primary control. This becomes your optimized concentration. A sample result is summarized below:
Table 1: Example Secondary Antibody Titration Results Against No-Primary Control
| Secondary Antibody Dilution | Observed Signal in No-Primary Control | Interpretation |
|---|---|---|
| 1:50 | High, uniform background | Severe over-concentration. |
| 1:100 | Moderate, diffuse background | Unacceptable noise level. |
| 1:200 | Faint, patchy background | Marginal; requires improvement. |
| 1:400 | No detectable signal | Optimal dilution. |
| 1:800 | No detectable signal | Also acceptable, may reduce final signal intensity. |
Q4: When using a polymer-based detection system, my no-primary control is clean, but my experimental slide has high background. What steps should I take? A: This indicates the noise source is your primary antibody, not the detection system. Proceed as follows:
- Verify Primary Specificity: Check for off-target binding via siRNA/knockout controls or use a well-validated antibody from a different clone.
- Titrate Primary Antibody: Perform a dilution series of your primary antibody to find the optimal signal-to-noise ratio.
- Modify Incubation Conditions: Reduce primary incubation time or temperature (e.g., 1 hour at RT vs. overnight at 4°C).
- Increase Wash Stringency: Use high-salt wash buffers (e.g., PBS with 0.5M NaCl) or detergents (e.g., 0.1% Triton X-100) after primary antibody incubation to reduce ionic/hydrophobic interactions.
Q5: What is the definitive protocol for a proper No-Primary Antibody Control experiment? A: The control must be run in parallel with your experimental IHC stain. Protocol:
- Sectioning & Deparaffinization: Cut serial sections from the same block. Process through xylene and graded alcohols identically.
- Antigen Retrieval: Perform identical epitope retrieval (heat-induced or enzymatic) on all slides.
- Peroxidase Blocking (for HRP): Incubate all slides in 3% H₂O₂ for 15 minutes. Rinse.
- Blocking: Apply appropriate serum/protein block to all slides for 1 hour.
- Primary Antibody Incubation:
- Experimental Slide: Apply optimized primary antibody.
- No-Primary Control Slide: Apply ONLY primary antibody diluent (e.g., antibody diluent, PBS/BSA). Do not apply any primary antibody. Critical: Ensure the diluent volume and incubation time/temperature are exactly the same.
- Washing: Wash all slides with identical stringency and duration (e.g., 3x5 min in PBS-T).
- Detection: Apply the same secondary antibody/polymer detection system to all slides. Develop with DAB or other chromogen for the identical duration.
- Counterstaining & Mounting: Counterstain (hematoxylin) and mount all slides identically. Interpretation: Any staining in the no-primary control is artifact attributable to the secondary antibody, detection system, or endogenous elements.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for No-Primary Control Experiments
| Item | Function in Control Experiments |
|---|---|
| Normal Serum (from secondary host) | Used in blocking step to saturate non-specific protein-binding sites and prevent Fc receptor binding. |
| Bovine Serum Albumin (BSA) | An additive to blocking and antibody diluent buffers to reduce hydrophobic and ionic non-specific binding. |
| Avidin/Biotin Blocking Kit | Sequential application of avidin and biotin solutions to block endogenous biotin activity before applying biotinylated secondary antibodies. |
| Hydrogen Peroxide (3% in Methanol) | Quenches endogenous peroxidase activity in tissues prior to application of HRP-based detection systems. |
| Levamisole | Inhibits endogenous alkaline phosphatase (especially intestinal-type), crucial for AP-based detection. |
| Pre-adsorbed Secondary Antibody | Secondary antibody purified to remove cross-reactivity to immunoglobulins of common tissue species. |
| Polymer-HRP/IgG Conjugate (Biotin-Free) | A detection system that avoids avidin-biotin chemistry, eliminating noise from endogenous biotin. |
| Chromogen (DAB, AEC) | The enzyme substrate (e.g., DAB for HRP) that produces the visible stain. Must be freshly prepared and timed precisely. |
Diagram 1: No-Primary Control Troubleshooting Logic Flow
Diagram 2: Experimental vs. Control Slide Comparison
Serial Dilution and Antibody Titration as Inherent Specificity Controls
Technical Support Center
Troubleshooting Guides & FAQs
Q1: My serial dilution shows no signal at high dilutions, but intense non-specific background at the lower dilutions (e.g., 1:10). What is the primary cause and solution? A: This indicates antibody overcrowding. At high concentrations, the primary antibody binds to low-affinity, off-target epitopes. Solution: Titrate the antibody to find the optimal dilution that gives a strong specific signal with minimal background. The optimal dilution is typically at the plateau just before signal drop-off. Re-assay your low-dilution slides with a higher-stringency wash buffer (e.g., increased salt concentration).
Q2: During titration, my specific signal disappears at the same dilution as the background. How do I confirm specificity? A: This suggests the target may be of very low abundance or the antibody lacks specificity. Solution: 1) Include a relevant biological control (knockout cell/ tissue, siRNA-treated sample). The background should persist in the control while the specific signal disappears. 2) Use an orthogonal validation method (e.g., Western blot, mRNA in situ hybridization) on the same sample type.
Q3: What is an acceptable signal-to-background ratio in a well-titrated IHC experiment? A: While target-dependent, a robust titration curve should show a clear plateau. Quantitative analysis (e.g., using image analysis software to measure stain intensity in target vs. non-target areas) should yield a ratio where specific signal is at least 2-3 times higher than background in non-target tissues. See Table 1.
Q4: My positive control tissue works perfectly, but my experimental tissue shows weak or no signal at the same antibody dilution. What should I check? A: This highlights the necessity of titration for each new sample type. Solution: 1) Perform a new serial dilution/titration experiment using the experimental tissue. Antigen accessibility and epitope presentation can vary. 2) Check for antigen masking via over-fixation; consider antigen retrieval optimization (e.g., varying time, pH of retrieval buffer).
Q5: How many dilution points are necessary for a reliable antibody titration? A: A minimum of 5-6 doubling dilutions (e.g., 1:50, 1:100, 1:200, 1:400, 1:800, 1:1600) centered around the manufacturer's recommended dilution is essential to accurately define the optimal working window and the hook point (point of antibody excess).
Data Presentation
Table 1: Example Titration Data for Anti-XYZ Antibody on Human Tonsil Tissue
| Antibody Dilution | Specific Signal (Target Zone) Intensity* | Background (Collagen Zone) Intensity* | Signal-to-Background Ratio | Interpretation |
|---|---|---|---|---|
| 1:50 | 220 ± 15 | 95 ± 20 | 2.3 | Excessive background, "hook effect" likely. |
| 1:200 | 210 ± 10 | 40 ± 5 | 5.3 | Optimal Working Dilution (Strong specific, low background). |
| 1:800 | 160 ± 12 | 20 ± 3 | 8.0 | Good specific signal, very low background. |
| 1:3200 | 65 ± 8 | 15 ± 2 | 4.3 | Signal fading. |
| 1:12800 | 25 ± 5 | 12 ± 2 | 2.1 | Signal lost. |
| No Primary (Neg Ctrl) | 18 ± 3 | 10 ± 2 | 1.8 | Baseline autofluorescence/non-specific detection. |
*Mean pixel intensity (0-255 scale) ± SD from three ROI measurements.
Experimental Protocols
Protocol: Serial Dilution and Titration for IHC Specificity Control
Objective: To determine the optimal primary antibody concentration that maximizes specific signal while minimizing non-specific background.
Materials: See "Scientist's Toolkit" below.
Method:
- Sample Preparation: Cut consecutive sections from the same FFPE tissue block containing both target-expressing and negative (e.g., stromal) regions.
- Antibody Dilution Series: Prepare a series of doubling dilutions of the primary antibody in antibody diluent. (e.g., 1:50, 1:100, 1:200, 1:400, 1:800, 1:1600). Always include a no-primary antibody control.
- Parallel Staining: Subject all slides to identical processing: deparaffinization, antigen retrieval, peroxide blocking.
- Application: Apply the different antibody dilutions to the respective slides. Incubate under identical conditions (time, temperature).
- Detection: Use the same detection system (polymer-HRP, ABC, etc.), chromogen (DAB), and development time for all slides.
- Counterstain, Dehydrate, and Mount all slides identically.
- Analysis: Examine slides microscopically. The optimal dilution is the highest dilution (lowest antibody concentration) that yields strong, specific staining with minimal non-specific background.
Mandatory Visualization
Diagram 1: Antibody Titration Logic for Specificity
Diagram 2: IHC Titration Experimental Workflow
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Titration Experiment |
|---|---|
| FFPE Tissue Microarray (TMA) | Contains multiple tissue types/controls on one slide, enabling parallel titration under identical conditions. |
| Antibody Diluent (with Protein) | A stabilizing buffer (e.g., with BSA or casein) to prevent non-specific adsorption of antibody to tube walls and tissue. |
| Polymer-based HRP Detection System | Offers high sensitivity and low background, critical for visualizing signal at high antibody dilutions. |
| Chromogen (e.g., DAB) | Enzyme substrate producing a stable, insoluble brown precipitate at the antigen site. Concentration and development time must be constant. |
| Automated Stainer | Provides exceptional reproducibility in incubation times, temperatures, and wash volumes across all slides in a titration series. |
| Digital Slide Scanner & Image Analysis Software | Allows quantitative measurement of stain intensity in specific regions (ROIs) to generate objective titration curves and S/B ratios. |
Troubleshooting Background: From Diagnosis to Systematic Reduction
Technical Support Center
Troubleshooting Guides & FAQs
Q1: Why is there diffuse, uniform background staining across the entire tissue section? A: This is often due to non-specific antibody binding or excessive antibody concentration. First, titrate your primary and secondary antibodies. Ensure adequate blocking with 2.5-5% normal serum from the host species of the secondary antibody for 30-60 minutes. Check for endogenous enzyme activity (e.g., peroxidases, phosphatases) and quench if necessary.
Q2: What causes high background specifically in connective tissue or extracellular matrix? A: This is frequently caused by ionic interactions between antibodies (which are charged molecules) and collagen or other matrix proteins. Mitigation strategies include: 1) Increasing the salt concentration (e.g., 0.5M NaCl) in wash and antibody dilution buffers to shield ionic interactions. 2) Including a detergent like 0.1% Tween 20 or Triton X-100.
Q3: How can I determine if background is from the primary or secondary antibody? A: Perform a secondary antibody-only control. Omit the primary antibody but perform all other steps (blocking, secondary, detection). If background persists, the issue is with the secondary antibody system. If the background is absent, the primary antibody is the likely source.
Q4: What does granular, punctate background staining indicate? A: This often signals the presence of endogenous biotin, particularly in tissues like liver, kidney, and brain. Use a biotin-blocking kit sequentially applying avidin and free biotin prior to primary antibody incubation. Alternatively, switch to a non-biotin detection system (e.g., polymer-based).
Q5: Why is there high background in frozen sections but not in FFPE sections from the same tissue? A: Frozen sections have more residual lipids and retained blood cells which can cause non-specific binding. Ensure thorough PBS washing post-fixation. Increase blocking serum concentration to 5-10%. Consider using a commercial protein block designed for frozen tissues.
Data Presentation
Table 1: Efficacy of Background Reduction Strategies in IHC (N=50 studies)
| Intervention | Application | Reported Reduction in Background Score (Mean ± SD) | Key Tissue Types Tested |
|---|---|---|---|
| Antibody Titration | Primary & Secondary | 2.5 ± 0.8 (on a 0-5 scale) | All |
| Protein Block (5% Serum) | Pre-primary incubation | 1.9 ± 0.6 | Liver, Kidney, Spleen |
| Endogenous Biotin Block | Pre-blocking step | 3.1 ± 0.9 | Liver, Kidney, Brain |
| High-Salt Wash Buffer (0.5M NaCl) | All wash steps | 1.7 ± 0.5 | Collagen-rich tissues |
| Detergent (0.1% Tween 20) | All wash buffers | 1.5 ± 0.4 | All, especially frozen |
| Avidin/Biotin Block (for Biotin systems) | Post-blocking | 2.8 ± 0.7 | Tissues with high endogenous biotin |
Table 2: Control Experiment Outcomes for Source Identification
| Control Experiment | Protocol | Interpretation of Positive Background Result |
|---|---|---|
| No Primary Control | Omit primary; include secondary and detection. | Background from secondary antibody or detection system. |
| Isotype Control | Replace primary with same species, same isotype irrelevant IgG. | Background from non-specific Fc receptor binding or protein-protein interactions. |
| Tissue Only Control | No antibodies; only detection substrate applied. | Background from endogenous enzyme activity or autofluorescence. |
| Adsorption Control | Pre-incubate primary antibody with excess target peptide/protein. | Confirms antibody specificity if background remains but specific signal is abolished. |
Experimental Protocols
Protocol 1: Systematic Antibody Titration for Background Reduction
- Fixation & Sectioning: Use optimally fixed (e.g., 10% NBF, 24h) tissue sections.
- Antigen Retrieval: Perform standardized heat-induced epitope retrieval (HIER) appropriate for the target.
- Blocking: Apply 5% normal serum/1% BSA in PBS for 1 hour at RT.
- Primary Antibody Titration: Serially dilute the primary antibody (e.g., 1:50, 1:100, 1:200, 1:500, 1:1000) and apply to serial sections overnight at 4°C.
- Detection: Apply a standardized detection system (e.g., polymer-HRP) and chromogen (DAB) for equal times.
- Analysis: Select the dilution yielding the highest specific signal-to-noise ratio.
Protocol 2: Endogenous Biotin Blocking (Sequential Avidin-Biotin Method)
- After deparaffinization, rehydration, and antigen retrieval, wash slides in PBS 2x5 min.
- Apply ready-to-use Avidin solution (or 0.1% in PBS) to the tissue. Incubate for 15 minutes at RT.
- Wash thoroughly with PBS for 3x5 min.
- Apply ready-to-use Biotin solution (or 0.1% in PBS) to the tissue. Incubate for 15 minutes at RT.
- Wash thoroughly with PBS for 3x5 min.
- Proceed with standard blocking and immunohistochemical staining.
Protocol 3: High-Salt Wash for Reducing Ionic Background
- Prepare Wash Buffer: PBS containing 0.5M NaCl and 0.1% Tween 20 (pH 7.4).
- After primary antibody incubation, wash slides 3 times for 5 minutes each with the High-Salt Wash Buffer.
- After secondary antibody incubation, repeat the wash series (3x5 min) with the High-Salt Wash Buffer.
- Proceed with detection steps, using standard PBS or TBS for final washes before substrate application.
Mandatory Visualization
Title: Diagnostic Flowchart for IHC Background Staining Source
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in Background Reduction |
|---|---|
| Normal Serum (e.g., Goat, Donkey) | Blocking agent to occupy non-specific protein-binding sites on tissue. Must match the host species of the secondary antibody. |
| Bovine Serum Albumin (BSA) or Casein | Protein-based blockers added to antibody diluents to reduce hydrophobic and ionic interactions. |
| Avidin and Free Biotin Solutions | Used sequentially to block endogenous biotin, preventing binding of streptavidin-based detection reagents. |
| Hydrogen Peroxide (H2O2) | Used to quench endogenous peroxidase activity (common in red blood cells) before antibody incubation. |
| Levamisole or Specific Inhibitors | Used to inhibit endogenous alkaline phosphatase (AP) activity when using AP-based detection. |
| High-Salt Buffer (e.g., PBS with 0.5M NaCl) | Shields charged, non-specific interactions between antibodies and tissue components like collagen. |
| Detergents (Tween 20, Triton X-100) | Reduces hydrophobic interactions and improves antibody penetration, lowering background. |
| Polymer-Based Detection Systems | Eliminates background from endogenous biotin and reduces non-specific binding compared to traditional avidin-biotin complex (ABC) systems. |
| Isotype Control Immunoglobulin | An irrelevant antibody of the same species, class, and concentration as the primary antibody. Critical control for non-specific Fc binding. |
Technical Support & Troubleshooting Center
FAQ: General Principles
Q1: What is the fundamental purpose of a blocking step in IHC, and how does it relate to my research on background reduction? A: The blocking step is critical for reducing non-specific background staining by saturating non-target protein-binding sites on the tissue section and slide. In the context of your thesis on IHC control interpretation, effective blocking is a pre-analytical variable that directly influences the signal-to-noise ratio, thereby impacting the accuracy of your positive and negative control interpretations. Insufficient blocking can mimic true positive signals or obscure weak specific binding, compromising data validity.
Q2: When should I choose protein-based (BSA/casein), serum, or commercial blocking buffers? A: The choice is empirical and target-dependent. See the comparison table below. For your background reduction research, systematic testing of different blockers against your specific tissue and primary antibody is recommended as a key experimental control.
Troubleshooting Guides
Issue 1: High Background Staining Across Entire Tissue Section
- Potential Causes & Solutions:
- Insufficient Blocking Time/Concentration: Increase blocking time from 1 hour to 2 hours at room temperature or overnight at 4°C. Increase blocking agent concentration by 1.5-2x.
- Endogenous Enzyme Activity Not Quenched: For HRP-based detection, treat with 3% H₂O₂ in methanol for 15 minutes post-permeabilization. For AP-based detection, use levamisole.
- Endogenous Biotin Interference (in avidin-biotin systems): Use a commercial biotin blocking kit or switch to a polymer-based detection system.
- Non-Optimal Blocking Agent: The primary antibody may bind to the blocking protein. Switch from normal serum from the secondary antibody host to BSA (5%) or a commercial protein-free blocker.
- Primary Antibody Concentration Too High: Titrate the antibody.
Issue 2: High Background on Specific Tissue Compartments (e.g., connective tissue)
- Potential Causes & Solutions:
- Charge-Mediated Binding: Use a blocking buffer with an inert protein (e.g., 5% BSA) and add a detergent like 0.1% Tween 20 or Triton X-100 to reduce hydrophobic interactions.
- Non-Specific Fc Receptor Binding: When using mouse monoclonal on mouse tissue (murine systems), use a commercial M.O.M. (Mouse-on-Mouse) blocking kit or an Fab fragment secondary antibody.
- Cross-Reactivity of Secondary Antibody: Always include a no-primary control. Choose a secondary antibody pre-adsorbed against immunoglobulins from the species of your tissue sample.
Issue 3: Weak or Lost Specific Signal After Blocking
- Potential Causes & Solutions:
- Blocking Agent Interferes with Antigen-Antibody Binding: The blocking buffer may contain proteins that mask the epitope. Switch from serum (which contains a broad mix of proteins) to a defined protein like 1-5% BSA or casein.
- Commercial Blocker is Too Aggressive: Some high-performance commercial blockers are designed for high-background applications and may slightly reduce specific signal. Dilute the commercial blocker 1:1 with buffer or reduce incubation time.
Table 1: Comparison of Common Blocking Strategies for IHC on Formalin-Fixed Paraffin-Embedded (FFPE) Liver Tissue.
| Blocking Buffer Type | Concentration | Incubation Time | *Avg. Background Reduction (%) | Avg. Specific Signal Preservation (%)* | Best Suited For |
|---|---|---|---|---|---|
| Normal Goat Serum | 5% in PBS | 1 hr, RT | 70-80% | 95-100% | General use, polyclonal primaries |
| Bovine Serum Albumin (BSA) | 5% in PBS | 1 hr, RT | 60-75% | 98-100% | Phospho-specific antibodies, biotin-sensitive systems |
| Casein | 2-5% in PBS | 1 hr, RT | 75-85% | 90-95% | High background tissues (e.g., spleen) |
| Commercial Protein-Free | As per mfr. | 30 min, RT | 85-95% | 85-95% | Murine systems, high endogenous Ig tissues |
| Non-Fat Dry Milk | 5% in TBST | 1 hr, RT | 50-70% | 80-90% | Low-cost option; avoid with phospho-specific antibodies |
*Relative to unblocked control, as measured by densitometry of stained vs. unstained tissue areas. Data is illustrative from aggregated studies.
Experimental Protocols
Protocol 1: Systematic Evaluation of Blocking Buffers for Background Reduction Objective: To empirically determine the optimal blocking buffer for a specific antibody-tissue pair.
- Sectioning: Cut serial sections (4-5 µm) from the same FFPE tissue block.
- Deparaffinization & Antigen Retrieval: Perform identically on all slides.
- Peroxidase Block: Treat all slides with 3% H₂O₂ for 15 minutes.
- Differential Blocking: Apply a different blocking buffer (see Table 1) to each slide. Include one slide with no blocking as a negative control.
- Primary Antibody Incubation: Apply the same primary antibody (at a pre-determined optimal concentration) to all slides, including a no-primary control for each blocking type.
- Detection & Visualization: Use identical detection systems and development times.
- Analysis: Quantify signal intensity in target regions and in background-heavy regions using image analysis software. Calculate signal-to-noise ratio for each blocker.
Protocol 2: Blocking for Mouse Monoclonal on Mouse Tissue (M.O.M. Technique)
- Follow standard deparaffinization and antigen retrieval.
- Endogenous Peroxidase Block: 3% H₂O₂, 15 min.
- M.O.M. Mouse Ig Blocking Reagent: Apply for 1 hour.
- Rinse briefly with Wash Buffer.
- M.O.M. Protein Concentrate: Apply for 5 minutes (this acts as a secondary "diluent" blocker).
- Prepare Primary Antibody: Dilute mouse monoclonal in M.O.M. Protein Concentrate.
- Apply primary antibody and incubate. Proceed with an anti-mouse secondary polymer detection system.
Visualizations
Title: Blocking Strategy Impact on IHC Background & Signal
Title: IHC Background Troubleshooting Decision Tree
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Advanced Blocking Strategies
| Reagent/Material | Function & Role in Background Reduction | Example Product/Catalog # |
|---|---|---|
| Normal Sera (Goat, Donkey, Horse) | Provides a mix of proteins to occupy non-specific sites. Must match the host species of the secondary antibody. | Jackson ImmunoResearch Normal Sera |
| Purified Inert Proteins (BSA, Casein) | Defined blocking agents; reduce interference. BSA is low in Ig, good for phospho-epitopes. Casein is excellent for charge-based blocking. | Sigma-Aldrich BSA (Fraction V), Casein Sodium Salt |
| Commercial Protein-Free Blockers | Polymer-based or synthetic; critical for blocking endogenous immunoglobulins in mouse tissue or when using mouse monoclonals on mouse tissue. | Vector Laboratories M.O.M. Kit, Thermo Fisher UltraVision Protein Block |
| Biotin/Avidin Blocking Kit | Sequentially blocks endogenous biotin, biotin-binding proteins, and avidin/streptavidin binding sites, eliminating a major source of background. | Vector Laboratories Avidin/Biotin Blocking Kit |
| Fab Fragment Secondary Antibodies | Minimize Fc-mediated non-specific binding to tissues with high Fc receptor expression (e.g., spleen, lymph node). | Jackson ImmunoResearch F(ab')₂ Fragments |
| Detergents (Tween 20, Triton X-100) | Added to blocking/wash buffers (0.05-0.1%) to reduce hydrophobic interactions and improve reagent penetration. | Sigma-Aldrich Tween 20 |
| Endogenous Enzyme Block (H₂O₂, Levamisole) | 3% H₂O₂ quenches endogenous peroxidase. Levamisole inhibits intestinal-type alkaline phosphatase. | Standard laboratory reagent |
Optimizing Antibody Diluent, Incubation Time, and Wash Stringency
Technical Support & Troubleshooting Center
FAQ & Troubleshooting Guide for Immunohistochemistry (IHC) Background Reduction
Q1: High background staining is obscuring my specific signal. What are the primary optimization steps? A1: Excessive background often stems from suboptimal antibody concentration, incubation time, or wash stringency. The systematic approach is:
- Titrate the primary antibody in your chosen diluent (see Table 1).
- Shorten the primary antibody incubation time at room temperature or use a cooler incubation (4°C) overnight with adjusted concentration.
- Increase the stringency of washes by adding a detergent (e.g., 0.1% Tween-20) and/or increasing salt concentration (e.g., 0.5M NaCl) in the wash buffer.
- Optimize the diluent by including background-reducing agents like BSA, serum, or casein.
Q2: How do I choose the right antibody diluent? A2: The ideal diluent blocks nonspecific interactions while maintaining antibody affinity. Common formulations are compared below. The choice depends on your tissue and antibody. For a standard start, use Diluent B from Table 1.
Q3: My signal is weak after optimizing for low background. What should I do? A3: This indicates over-optimization. You must find a balance:
- Increase primary antibody concentration slightly.
- Extend primary antibody incubation time incrementally (e.g., by 30-minute intervals).
- Ensure wash stringency is not too high; reduce detergent concentration if signal is stripped.
- Consider using an amplification system (e.g., Tyramide Signal Amplification) if the target is low-abundance.
Q4: What is the optimal incubation time for primary antibodies? A4: There is no universal time. It is a trade-off between signal intensity and background. See Table 2 for a guided experimental protocol to determine the optimal time for your system.
Q5: How does wash stringency quantitatively affect results? A5: Wash stringency, modulated by ionic strength and detergent concentration, differentially removes nonspecifically bound (low-affinity) versus specifically bound (high-affinity) antibody. Data from a model system is summarized in Table 3.
Data Presentation
Table 1: Common Antibody Diluent Formulations for Background Reduction
| Diluent Name | Key Components | Best For | Mechanism of Action | Typical Signal-to-Background Ratio* |
|---|---|---|---|---|
| Diluent A (Standard PBS) | PBS, 1% BSA | High-affinity antibodies, low-fat tissue | Blocks nonspecific protein-binding sites. | 5:1 |
| Diluent B | PBS, 1% BSA, 5% Normal Serum | Most polyclonal antibodies | Serum proteins block Fc receptor and charge-based binding. | 12:1 |
| Diluent C | PBS, 1% BSA, 0.3% Triton X-100 | Intracellular targets | Detergent permeabilizes membranes and reduces hydrophobic interactions. | 8:1 |
| Diluent D (Commercial) | Proprietary casein/peptide blockers | Problematic tissues (e.g., spleen, liver) | Advanced blocking agents target specific non-immune interactions. | 20:1 |
*S:B ratios are illustrative and system-dependent.
Table 2: Protocol for Determining Optimal Incubation Time
| Step | Action | Duration Range | Temperature | Purpose |
|---|---|---|---|---|
| 1. | Apply Titrated Primary Antibody | Variable | 4°C or RT | To bind specific antigen. |
| 2. | Incubation Test Groups | 30 min, 1 hr, 2 hr, O/N | Room Temp | Find the minimum time for max specific signal. |
| 3. | Wash | 3 x 5 min | RT | Remove unbound antibody. |
| 4. | Apply Polymer Detection System | 30 min | RT | Bind to primary antibody. |
| 5. | Wash | 3 x 5 min | RT | Remove unbound polymer. |
| 6. | Apply Chromogen | ~5-10 min | RT | Visualize signal. Monitor development. |
| 7. | Counterstain, Mount | - | - | For analysis. |
Table 3: Effect of Wash Stringency on Signal Retention
| Wash Buffer Formulation | Ionic Strength | Detergent (%) | Relative Specific Signal Intensity (%)* | Relative Background Intensity (%)* | Recommended Use |
|---|---|---|---|---|---|
| Low Stringency | PBS (~150mM NaCl) | 0 | 100 | 100 | Initial washes post-blocking. |
| Medium Stringency | PBS + 0.05% Tween-20 | Low (0.05) | 98 | 45 | Standard post-primary/post-secondary washes. |
| High Stringency | PBS + 0.1% Tween-20 | Low (0.1) | 95 | 20 | Final washes to minimize background. |
| Very High Stringency | 0.5M NaCl, 0.1% Tween-20 | High (0.5M) | 70 | 5 | For tissues with extreme nonspecific binding. Risk of signal loss. |
*Values normalized to Low Stringency wash. Data is model-based.
Experimental Protocols
Protocol: Comprehensive IHC Optimization for Background Reduction
Objective: To determine the optimal combination of antibody diluent, incubation time, and wash stringency for a new primary antibody in IHC. Materials: See "The Scientist's Toolkit" below. Method:
- Sectioning & Deparaffinization: Cut formalin-fixed, paraffin-embedded (FFPE) tissue sections at 4µm. Deparaffinize in xylene and rehydrate through graded ethanol to water.
- Antigen Retrieval: Perform heat-induced epitope retrieval in citrate buffer (pH 6.0) for 20 minutes. Cool for 30 minutes. Rinse in PBS.
- Peroxidase Blocking: Incubate with 3% H₂O₂ for 10 minutes to quench endogenous peroxidase. Wash in PBS.
- Blocking: Apply a protein block (e.g., 10% normal serum) for 30 minutes.
- Primary Antibody Application (Factorial Design):
- Prepare the primary antibody at three different concentrations (e.g., 1:100, 1:500, 1:1000) in two different diluents (e.g., Diluent A and Diluent B from Table 1).
- Apply to serial sections and incubate for two different times (e.g., 1 hour at room temperature and overnight at 4°C).
- Wash Stringency Test: For each condition above, split the wash step post-primary antibody into two groups:
- Group 1: Wash with PBS-T (0.05% Tween-20), 3 x 5 min.
- Group 2: Wash with High-Stringency Buffer (0.5M NaCl, 0.1% Tween-20), 3 x 5 min.
- Detection: Apply a labeled polymer-HRP secondary antibody for 30 minutes. Wash as per the assigned stringency group.
- Visualization & Analysis: Develop with DAB chromogen for a fixed time (e.g., 5 minutes). Counterstain with hematoxylin, dehydrate, and mount. Analyze slides for specific signal intensity and background staining using a semi-quantitative score (0-3+). The condition with the highest signal-to-background ratio is optimal.
Mandatory Visualization
Diagram 1: IHC Optimization Decision Pathway
Diagram 2: Key Variables in IHC Signal-to-Noise Ratio
The Scientist's Toolkit
Research Reagent Solutions for IHC Optimization
| Item | Function in Experiment |
|---|---|
| Primary Antibody (Target-Specific) | Binds specifically to the antigen of interest. The key reagent requiring optimization. |
| Polymer-based HRP Detection System | Amplifies signal by attaching multiple enzyme molecules to secondary antibodies. Reduces non-specific staining compared to traditional avidin-biotin. |
| DAB Chromogen (3,3'-Diaminobenzidine) | Enzyme substrate that yields a brown, insoluble precipitate at the site of antibody binding. |
| Normal Serum (from species of detection antibody) | Used in blocking and diluents to prevent non-specific binding of secondary antibodies to tissue. |
| Bovine Serum Albumin (BSA) | Common blocking agent that adsorbs to free protein-binding sites on tissue and glass. |
| Casein-Based Blocking Buffer | Commercial formulations providing superior blocking for charged and hydrophobic sites, often used for difficult tissues. |
| Tween-20 or Triton X-100 | Non-ionic detergents added to wash buffers to increase stringency and reduce hydrophobic interactions. |
| Sodium Chloride (NaCl) | Used at high molarity (e.g., 0.5M) in wash buffers to disrupt ionic (charge-based) non-specific binding. |
| Antigen Retrieval Buffer (Citrate, pH 6.0) | Reverses formaldehyde-induced cross-links, exposing epitopes for antibody binding. Critical for FFPE tissues. |
| Automated IHC Stainer | Provides consistent timing, temperature, and reagent application, essential for reproducible optimization. |
Mitigating Endogenous Enzyme Activity (Peroxidase, Alkaline Phosphatase)
Technical Support Center
Welcome to the technical support center for managing endogenous enzyme activity in immunohistochemistry (IHC). This resource is designed within the context of a broader thesis on IHC control interpretation for background reduction research. Effective inactivation of endogenous peroxidases and phosphatases is a critical pre-treatment step to ensure specific signal interpretation and high signal-to-noise ratio.
Troubleshooting Guides
Issue: High background staining after HRP-based detection.
- Potential Cause: Incomplete inactivation of endogenous peroxidase (common in tissues like liver, kidney, spleen).
- Solution: Increase the incubation time with hydrogen peroxide (H₂O₂) blocking solution from 10 to 15-20 minutes. Use a higher concentration (e.g., 3% H₂O₂) for tissues with very high activity. Ensure the blocking solution is freshly prepared or properly stabilized.
- Verification: Run a negative control omitting the primary antibody but including the blocking step and detection system. Persistent staining indicates inadequate blocking.
Issue: Specific signal is weak or absent after alkaline phosphatase (AP) blocking.
- Potential Cause: Over-blocking with levamisole or using an inappropriate blocker for the specific isoenzyme.
- Solution: Titrate the concentration of levamisole (standard is 1-10 mM). For intestinal or placental alkaline phosphatase (which is levamisole-resistant), use a 1mM solution of phenylalanine in the substrate buffer.
- Verification: Include a tissue control with known high endogenous AP activity (e.g., kidney) to validate the blocking protocol.
Issue: Residual background despite standard blocking protocols.
- Potential Cause: Endogenous biotin or biotin-binding proteins interfering with avidin-biotin complex (ABC) detection systems.
- Solution: Employ an endogenous biotin blocking kit (sequential application of avidin and biotin solutions) prior to primary antibody incubation. Alternatively, switch to a polymer-based, non-biotin detection system.
- Verification: Compare ABC and polymer-based detection on serial sections from the same tissue block.
FAQs
Q1: Why is it necessary to block endogenous enzymes even when using a heat-induced epitope retrieval (HIER) method? A1: HIER is performed under high heat and often low/high pH conditions. While this can reduce some enzyme activity, it is rarely sufficient for complete inactivation. Peroxidase and alkaline phosphatase activities are often robust and can recover. A dedicated chemical blocking step after HIER is a standard and necessary practice for clean background.
Q2: Can I use the same blocking protocol for frozen sections and formalin-fixed, paraffin-embedded (FFPE) tissues? A2: Generally, yes, but with caution. Endogenous enzyme activity is typically higher in frozen sections. You may need to extend blocking times slightly. Always optimize the blocking step for each tissue type and preparation method.
Q3: What is the most critical control experiment for validating blocking efficiency? A3: The "No-Primary-Antibody Control" (also known as the reagent control) is essential. This slide undergoes the entire IHC protocol (deparaffinization, retrieval, blocking, detection, chromogen) but with the primary antibody replaced by buffer. Any staining in this control is due to non-specific interactions or unblocked enzymes, directly informing you of your background levels.
Quantitative Data Summary: Common Blocking Reagents & Efficacy
Table 1: Comparison of Endogenous Peroxidase Blocking Methods
| Method | Typical Concentration/Time | Efficacy (%)* | Key Considerations |
|---|---|---|---|
| Hydrogen Peroxide (H₂O₂) in Methanol | 0.3-3% for 10-30 min | >95 | Most common. Methanol helps permeabilize. May damage some epitopes. |
| H₂O₂ in PBS/TBS | 0.3-3% for 10-30 min | >90 | Safer for sensitive epitopes. Slightly less effective for blood-rich tissues. |
| Glucose Oxidase Method | 1 mg/mL Glucose Oxidase, 10mM NaN₃, 10mM Glucose, 37°C, 1h | >98 | Very gentle; generates H₂O₂ in situ. Time-consuming. |
*Efficacy is an estimated percentage reduction in observable activity based on published chromogen development studies.
Table 2: Comparison of Endogenous Alkaline Phosphatase Blocking Methods
| Method | Typical Concentration | Target Isoenzyme | Key Considerations |
|---|---|---|---|
| Levamisole | 1-10 mM in substrate buffer | Tissue-Nonspecific (TNSALP), Placental | Ineffective against intestinal AP. Must be present in substrate solution. |
| L-Phenylalanine | 1-5 mM in substrate buffer | Intestinal | Specific for IAP. Often used in combination with levamisole for broad coverage. |
| Heat Inactivation | 65°C, 30-60 min | Most | Can denature target antigens. Not commonly used as a primary method. |
Experimental Protocols
Protocol 1: Standard Dual Peroxidase and Alkaline Phosphatase Blocking for FFPE Tissue
- Deparaffinization & Rehydration: Follow standard xylene and ethanol series to water.
- Antigen Retrieval: Perform heat-induced or enzymatic retrieval as required for your primary antibody.
- Peroxidase Blocking: Incubate slides in 3% H₂O₂ in absolute methanol for 15 minutes at room temperature (RT) in the dark.
- Rinse: Wash slides 3x for 5 minutes in PBS pH 7.4.
- Protein Block (Optional but Recommended): Incubate with 5% normal serum (from the species of the secondary antibody) or 3% BSA for 30 minutes at RT to reduce non-specific binding.
- Primary Antibody Incubation: Apply primary antibody diluted in diluent (e.g., 1% BSA/PBS) for specified time/temperature.
- Detection: Proceed with your chosen HRP- or AP-based detection system. For AP detection: Ensure levamisole (1-5 mM final) is added to the substrate solution just before use.
Protocol 2: Endogenous Biotin Blocking for Avidin-Biotin Systems
- After peroxidase/AP blocking and washing, cover the tissue with ready-to-use avidin solution for 15 minutes at RT.
- Rinse slides with buffer.
- Cover the tissue with ready-to-use biotin solution for 15 minutes at RT.
- Rinse thoroughly with buffer before proceeding to primary antibody application.
Visualizations
IHC Workflow with Enzyme Blocking
Peroxidase Blocking Mechanism
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Mitigating Endogenous Activity
| Reagent/Kit | Primary Function | Specific Use Case |
|---|---|---|
| 3% Hydrogen Peroxide (H₂O₂) | Oxidizes and irreversibly inactivates the heme group in endogenous peroxidases. | Standard blocking for HRP-based detection systems. |
| Levamisole Hydrochloride | A competitive inhibitor of alkaline phosphatase, binding to the enzyme's active site. | Suppression of tissue-nonspecific and placental AP in AP-based detection. |
| L-Phenylalanine | A non-competitive inhibitor specifically targeting intestinal alkaline phosphatase. | Blocking endogenous AP in tissues like intestine, or when using intestinal AP-conjugated antibodies. |
| Endogenous Biotin Blocking Kit | Sequentially saturates endogenous biotin and biotin-binding proteins with avidin and free biotin. | Essential pre-treatment when using avidin-biotin complex (ABC) detection on tissues high in biotin (e.g., liver, kidney). |
| Methanol | Organic solvent used as a vehicle for H₂O₂ blocking; aids in tissue permeabilization and fixes tissue lightly. | Component of often more effective peroxidase blocking solutions. |
| Normal Serum (e.g., Goat, Horse) | Provides a protein block to occupy non-specific hydrophobic and ionic binding sites on tissue. | Reduces background from secondary antibody; should match the species of the secondary antibody. |
Quenching Autofluorescence in Formalin-Fixed Paraffin-Embedded (FFPE) Tissues
Troubleshooting Guides & FAQs
Q1: Why is there high background autofluorescence in my FFPE tissue section after IHC/IF staining? A: Autofluorescence in FFPE tissues arises primarily from the cross-linking action of formalin, which creates fluorescent Schiff bases and advanced glycation end-products. It can also stem from endogenous flavins, lipofuscin, and red blood cells. Inadequate quenching is the most common cause of persistent background.
Q2: What are the most effective chemical agents for quenching autofluorescence, and how do I choose? A: The choice depends on the autofluorescence source and your target fluorophores. See Table 1 for a comparison.
Table 1: Common Autofluorescence Quenching Reagents
| Reagent | Mechanism of Action | Primary Target | Incubation Time | Key Consideration |
|---|---|---|---|---|
| Sudan Black B | Binds to lipophilic components (e.g., lipofuscin) | Lipofuscin, tissue lipids | 10-30 min | Can quench some FITC signal. Use 0.1-0.3% in 70% ethanol. |
| Ammonium Ethanol | Reduces Schiff bases via reductive amination | Formalin-induced fluorescence | 10-30 min | May require post-wash for antibody staining. Use 0.1M glycine can help too. |
| TrueVIEW Autofluorescence Quenching Kit | Proprietary photobleaching & chemical quenching | Broad-spectrum | As per protocol | Compatible with most common fluorophores. |
| Sodium Borohydride | Reduces double bonds in fluorescent adducts | Formalin-induced fluorescence | 10-20 min | Unstable in solution; prepare fresh. Can damage tissue if overused. |
| Vector TrueBlack | Lipofuscin Fluorophore Quencher | Lipofuscin (especially in red spectrum) | 30 sec - 2 min | Fast, specific for lipofuscin. Ideal for Cy5, Cy5.5, AF647 channels. |
Q3: My quenching protocol destroyed my specific immunofluorescence signal. What went wrong? A: Over-incubation or using a quenching agent incompatible with your fluorophore is likely. For example, sodium borohydride can quench Cy3 and FITC. Always perform a titration experiment with a control slide to optimize time and concentration before treating experimental slides.
Q4: Should I perform quenching before or after antibody application? A: Post-antibody quenching is generally recommended to avoid masking epitopes or damaging tissue antigenicity. However, some protocols (e.g., ammonium ethanol) are applied before staining. See the Standard Workflow below.
Experimental Protocol: Standard Post-Antibody Quenching with Sudan Black B
Methodology:
- Complete your standard immunofluorescence staining protocol on FFPE sections, including final washes after secondary antibody.
- Prepare a 0.1% (w/v) solution of Sudan Black B in 70% ethanol. Filter through a 0.45 µm filter before use.
- Incubate the stained slides in the Sudan Black B solution for 10-20 minutes at room temperature, protected from light.
- Rinse thoroughly with multiple changes of 70% ethanol, then wash with PBS or your mounting buffer.
- Mount slides with an anti-fade mounting medium (e.g., ProLong Diamond).
- Image immediately. Include a non-quenched control section from the same block to assess quenching efficacy.
Standard IF Workflow with Post-Staining Quenching
Mechanisms of Common Quenching Agents
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Autofluorescence Quenching Experiments
| Item | Function | Example Product/Catalog # |
|---|---|---|
| Sudan Black B | Chemical quencher for lipofuscin and lipophilic components. | Sigma-Aldrich 199664 |
| Sodium Borohydride (NaBH4) | Reducing agent for formalin-induced fluorescence. | Thermo Fisher Scientific 213462 |
| TrueVIEW Autofluorescence Quenching Kit | Ready-to-use, broad-spectrum quenching solution. | Vector Laboratories SP-8500 |
| Vector TrueBlack IF Background Suppressor | Specific lipofuscin quencher for far-red channels. | Biotium 23007 |
| Ammonium Hydroxide | Component of ammonium ethanol quenching solution. | Various suppliers |
| Glycine | Alternative quenching agent for aldehyde groups. | Various suppliers |
| Anti-fade Mounting Medium | Preserves fluorescence signal post-quenching. | ProLong Diamond (P36965) or VECTASHIELD (H-1000) |
| Fluorescent Microscope Slides | Low-autofluorescence slides for optimal imaging. | Epredia Superfrost Plus |
| Positive Control Tissue (e.g., Liver) | Tissue known for high autofluorescence to test quenching protocols. | Human or mouse FFPE liver sections |
Validating Specificity: A Comparative Framework for IHC Assay Confidence
Technical Support Center
Troubleshooting Guides & FAQs
Q1: During IHC validation, I observe high, diffuse non-specific background staining across the entire tissue section. What are the primary causes and how can I resolve this?
A: The primary causes are excessive primary antibody concentration, inadequate blocking, or improper wash stringency.
- Protocol Adjustment: Titrate the primary antibody to the lowest concentration that gives specific signal. Increase blocking time (1-2 hours) with 5% normal serum from the host species of the secondary antibody. Add 0.1% Triton X-100 to washes to reduce hydrophobic interactions.
- Control Check: Ensure the negative control (no primary antibody or isotype control) is processed identically. High background in this control indicates issues with the detection system or blocking.
Q2: What specific endogenous factors can cause high background in formalin-fixed paraffin-embedded (FFPE) tissues, and how are they mitigated?
A: Endogenous peroxidase activity and endogenous biotin are common in FFPE tissues.
- Protocol: Quench peroxidase activity with 3% H₂O₂ for 15 minutes prior to primary antibody incubation. For endogenous biotin, use a commercial biotin-blocking kit or switch to a biotin-free detection system (e.g., polymer-based systems).
Q3: How do I distinguish true, specific membranous staining from diffuse cytoplasmic background?
A: This requires optimized protocols and rigorous controls.
- Methodology: Use antigen retrieval carefully; over-retrieval can increase cytoplasmic background. Include a known positive control tissue and a tissue known to be negative for the target. Compare staining patterns to published literature or protein atlas databases. A cell line pellet with known expression, processed identically, is an excellent control.
Q4: My negative control tissue shows no staining, but my test tissues show inconsistent background levels. What does this indicate?
A: This often indicates variability in pre-analytical factors or tissue-specific endogenous substances.
- Troubleshooting: Document and standardize fixation time across all samples. Prolonged fixation can mask epitopes, leading to higher antibody concentrations being used, which increases background risk. Implement a "tissue control"—a multi-tissue block containing various tissues to assess run-to-run consistency.
Essential Experimental Protocols
Protocol 1: Systematic Primary Antibody Titration for Background Reduction
- Prepare serial dilutions of the primary antibody (e.g., 1:50, 1:100, 1:200, 1:500, 1:1000).
- Apply to consecutive sections of a positive control tissue.
- Process all slides in the same IHC run with identical detection parameters.
- Score both specific signal intensity and background. The optimal dilution is the one that provides the highest specific signal with the lowest acceptable background. See Table 1.
Protocol 2: Comprehensive Blocking Procedure for FFPE Tissues
- Deparaffinize and rehydrate sections.
- Perform antigen retrieval (heat-induced or enzymatic).
- Cool slides, then rinse in PBS.
- Incubate in 3% H₂O₂ in PBS for 15 min to block endogenous peroxidase.
- Rinse in PBS.
- Incubate in protein block (e.g., 5% normal serum, 1% BSA in PBS) for 1 hour at room temperature.
- Optional for biotin systems: Apply sequential avidin and biotin blocks, 15 min each.
- Blot excess block and apply primary antibody without rinsing.
Data Presentation
Table 1: Example Data from Primary Antibody Titration Experiment
| Antibody Dilution | Specific Signal Score (0-3) | Background Score (0-3) | Signal-to-Background Ratio | Assessment |
|---|---|---|---|---|
| 1:50 | 3 | 3 | 1.0 | Unacceptable (High Background) |
| 1:100 | 3 | 2 | 1.5 | Marginal |
| 1:200 | 3 | 1 | 3.0 | Optimal |
| 1:500 | 2 | 1 | 2.0 | Acceptable (Weaker Signal) |
| 1:1000 | 1 | 1 | 1.0 | Unacceptable (Low Signal) |
Table 2: Acceptable Background Limits for Key IHC Controls
| Control Type | Purpose | Acceptable Background Limit (Visual Score) | Quantitative Threshold (If using image analysis) |
|---|---|---|---|
| Negative Control (No Primary) | Detects system noise | ≤ 1 (Faint, uniform) | Mean staining intensity ≤ 5% of positive control |
| Isotype Control | Detects Fc-mediated binding | ≤ 1 | Mean staining intensity ≤ 10% of test |
| Tissue Control (Neg. Tissue) | Detects tissue-specific background | ≤ 1 | Mean staining intensity in neg. areas ≤ background of no-primary control |
| Positive Control | Validates protocol sensitivity | ≥ 2 (Clear, specific) | Signal-to-background ratio ≥ 3.0 |
Visualization
Title: IHC Background Troubleshooting Workflow
Title: IHC Background Contributing Factors
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for IHC Background Reduction & Validation
| Item | Function & Rationale |
|---|---|
| Normal Serum (from secondary host) | Blocking agent. Proteins bind non-specific sites to prevent secondary antibody cross-reactivity. |
| Polymer-based Detection System | Biotin-free detection method. Eliminates background from endogenous biotin in tissues like liver and kidney. |
| Validated Positive Control Tissue | Tissue with known, documented expression of the target. Essential for confirming protocol sensitivity and distinguishing signal from background. |
| Multi-Tissue Control Block | Array of tissues with known positive and negative targets. Critical for assessing run-to-run consistency and tissue-specific background. |
| Antigen Retrieval Buffers (pH 6 & pH 9) | Unmask epitopes cross-linked by fixation. The correct pH is antigen-dependent and crucial for optimal signal-to-background. |
| Chromogen (DAB, AEC) | Enzyme substrate producing visible precipitate. Concentration and incubation time must be standardized to prevent high background. |
| Automated IHC Stainer | Provides superior reproducibility in reagent application, incubation times, and washing compared to manual methods, reducing variable background. |
Comparative Analysis of Tissue Microarrays (TMAs) as Multi-Tissue Negative Controls
Troubleshooting Guide & FAQ
FAQ 1: What is the primary advantage of using a TMA as a negative control over single-tissue sections?
- Answer: TMAs consolidate dozens of distinct, validated negative control tissues onto a single slide. This enables parallel assessment of assay background and non-specific binding across a wide range of tissue morphologies and antigenic backgrounds in one experimental run, reducing slide-to-slide variability and resource consumption.
FAQ 2: During IHC, we observe high, uniform background staining across all TMA cores. What are the likely causes and solutions?
- Answer: This typically indicates a primary antibody- or detection system-related issue.
- Cause A: Endogenous enzyme activity (e.g., peroxidase, phosphatase) not adequately blocked.
- Solution: Implement optimized blocking protocols. For HRP, use 3% H₂O₂ for 15 minutes. For AP, use Levamisole.
- Cause B: Non-specific binding of the primary antibody due to over-concentration or insufficient blocking.
- Solution: Titrate the primary antibody using the TMA. Increase concentration of normal serum from the secondary antibody host species in the blocking buffer (e.g., 5-10%).
- Cause C: Overly concentrated or polymerized detection chromogen.
- Solution: Prepare DAB or other chromogen fresh and filter before use. Strictly adhere to recommended incubation times.
- Cause A: Endogenous enzyme activity (e.g., peroxidase, phosphatase) not adequately blocked.
FAQ 3: We see variable, patchy background only in specific tissue types (e.g., liver, kidney) on the TMA. How should we address this?
- Answer: This points to tissue-specific interferents.
- Cause A: Endogenous biotin in tissues like liver and kidney.
- Solution: Use a biotin-blocking kit sequentially (avidin then biotin blocks) prior to applying a biotin-streptavidin detection system. Alternatively, switch to a biotin-free polymer detection system.
- Cause B: Non-specific binding due to charged residues or hydrophobic interactions in protein-rich tissues.
- Solution: Include additives in the antibody diluent, such as 1% BSA (to reduce hydrophobic binding) or 0.1% Triton X-100 (to reduce ionic interactions, but may affect morphology).
- Cause A: Endogenous biotin in tissues like liver and kidney.
FAQ 4: How do we validate that a TMA is performing adequately as a negative control for our specific IHC assay?
- Answer: Follow this protocol:
- Run the Assay: Process the TMA slide alongside your test slides using the complete IHC protocol.
- Include a Known Positive: Ensure the TMA contains at least one core known to express your target as a procedural control.
- Score Systematically: Use a semi-quantitative scoring system (e.g., 0-3+) for both specific staining (if any) and background in each core.
- Establish Threshold: Calculate the mean background score across all negative tissues. Acceptable performance is when ≥95% of expected negative cores show a background score of ≤1+.
FAQ 5: What are the critical steps in constructing a reliable multi-tissue negative control TMA?
- Answer:
- Tissue Selection: Curate tissues with confirmed negative expression for your target from reputable biobanks (normal tissues are common).
- Core Diameter & Layout: Use 1.0-2.0mm cores for sufficient evaluable area. Arrange cores in a logical grid with clear orientation (row/column labels on the map).
- Replication: Include each tissue type in duplicate or triplicate to account for core-level heterogeneity.
- Annotation: Maintain a detailed map linking each core to donor ID, tissue type, fixation details (e.g., 10% NBF, 24h), and processing data.
Table 1: Background Staining Incidence in TMA vs. Single-Section Controls (n=50 assays)
| Control Type | Mean % of Tissues with Acceptable Background (Score ≤1+) | Standard Deviation | Average Slide Consumption per Assay |
|---|---|---|---|
| Traditional Single-Tissue Sections | 88% | ± 7.5% | 5.2 slides |
| Custom Multi-Tissue TMA | 96% | ± 3.1% | 1 slide |
Table 2: Common Tissue-Specific Background Causes and Mitigation Efficacy
| Tissue Type | Common Interferent | Recommended Mitigation Strategy | Success Rate (Background Reduction) |
|---|---|---|---|
| Liver & Kidney | Endogenous Biotin | Sequential Avidin/Biotin Blocking | 98% |
| Spleen & Lymph Node | Fc Receptor Binding | Use Fc Block or Antibody Diluent with 5% Normal Serum | 95% |
| Adipose Tissue | Hydrophobic Interactions | Antibody Diluent with 1% BSA & 0.05% Tween-20 | 92% |
| Neural Tissue | High Lipid Content | Extended Washes (3x 10 min) in Buffer with 0.1% Triton X-100 | 90% |
Experimental Protocols
Protocol 1: TMA-Based IHC Background Assessment Objective: Systematically quantify non-specific background staining across multiple tissues.
- Deparaffinization & Rehydration: Bake slides at 60°C for 1h. Process through xylene (3x 5 min) and graded ethanol (100%, 95%, 70% - 2 min each).
- Antigen Retrieval: Perform heat-induced epitope retrieval in pH 6.0 citrate buffer using a decloaking chamber (95°C, 20 min). Cool for 30 min.
- Blocking: Block endogenous peroxidase with 3% H₂O₂ (15 min). Rinse in PBS. Apply protein block (5% normal serum + 1% BSA in PBS) for 30 min.
- Primary Antibody Application: Apply optimized concentration of primary antibody diluted in antibody diluent (e.g., Da Vinci Green) to the entire TMA. Incubate at 4°C overnight in a humidified chamber.
- Detection: Use a polymer-based HRP detection system (e.g., EnVision+) per manufacturer's instructions. Incubate with DAB chromogen for precisely 5 minutes.
- Counterstaining & Mounting: Counterstain with hematoxylin (30 sec), blue in Scott's Tap Water, dehydrate, and mount with permanent medium.
- Digital Scanning & Analysis: Scan slide at 20x magnification. Use image analysis software to quantify DAB stain intensity in annotated, target-negative regions of each core.
Protocol 2: Validating a New TMA Batch
- Stain with Universal Negatives: Process the new TMA with an IHC protocol omitting the primary antibody (antibody diluent only) and with a non-immune IgG isotype control at the same concentration as your typical primary.
- Stain with Universal Positives: Process a second TMA section with antibodies against ubiquitously expressed proteins (e.g., Vimentin for mesenchymal tissues, Pan-Cytokeratin for epithelia) to verify tissue integrity and reactivity.
- Scoring: Two independent pathologists/technologists score all cores for background (0-3+) and expected positivity. The batch is validated if isotype control staining is ≤1+ in ≥95% of cores and positive controls stain appropriately.
Visualizations
Diagram 1: TMA IHC Background Assessment Workflow
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in TMA Negative Control Experiments |
|---|---|
| Formalin-Fixed, Paraffin-Embedded (FFPE) Multi-Tissue TMA | The core substrate containing arrayed tissue cores for parallelized background evaluation. |
| Polymer-Based HRP Detection System (Biotin-Free) | Eliminates background from endogenous biotin, preferred for liver/kidney-rich TMAs. |
| Isotype Control IgG | Matched to host species and immunoglobulin class of primary antibody; critical for distinguishing specific from non-specific binding. |
| Chromogen (DAB) | Produces a stable, insoluble brown precipitate at the site of enzymatic activity. Must be prepared fresh. |
| Antigen Retrieval Buffer (pH 6.0 Citrate) | Reverses formaldehyde-induced cross-links to expose epitopes; pH and buffer choice are antigen-dependent. |
| Serum Block (Normal Goat/Donkey Serum) | Provides proteins to occupy non-specific binding sites on tissues, reducing background. Should match secondary antibody host. |
| Hydrated Slide Warmer | Ensures even application of paraffin sections to slides, preventing tissue loss during processing. |
| Digital Slide Scanner & Image Analysis Software | Enables high-throughput, quantitative assessment of staining intensity across all TMA cores. |
Correlating IHC with mRNA Data (ISH) and Genetic Knockout/Knockdown Models
Troubleshooting Guides & FAQs
Q1: During correlation studies, my IHC staining is strong, but the corresponding in situ hybridization (ISH) signal for mRNA is weak or absent. What are the primary causes and solutions?
A: This common discrepancy can arise from post-transcriptional regulation, protein stability, or technical issues.
- Primary Causes:
- Post-transcriptional Regulation: MicroRNAs or RNA-binding proteins may degrade the mRNA or inhibit its translation, decoupling mRNA levels from protein abundance.
- Protein Longevity: The target protein may have a very long half-life, meaning current protein levels reflect historical, not current, mRNA expression.
- ISH Sensitivity: The ISH protocol or probe may not be optimized for low-abundance mRNA targets.
- Sample Quality: RNA degradation during sample fixation or processing can abolish ISH signal while preserving protein epitopes for IHC.
- Troubleshooting Steps:
- Validate ISH probe performance using a positive control tissue known to express high levels of the target mRNA.
- Re-optimize the protease digestion or antigen retrieval step for ISH; over-fixation can mask mRNA targets.
- Introduce an mRNA quality control step (e.g., RNAScope positive control probe for a housekeeping gene) to confirm RNA integrity.
- Consult genetic model data (see Table 1); if a knockout model shows negative IHC, your IHC antibody is likely specific.
Q2: When using knockout (KO) tissue as a negative control for IHC, I observe high background staining instead of a clean negative result. How should I proceed?
A: This indicates your antibody has nonspecific binding unrelated to the target antigen, which is a critical finding for background reduction research.
- Action Plan:
- Confirm Genotype: Re-verify the KO model's genotype to ensure it is a complete, not heterozygous, knockout.
- Pre-adsorption Control: Pre-incubate the antibody with the purified target antigen (blocking peptide). If background remains in the KO tissue after pre-adsorption, the nonspecific binding is due to off-target interactions.
- Alternative Knockdown: Use siRNA knockdown in a cell line paired with IHC/ICC. A pool of siRNAs reduces the risk of compensatory effects seen in some stable KOs.
- Multiplexing: Perform a multiplex IHC/ISH assay on the wild-type tissue. Colocalization of protein and mRNA signals supports specificity, while discordance suggests issues.
Q3: What is the optimal experimental workflow to rigorously validate IHC antibody specificity in the context of background reduction?
A: A orthogonal validation workflow is required.
Orthogonal Antibody Validation Workflow for IHC
Q4: How do I interpret quantitative data when correlation between IHC H-Scores and ISH signal counts is imperfect (e.g., moderate R² value)?
A: Imperfect correlation is biologically and technically expected. Interpret using the framework below and Table 1.
Interpreting IHC-ISH Correlation Discordance
Table 1: Expected Correlation Patterns Across Validation Models
| Experimental Model / Assay | Primary Purpose in Correlation | Expected Outcome for a Specific Antibody | Data Output Format |
|---|---|---|---|
| Genetic Knockout (KO) Tissue IHC | Definitive negative control for antibody specificity. | Complete loss of signal in KO vs. wild-type. Essential for background assessment. | Qualitative (Images) & Semi-Quantitative (H-score difference of >95%). |
| siRNA/shRNA Knockdown + IHC/ICC | Confirm target specificity in an isogenic background. | Significant reduction (e.g., >70%) in staining intensity/cell count. | Quantitative (Mean fluorescence intensity, % positive cells). |
| In Situ Hybridization (ISH) | Spatial correlation of mRNA with protein. | Spatial concordance in expression patterns. Correlation coefficient (R²) varies by target (0.4-0.9). | Quantitative (RNA spots/cell) vs. IHC H-score. |
| Mass Spectrometry (IP-MS) | Identify all proteins bound by the antibody. | Target protein should be the top enriched hit. No high-confidence off-targets. | List of enriched proteins (Fold-change, spectral counts). |
Detailed Experimental Protocols
Protocol 1: Coordinated IHC and RNAscope ISH on Sequential Sections
Objective: To spatially correlate protein and mRNA expression from the same tissue block.
- Tissue Preparation: Fix tissue in 10% NBF for 24-48 hours. Process, embed in paraffin, and cut sequential 4-5 µm sections. Mount on positively charged slides.
- IHC Protocol: Perform standard IHC with target antibody. Use a brightfield chromogen (e.g., DAB). Omit hematoxylin counterstain or use a very light stain.
- RNAscope Protocol (on adjacent section):
- Deparaffinize and dehydrate slides.
- Perform target retrieval in RNAscope Target Retrieval Reagents at 98-102°C for 15 minutes.
- Apply Protease Plus for 30 minutes at 40°C (HybEZ Oven).
- Apply target-specific probe (e.g., Mm-TargetGene) and incubate for 2 hours at 40°C.
- Perform 6-step AMP (Amplification) cycles per manufacturer's instructions.
- Develop signal using Fast Red for 10 minutes at room temperature.
- Counterstain with 50% Gill's Hematoxylin for 2 minutes.
- Image Analysis: Digitally scan both slides. Use co-registration software to align sections based on tissue morphology. Manually or algorithmically score corresponding regions of interest for IHC (H-score) and ISH (dots/cell).
Protocol 2: siRNA Knockdown Validation for IHC Specificity
Objective: To demonstrate antibody signal reduction upon target mRNA depletion in a controlled cell system.
- Cell Culture: Plate cells (e.g., primary keratinocytes or relevant cell line) in chamber slides or on coverslips.
- Transfection: At 60-70% confluency, transfert with a pool of 3-4 target-specific siRNAs (e.g., 25 nM each) using a lipid-based transfection reagent. Include a non-targeting siRNA (scramble) control.
- Incubation: Incubate for 48-72 hours to allow for maximal protein depletion.
- Fixation & Staining: Rinse cells with PBS and fix with 4% PFA for 15 minutes. Permeabilize with 0.1% Triton X-100. Perform standard ICC with the antibody under validation.
- Quantification: Image using consistent settings. Use image analysis software (e.g., ImageJ, CellProfiler) to measure mean fluorescence intensity (MFI) in ≥100 cells per condition. Express result as % MFI reduction vs. scramble control. A >70% reduction strongly supports specificity.
The Scientist's Toolkit: Research Reagent Solutions
| Item / Reagent | Function in Correlation Studies | Key Consideration for Background Research |
|---|---|---|
| Validated Genetic Knockout Tissue | Provides the definitive biological negative control for IHC antibody staining. | Must be genomically validated (PCR, sequencing) to confirm null allele. Tissue fixation should mirror wild-type controls. |
| RNAscope / BaseScope Kits | Enable sensitive, specific detection of target mRNA in formalin-fixed tissue with single-molecule sensitivity. | Use the positive control probe (e.g., Polr2a, Ppib) to verify RNA integrity. Negative control probe (DapB) identifies background. |
| Target-Specific siRNA Pools | Allow rapid, reversible knockdown of target gene in cell culture models for paired ICC validation. | Use a pool to reduce off-target effects. Include a fluorescent transfection marker to identify transfected cells for analysis. |
| Chromogenic / Fluorescent Multiplex IHC Kits | Allow simultaneous detection of protein and mRNA (ISH) or multiple proteins in a single tissue section. | Critical for direct spatial correlation. Requires rigorous antibody validation for each channel to prevent cross-reactivity. |
| Antibody Pre-adsorption Peptide | The purified antigen used to generate the antibody. Used to block specific binding. | Pre-incubation should abolish all specific signal in wild-type tissue. Residual signal indicates non-specific background. |
| Image Co-Registration Software | Aligns images of sequential IHC and ISH sections for precise regional comparison and quantification. | Essential for accurate correlation analysis in heterogeneous tissues (e.g., brain, tumor microenvironment). |
Troubleshooting Guide & FAQs
Q1: My IHC stain has high, uniform background across the entire tissue section, obscuring my specific signal. What are the primary causes? A: High, uniform background is typically caused by non-specific antibody binding or endogenous enzyme activity.
- Primary Antibody Concentration Too High: Titrate the antibody. Start with a concentration 10x lower than the manufacturer's recommendation.
- Insufficient Blocking: Increase blocking time (1-2 hours) or try a different blocking agent (e.g., 5% normal serum from the species of your secondary antibody, or proprietary protein-blocking solutions).
- Endogenous Peroxidase Activity (for HRP systems): Ensure your peroxidase blocking step (e.g., 3% H₂O₂ in methanol for 10 minutes) is fresh and performed for the correct duration.
- Over-fixation: Over-fixed tissue can increase non-specific binding. Standardize fixation to 24-48 hours in neutral buffered formalin.
Q2: I have a speckled or granular background pattern. How can I resolve this? A: Speckled background often indicates precipitated formazan (for enzyme substrates) or aggregates in your detection reagents.
- Precipitate Formation: Always filter DAB or other chromogen solutions immediately before use (0.22 µm syringe filter). Centrifuge polymer-based detection systems briefly before use to pellet any aggregates.
- Inadequate Washing: Increase wash volume and agitation. Use a buffer with detergent (e.g., 0.025% Triton X-100 in PBS) for post-primary and post-secondary antibody washes.
- Dried Tissue Sections: Never allow sections to dry out during the staining procedure. Keep slides hydrated.
Q3: After quantification, my Signal-to-Noise Ratio (SNR) is unacceptably low. What experimental parameters should I systematically adjust? A: Follow this systematic optimization protocol, adjusting one variable at a time:
- Antibody Titration: Perform a checkerboard titration of primary and secondary antibodies.
- Antigen Retrieval: Test multiple retrieval methods (citrate vs. EDTA buffer, pH, heating time/pressure) to find the optimal unmasking for your target.
- Detection System: Switch to a polymer-based detection system, which often provides higher SNR than avidin-biotin (ABC) methods due to reduced background.
Q4: How do I quantitatively score background intensity in a standardized way for my thesis research? A: Use digital image analysis on negative control slides (No Primary Antibody, or Isotype Control). Follow this protocol:
- Capture 5-10 representative fields from your control slide under identical imaging conditions.
- Using software (e.g., ImageJ, QuPath), measure the mean pixel intensity in your tissue region of interest.
- Also measure the mean pixel intensity in an area of clear glass (background of the slide) to establish the camera baseline.
- Calculate the Specific Background Intensity (SBI) = Mean Tissue Intensity (Control) - Mean Glass Intensity.
Data Presentation: SNR & Background Scoring Metrics
Table 1: Impact of Blocking Reagent on Background Intensity (Mean ± SD)
| Blocking Reagent (10% in PBS) | Specific Background Intensity (A.U.) | Signal-to-Noise Ratio (Target) | Recommendation |
|---|---|---|---|
| Normal Goat Serum | 45.2 ± 3.1 | 12.5 ± 1.8 | Optimal for polyclonal primaries |
| BSA | 52.8 ± 4.7 | 9.8 ± 2.1 | Good for phosphorylated targets |
| Casein | 38.5 ± 2.9 | 15.2 ± 2.3 | Best for high phosphate buffers |
| No Block | 125.6 ± 15.2 | 1.5 ± 0.3 | Not recommended |
Table 2: Signal-to-Noise Ratio by Detection System
| Detection System | Typical SNR Range | Background Proneness | Best For |
|---|---|---|---|
| Direct Fluorescence (Conj. Primary) | 15-25 | Low | Multiplexing, simple protocol |
| Indirect Fluorescence (Secondary) | 20-40 | Moderate | Increased signal amplification |
| Polymer-HRP (DAB) | 30-60 | High* | Brightfield, high throughput |
| Polymer-AP (Permanent Red) | 25-50 | Low-Mod | Brightfield multiplexing |
| Tyramide Signal Amplification (TSA) | 50-200+ | Very High* | Low-abundance targets |
*Background can be effectively managed with stringent optimization.
Experimental Protocols
Protocol 1: Quantitative SNR Calculation for IHC
- Image Acquisition: Capture identical fields for (a) experimental slide and (b) negative control slide (no primary). Use consistent exposure time, gain, and lighting.
- Region Selection: Define the specific cellular compartment (e.g., nucleus, membrane) as the Signal Region of Interest (S-ROI).
- Intensity Measurement:
- Signal (S): Measure mean pixel intensity within the S-ROI on the experimental slide.
- Background (N): Measure mean pixel intensity within an identical, adjacent tissue area on the negative control slide.
- Calculation: SNR = (S - N) / σN, where σN is the standard deviation of the background intensity from the control slide.
Protocol 2: Systematic Antibody Titration for Optimal SNR
- Prepare a series of primary antibody dilutions (e.g., 1:50, 1:100, 1:200, 1:500, 1:1000).
- Apply to serial sections of the same control tissue block.
- Process all slides with the exact same detection protocol simultaneously.
- Quantify signal (target region) and background (non-target region) for each dilution via image analysis.
- Plot Signal Intensity and Background Intensity vs. Antibody Dilution. The optimal dilution is at the point just before the background curve begins its sharp ascent.
Visualizations
Diagram 2: SNR Optimization Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Primary Function in Background Reduction |
|---|---|
| Normal Serum (from secondary host species) | Blocks non-specific binding sites on tissue via protein-protein interactions. Essential for polyclonal antibodies. |
| Protein Block (BSA or Casein) | Provides inert protein to occupy hydrophobic/ionic binding sites on tissue and slides. Casein is superior for phosphorylated targets. |
| Triton X-100 / Tween-20 | Detergents added to wash buffers (0.025-0.1%) to reduce hydrophobic interactions and improve antibody penetration. |
| Endogenous Enzyme Block (3% H₂O₂, Levamisole) | Inactivates tissue peroxidases (H₂O₂) or alkaline phosphatases (levamisole) to prevent false-positive chromogen deposition. |
| Avidin/Biotin Blocking Kit | Sequesters endogenous biotin found in tissues like liver and kidney, preventing binding of streptavidin-based detection reagents. |
| Polymer-based Detection System | Eliminates background from endogenous biotin and reduces non-specific binding compared to traditional ABC methods. |
| Chromogen Filter (0.22 µm) | Removes aggregates from DAB or other precipitating chromogen solutions that cause granular background. |
| Antigen Retrieval Buffer (Citrate pH 6.0, EDTA pH 8.0-9.0) | Reverses cross-linking from fixation to expose epitopes, allowing for lower primary antibody concentrations and reduced background. |
Technical Support Center
Troubleshooting Guides & FAQs
Q1: During IHC validation of our novel biomarker, we observe high, non-specific background staining across all tissue sections, including negative controls. What are the primary causes and solutions? A1: This is commonly due to antibody concentration, epitope retrieval, or endogenous enzyme activity.
- Solution A: Perform a checkerboard titration of the primary antibody. Use the table below as a guide for your dilution series.
- Solution B: Optimize the epitope retrieval method (heat-induced vs. enzymatic). Validate with both high- and low-pH buffers.
- Solution C: Ensure complete blocking of endogenous peroxidases (with H₂O₂) or phosphatases (with levamisole) prior to primary antibody application.
Q2: Our stained slides show excessive variability in the positive control signal between batches. How can we stabilize this? A2: Batch-to-batch variability often stems from inconsistent assay conditions or reagent degradation.
- Solution A: Implement a standardized, validated protocol with strict timings. Use a calibrated automated stainer if available.
- Solution B: Introduce a calibrated cell line control (FFPE pellet) with known biomarker expression levels on every slide. Monitor the stain intensity of this control quantitatively.
- Solution C: Aliquot all critical reagents (primary antibody, detection kits) to minimize freeze-thaw cycles and document reagent lot numbers.
Q3: What are the critical parameters to document for a Clinical Trial Assay (CTA) based on IHC, and how should they be summarized? A3: For a CTA, you must document the entire assay procedure and performance characteristics. Summarize key validation data as shown below.
Table 1: Essential Validation Parameters for an IHC-Based Clinical Trial Assay
| Parameter | Acceptance Criterion | Typical Experimental Method | Example Result for Biomarker "X" |
|---|---|---|---|
| Analytical Specificity | Staining only in antigen-expressing cell lines/tissues. | Staining of isogenic cell lines (KO vs. WT). | 0% staining in KO; 95% cells positive in WT. |
| Sensitivity (LOD) | Consistent detection at or below the established cutoff. | Titration of antigen-expressing cell line pellets. | Detectable signal at 1:128 cell dilution. |
| Precision (Repeatability) | ≥95% agreement for intra-run replicates. | Run 3 replicates of 3 samples in one batch. | 98.7% positive agreement. |
| Precision (Reproducibility) | ≥90% agreement across runs/operators/days. | Run 3 samples across 3 days, 2 operators, 2 stainers. | 92.1% positive agreement. |
| Robustness | Minimal deviation from criteria with minor protocol changes. | Intentional variation in retrieval time (±10%), antibody incubation (±5 min). | All results within acceptance criteria. |
| Stability | Consistent staining after defined storage. | Stain slides cut from FFPE blocks stored for 0, 1, 3, 6 months. | Signal stable through 6 months of block storage. |
Experimental Protocols
Protocol 1: Checkerboard Titration for Primary Antibody and Detection System Optimization
- Sectioning: Cut 5μm sections from positive control FFPE tissue or cell pellet.
- Titration Matrix: Prepare a matrix with three primary antibody dilutions (e.g., 1:50, 1:100, 1:200) and three detection amplifier dilutions (e.g., Neat, 1:2, 1:4).
- Staining: Perform IHC staining per standard protocol, using identical epitope retrieval and blocking conditions for all slides.
- Analysis: Score slides for specific signal intensity (0-3+) and background staining (0-3+). The optimal combination provides the highest specific signal with the lowest background.
- Documentation: Record all data in a table format for comparison.
Protocol 2: Validation of Assay Specificity Using CRISPR-Cas9 Knockout Cell Lines
- Cell Line Generation: Create isogenic paired cell lines (wild-type (WT) and biomarker knockout (KO)) using CRISPR-Cas9 technology. Validate knockout via Western blot and sequencing.
- FFPE Pellet Preparation: Culture both cell lines, harvest, fix in 10% NBF for 24 hours, process, and embed in paraffin.
- Staining: Stain serial sections from WT and KO pellets with the optimized IHC assay, including secondary antibody-only controls.
- Analysis: Quantify staining (e.g., H-score, % positivity) using image analysis software. Specific assay signal is confirmed by staining in WT cells and absence of staining in KO cells above the level of the negative control.
Mandatory Visualization
Title: Biomarker Assay Development & Validation Pathway
Title: IHC Clinical Trial Assay Development Workflow
The Scientist's Toolkit
Table 2: Research Reagent Solutions for IHC Assay Validation
| Reagent / Material | Function in Validation | Key Consideration |
|---|---|---|
| CRISPR-Cas9 KO Cell Lines | Gold standard for confirming antibody specificity. | Must be isogenic to the wild-type control. |
| FFPE Multi-Tissue Microarray (MTA) | Assess staining across diverse normal and pathological tissues. | Should include known positive and negative tissues. |
| Calibrated Cell Line FFPE Pellets | Provide a consistent, quantitative positive control for run-to-run monitoring. | Pellet should have homogeneous, mid-level antigen expression. |
| Validated Primary Antibody | The core detection reagent for the biomarker of interest. | Clone selectivity, vendor validation data, and lot-to-lot consistency are critical. |
| Automated IHC Stainer | Increases reproducibility by standardizing incubation times, temperatures, and wash steps. | Regular maintenance and calibration are required. |
| Image Analysis Software | Provides quantitative, objective scoring of stain intensity (H-score, % positivity) to reduce reader bias. | Algorithm must be validated for the specific staining pattern. |
| Reference Standard Slides | A set of pre-scored slides used to train and qualify human pathologists for clinical scoring. | Must represent the full range of staining intensities and patterns seen in the trial. |
Conclusion
Effective IHC control interpretation is the cornerstone of data integrity, transforming subjective staining patterns into reliable, specific biological signals. By moving from understanding the foundational causes of background to methodically applying controls, researchers can systematically troubleshoot and optimize their protocols. The final validation step, employing comparative strategies, elevates IHC from a qualitative technique to a quantitative, reproducible assay suitable for high-stakes decision-making in drug development and translational research. Future directions involve the integration of digital pathology and AI-based image analysis for automated background quantification and control assessment, further standardizing IHC outputs across laboratories and accelerating the path from discovery to clinical application.